
Expression and clinical significance of threonine and tyrosine protein kinase (TTK) in osteosarcoma
Introduction
Osteosarcoma (OS), which usually happens in the distal femur and proximal tibia, is one of the most common types of malignant bone tumors in children and adults (1-3). It remains the second leading cause in this age group of tumor-related death(1,4,5). Although the advances of understanding in the molecular and pathological mechanisms, the prognosis of OS is still poor due to its highly invasive (4,6). Because of this, more investigates are necessary for the early diagnosis and targeted therapy of OS.
TTK, which is known as the human monopolar spindle 1 (hMps1/TTK), is a dual serine/ threonine and tyrosine protein kinase (7). In various types of malignant tumors it had been reported that TTK plays an important role in the occurrence and development of tumors, such as triple-negative breast cancer (8,9), hepatocellular carcinoma (7,10), pancreatic cancer (11) and so on. But until now no studies about the role of TTK in OS has been reported.
In the present study, we first analyzed the expression level of TTK in OS through GEO dataset and then verified the results by using IHC. The goal of the study was to clarify whether TTK could offer additional help for the early diagnosis or even targeted therapy of OS.
Methods
Cell culture and main reagents
The human osteosarcoma cell lines, MG-63 and U2-OS, were cultured in RMPI1640 (Invitrogen) with 10% fetal bovine serum at 37 °C, 5% CO2. Fetal bovine serum (FBS) was obtained from HyClone (Logan, UT, USA). RPMI-1,640 media and 0.25% trypsin solution were purchased from Invitrogen (Carlsbad, CA, USA). Cells in logarithmic growth phase were used for western blotting.
Gene expression profiles
GEO dataset GSE33382 (Genome-wide gene expression analysis of high-grade osteosarcoma), GSE39262 (Human sarcoma cell lines and untransformed cells), GSE21257 (Genome-wide gene expression profiling on pre-chemotherapy biopsies of osteosarcoma patients who developed metastases within 5 yrs and patients who did not develop metastases within 5 yrs), GSE39055 (mRNA profiling and clinical outcomes in human osteosarcoma), was considered to investigate the expression level of TTK in sarcoma tumor tissue and normal tissue, and to investigate the correlation between TTK expression level and the metastasis, recurrence and survival time of OS.
Patients selected and TMA
Two independent TMAs (Alenabio Biotechnology Co., LTD, Xi’an, China) (Production number: BO244d and OS804b) were purchased. The TMAs consisted of 11 bone tissues and 41 OS tissues, including 38 men and 14 women. The mean age was 37.5 years old (ranged from 11 to 84 years old) (Table 1).

Full table
Immunohistochemical staining
The TMAs was deparaffined with standard pure xylene for 15 minutes three times at room temperature and hydrated in graded alcohols, phosphate buffer saline (PBS) was used to wash the TMA. Antigen retrieval was performed in boiling citrate buffer (PH 6.0) for 15 minutes. Then TMA was cooled down to room temperature in the buffers. After washing TMA in PBS for 5 minutes three times, 0.3% hydrogen peroxide phosphate-citrate buffer was used to block endogenous peroxidase activity for 10 minutes. Rinsed with PBS for 5minutes, TMA was incubated with primary antibody TTK (Sigma-Aldrich; dilution 1:100) for 12 hours at 4 °C. The TMAs were incubated with Poly-HRP Goat anti-rabbit (Maixin. Bio, FuZhou, China) for 30 minutes. Slides were dyed with diaminobenzidine for 5 minutes. Haematoxylin was used to counterstain the nucleus, followed by dehydration and mounted. Images of TMAs were taken using an Olympus BX40 microscope and CC-12 Soft-Imaging System (Olympus, Tokyo, Japan).
Evaluation of immunohistochemical staining
TTK showed both cytoplasmic and membrane expression. The TMAs was analyzed and scored for intensity [0-3] and frequency [0-4]. The intensity was scored as grade [0], negative; grade [1], weak intensity; grade [2], moderate intensity; grade [3], strong intensity. The frequency scores were respectively assigned when 0–25%, 26–50%, 51–75% and 76–100% of the tumor cell were positive. To use statistical analysis, TTK protein intensity and frequency were transformed into a composite expression score (CES) utilizing the formula CES = Intensity × Frequency. The range of CES was from 0 to 12. The CES was scored as negative (0), weak positive (1~4), positive (5~8), strong positive (9~12). All the staining scores were analyzed and determined by two experienced pathologist.
Synthesis of siTTK gene and gene transfection
The siRNAs molecules used for suppression of MG-63 and U2-OS cells TTK gene and the negative control siRNA (which does not target any sequence present in the MG-63 and U2-OS cell genomes) were all obtained from RiboBio Co Ltd (Guangzhou, China). Transient transfection of siRNA was carried out with the lipofectamine 2000 regent (Invitrogen, Massachusetts, USA) according to the manufacturer’s instructions.
