Defining the individual internal gross tumor volume of hepatocellular carcinoma using 4DCT and T2-weighted MRI images by deformable registration
Introduction
As our understanding of science technology and liver cancer has developed, the management of hepatocellular carcinoma (HCC) with radiotherapy has been well established over the last ten years. An estimated 65% of patients with primary HCC require radiotherapy treatment over the course of the disease. However, the clinical curative effect of radiotherapy in HCC is limited by normal liver toxicity. Precise radiotherapy greatly improves the dose received by the tumor target, while at the same time reducing radiation exposure to the normal liver. Two main variables are involved in this precise radiotherapy treatment of HCC, the respiratory movement and delineation of target area.
The movement of the liver associated with respiration creates challenges for scanning and treatment protocols. The liver is the largest organ affected by respiratory movement and has been reported to move in the range of 0.5–4.1 cm cranial to caudal (1). A CT scan acquired with the patient breathing freely will less accurately display the size and location of the target and surrounding normal liver tissue than one in breath hold (2). Breath motion management methods for precision radiotherapy of HCC currently include active breathing control (ABC), abdominal compression and four-dimensional computed tomography (4DCT). 4DCT sorts the volumetric CT images according to breath cycle, which are recorded and segmented applying a respiratory position management (RPM) system. Use of transarterial chemoembolization (TACE) prior to radiotherapy may assist in the location and treatment of smaller liver cancer lesions and help to identify the tumor target area. However, in tumors not treated with TACE, the delineation of the target relies on CT and MRI. MRI has been reported to be superior to CT in detection, follow up and staging of lesions due to its higher sensitivity and specificity (3). HCC demonstrates high signal in T2-weighted MRI, and strong signal differences with normal liver tissue. Indeed, using MRI to evaluate liver cancer following radiotherapy, Mahmoud et al. (4) reported that 77% of recurrent lesions elicit a high signal, indicating that T2-weighted images are superior to the non-enhanced T1-weighted images in the detection of HCC. As such, in the present study we used T2-weighted MRI to achieve deformable registration (DR) with 4DCT, in order to delineate the target in HCC for radiotherapy treatment.
In the present study, we investigate a novel method of defining the individual internal gross tumor volume (IGTV) of HCC using 4DCT and T2-weighted MRI images using DR. The delineation of IGTV using contrast-enhanced 4DCT scanning remains in the study due to the difficulty associated with capturing the timing of contrast agent injection during the extended 4DCT image acquisition process.
Methods
Patient information
Ten patients (median age 50, range, 38–60 years) with HCC as confirmed by pathology or imaging, were recruited for this study from the Shandong Cancer Hospital between September 2015 and January 2016. The study was approved by the Research Ethics Board of the Shandong Cancer Hospital, with all patients providing written informed consent at enrollment. None of the patients included in the study received TACE. Prior to receiving initial radiotherapy, the patients’ Karnofsky performance status was greater than 80. The patient characteristics are summarized in Table 1.
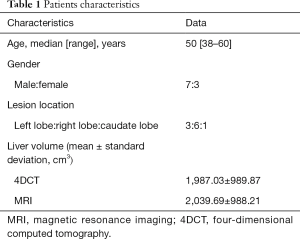
Full table
Imaging study
Image acquisition of 4DCT scans was performed using a Brilliance Big Bore CT scanner (Philips Medical Systems, Highland Heights, OH, USA) in free breathing. All patients were positioned in a supine position in an immobilization device during the scans. The CT scans were equipped with the RPM system (Varian Medical System, Palo Alto, CA, USA), using infrared cameras to record the trajectory of two fluorescent marker points placed near the Xiphoid process module and to record the respiratory signal. 4DCT images were sorted into ten series of CT images (CT00–CT90) according to the respiratory phase, with CT00 defined as end inspiration and CT50 end expiration. For all images, the following parameters were used: 120 KV, 300 mAs, 3 mm slice thickness, reconstruction matrix of 512×512 and a field of view (FOV) of 450–500 mm.
All MRI acquisitions were performed on a Philips 3.0-T Ahieva MR system (Philips Medical System, Best, Netherlands). The T