Cell proliferation
MG-63 and U2-OS cells were seeded at 1×103/well in 96-well plates in RPMI-1640 medium supplemented with 10% FBS. Twenty-four hours later, the cells were transfected, and proliferation was determined at 0, 24, 48 and 72 h post-transfection using the Cell Counting Kit-8 (CCK-8), according to the manufacturer’s instructions.
Flow cytometry for detection of cell apoptosis
Cells were harvested by trypsinization and washed twice with cold phosphate-buffered saline (PBS). Then the cells were resuspended in Annexin-V-binding buffer and then stained with 5 µL of Annexin-VFITC solution and 10 µL of propidium-iodide (PI) solution for 15 min in dark at room temperature. Fluorescence was analyzed on a FACSCantoTM II spectrometer (BD Biosciences, San Jose, CA). Cells stained with FITC/PI were counted as apoptotic cells.
Western blotting
Total cell lysate was performed according to standard instructions. The lysates were resolved using 10% SDS-PAGE, transferred to PVDF membranes and immunoblotted with primary antibodies against TTK, p-ERK, T-ERK, p-P38, T-P38, p-JNK, T-JNK and GAPDH. Following incubation with secondary antibodies, the protein bands were detected using an enhanced chemiluminescence reagent (Thermo Fisher Scientific, Rockford, IL, USA).
Statistic analysis
All statistical analyses were performed using the Statistical Package for Social Sciences (SPSS) version 20.0 software. The data was downloaded from GEO dataset and the differences were analyzed. TTK expression was compared with t-test. Probability values <0.05 were considered statistically significant.
Results
Analysis of TTK expression in GEO dataset
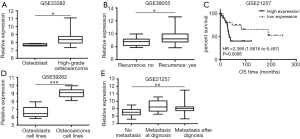
GEO dataset GSE33382, GSE39262, GSE39055, GSE21257 was analyzed and we compared the relative expression of TTK gene. GSE33382 and GSE39262 showed that TTK expression in osteoblast tissue was lower than that in osteosarcoma tissue (Figure 1A, P<0.05), as well as in their cell lines (Figure 1B, P<0.05). GSE39055 showed that TTK expression in patients with recurrence was higher than without (Figure 1C, P<0.05). GSE21257 showed that the higher expression of TTK was related to metastasis (Figure 1D, P<0.05) and short survival period (Figure 1E, P<0.05).
Analysis of TTK expression in TMAs
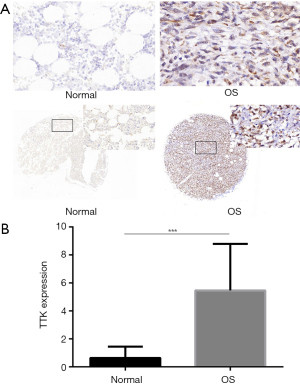
In order to investigate the expression level of TTK in clinic samples of OS, we used two TMAs to have a subset analysis. BO244d was consisted of 11 bone tissues and 1 OS tissue, and OS804b 40 OS tissues. The CES scores of every sample of the TMAs were measurement. We chose the images of TTK expression from the TMAs, representative images of TTK expression were shown in Figure 2A. TTK expression in OS was significantly higher than normal bone tissue (Figure 2B, P<0.05). The result was also consistent with what we analyzed from the GEO dataset.
TTK suppressing in MG-63 and U2-OS cells
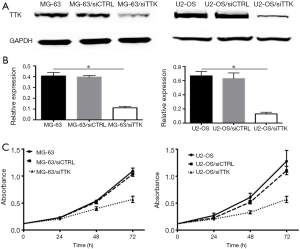
MG-63 and U2-OS cells stably transfected with TTK siRNA (named as MG-63/siTTK and U2-OS/siTTK). Control MG-63 and U2-OS cells were transfected with nonsense siRNA (named as MG-63/siCTRL and U2-OS/siCTRL). Western blot (Figure 3A,B) revealed that the expression of TTK protein was significantly reduced in MG-63/siTTK and U2-OS/siTTK cells relative to control cells. When TTK was suppressed, the proliferation of MG-63 cells inhibited in a time-dependent manner compared with the control cells (Figure 3C).
Analysis of the results of transwell chamber assay and flow cytometry
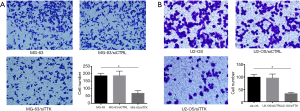
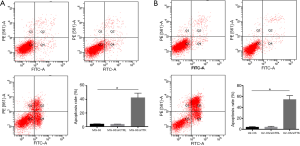
To further study the effect of TTK in MG-63 and U2-OS cells, the migration and apoptosis of cells was evaluated. The results showed that in both MG-63/siTTK and U2-OS/siTTK cells the migration was significantly reduced than in MG-63 (Figure 4A, P<0.05) and U2-OS cells (Figure 4B, P<0.05). The proportion of apoptotic cells of MG-63/siTTK and U2-OS/siTTK cells was significantly higher than that of MG-63 (Figure 5A, P<0.05) and U2-OS cells (Figure 5B, P<0.05).
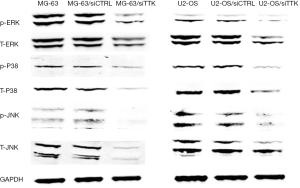
The expression of MAPK signal pathway related proteins
In order to explore the pathway behind, using Western blot we detected the expression of MAPK signal pathway related proteins, including p-ERK, T-ERK, p-P38, T-P38, p-JNK, T-JNK. The results revealed that after TTK suppressing, the expression of these genes were all decreased significantly (Figure 6).
Discussion
OS has been studied for many years, but still a highly malignant bone tumor that affect children and young adolescents with poor clinical outcomes (12). Recently it has been reported that the 5-year survival rate of OS is approximately 70% owe to the radical surgery and chemotherapy (2,13), but it remains unsatisfactory particularly of the metastasis which usually occur in the lung (12,14-16). About 40–50% of OS occur pulmonary metastasis and the 5-year survival rate is just 28% in such patients (17,18). Because of this, the study was designed to investigate the role of TTK in OS and the goal was to clarify whether TTK could offer additional help in the early diagnosis of OS or even be molecular targeted therapy.
Several studies have revealed that TTK was related to various kinds of human malignant tumors and always showed higher gene expression in tumor tissue than normal tissue, which was correspond with the results we observed in the present study (19). However there has few studies about the relation between OS and TTK which was more be investigated on hepatocellular carcinoma, triple-negative breast cancer or gastric/colorectal cancers (7,8,10,20).
Kaistha et al. (11) investigated the role of TTK in pancreatic cancer cells and reported that TTK function was vital for cell proliferation which was significantly attenuated after TTK knockdown. From it we suspect that high TTK expression could also promote the tumorigenesis of OS.
By using GEO dataset we have analyzed the TTK expression in sarcoma, it showed that TTK expression in sarcoma was significantly higher than normal tissue. And then the result of IHC had verified it. In order to explore the internal mechanism, we conducted cell experiments. The results revealed that in TTK suppressing cells the proliferation and migration were decreased with apoptosis increased. The mechanism may be related to MAPK signal pathway.
In a study about TTK gene in gastric/colorectal cancers, authors analyzed the mutation of TTK and revealed that frameshift mutations of TTK contribute to occurrence and development of tumors with high microsatellite instability (20). The mechanisms may also exist in OS and promote the tumorigenesis.
In addition, a few studies clarified that TTK could be a new approach of molecular targeting treatment in some other tumors, such as liver carcinoma, triple-negative breast cancer and so on. In a report about liver cancer, soft agar colony formation, cell growth and resistance to sorafenib were restrained following TTK knockdown(10). And in the present study, it had been verified by the results of cell experiment.
Furthermore, Maia et al. (9) used a well-defined mouse model represented human triple negative breast cancer and revealed that, the combination of inhibition of the spindle assembly checkpoint kinase TTK with a therapeutic dose of docetaxel could significantly slow tumor growth and increase survival rate. The results of an experiment about orally available Mps1 (TTK) kinase inhibitors also verified the anti-tumor effect (21). However, the internal mechanism should further be investigated in OS.
In a word, the present study revealed that TTK expression in OS was significantly higher than in normal tissue, it may offer additional help for the early diagnosis or even therapeutic target of OS. And the applicability and mechanisms need further investigation.
Acknowledgments
Funding: None.
Footnote
Conflicts of Interest: All authors have completed the ICMJE uniform disclosure form (available at http://dx.doi.org/10.21037/tcr.2017.03.73). The authors have no conflicts of interest to declare.
Ethical Statement: The authors are accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved. The study was conducted in accordance with the Declaration of Helsinki (as revised in 2013). Institutional ethical approval and informed consent were waived because what we used were two independent TMAs (Alenabio Biotechnology Co., LTD, Xi’an, China).
Open Access Statement: This is an Open Access article distributed in accordance with the Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License (CC BY-NC-ND 4.0), which permits the non-commercial replication and distribution of the article with the strict proviso that no changes or edits are made and the original work is properly cited (including links to both the formal publication through the relevant DOI and the license). See: https://creativecommons.org/licenses/by-nc-nd/4.0/.
References
- Li X, Xu R, Liu H, et al. CUL4A expression in pediatric osteosarcoma tissues and its effect on cell growth in osteosarcoma cells. Tumour Biol 2016;37:8139-44. [Crossref] [PubMed]
- Cheng HL, Hsieh MJ, Yang JS, et al. Nobiletin inhibits human osteosarcoma cells metastasis by blocking ERK and JNK-mediated MMPs expression. Oncotarget 2016;7:35208-23. [PubMed]
- Uzan VR. High Expression of HULC Is Associated with Poor Prognosis in Osteosarcoma Patients. PLoS One 2016;11:e0156774 [Crossref] [PubMed]
- Geller DS, Gorlick R. Osteosarcoma: a review of diagnosis, management, and treatment strategies. Clin Adv Hematol Oncol 2010;8:705-18. [PubMed]
- Lee DH, Qi J, Bradner JE, et al. Synergistic effect of JQ1 and rapamycin for treatment of human osteosarcoma. Int J Cancer 2015;136:2055-64. [Crossref] [PubMed]
- Zhang J, Wang X, Wu W, et al. Expression of the Nrf2 and Keap1 proteins and their clinical significance in osteosarcoma. Biochem Biophys Res Commun 2016;473:42-6. [Crossref] [PubMed]
- Liu X, Liao W, Yuan Q, et al. TTK activates Akt and promotes proliferation and migration of hepatocellular carcinoma cells. Oncotarget 2015;6:34309-20. [PubMed]
- Maire V, Baldeyron C, Richardson M, et al. TTK/hMPS1 is an attractive therapeutic target for triple-negative breast cancer. PLoS One 2013;8:e63712 [Crossref] [PubMed]
- Maia AR, de Man J, Boon U, et al. Inhibition of the spindle assembly checkpoint kinase TTK enhances the efficacy of docetaxel in a triple-negative breast cancer model. Ann Oncol 2015;26:2180-92. [Crossref] [PubMed]
- Liang XD, Dai YC, Li ZY, et al. Expression and function analysis of mitotic checkpoint genes identifies TTK as a potential therapeutic target for human hepatocellular carcinoma. PLoS One 2014;9:e97739 [Crossref] [PubMed]
- Kaistha BP, Honstein T, Müller V, et al. Key role of dual specificity kinase TTK in proliferation and survival of pancreatic cancer cells. Br J Cancer 2014;111:1780-7. [Crossref] [PubMed]
- Lucero CM, Vega OA, Osorio MM, et al. The cancer-related transcription factor Runx2 modulates cell proliferation in human osteosarcoma cell lines. J Cell Physiol 2013;228:714-23. [Crossref] [PubMed]
- Shu J, Li L, Sarver AE, et al. Imprinting defects at human 14q32 locus alters gene expression and is associated with the pathobiology of osteosarcoma. Oncotarget 2016;7:21298-314. [PubMed]
- Luo Z, Li D, Luo X, et al. Decreased Expression of miR-548c-3p in Osteosarcoma Contributes to Cell Proliferation Via Targeting ITGAV. Cancer Biother Radiopharm 2016;31:153-8. [Crossref] [PubMed]
- Tetreault MP, Yang Y, Katz JP. Krüppel-like factors in cancer. Nat Rev Cancer 2013;13:701-13. [Crossref] [PubMed]
- Isakoff MS, Bielack SS, Meltzer P, et al. Osteosarcoma: Current Treatment and a Collaborative Pathway to Success. J Clin Oncol 2015;33:3029-35. [Crossref] [PubMed]
- Hirahata M, Osaki M, Kanda Y, et al. PAI-1, a target gene of miR-143, regulates invasion and metastasis by upregulating MMP-13 expression of human osteosarcoma. Cancer Med 2016;5:892-902. [Crossref] [PubMed]
- Kempf-Bielack B, Bielack SS, Jürgens H, et al. Osteosarcoma relapse after combined modality therapy: an analysis of unselected patients in the Cooperative Osteosarcoma Study Group (COSS). J Clin Oncol 2005;23:559-68. [Crossref] [PubMed]
- Liu Y, Lang Y, Patel NK, et al. The Discovery of Orally Bioavailable Tyrosine Threonine Kinase (TTK) Inhibitors: 3-(4-(heterocyclyl)phenyl)-1H-indazole-5-carboxamides as Anticancer Agents. J Med Chem 2015;58:3366-92. [Crossref] [PubMed]
- Ahn CH, Kim YR, Kim SS, et al. Mutational analysis of TTK gene in gastric and colorectal cancers with microsatellite instability. Cancer Res Treat 2009;41:224-8. [Crossref] [PubMed]
- Kusakabe K, Ide N, Daigo Y, et al. Discovery of imidazo[1,2-b]pyridazine derivatives: selective and orally available Mps1 (TTK) kinase inhibitors exhibiting remarkable antiproliferative activity. J Med Chem 2015;58:1760-75. [Crossref] [PubMed]


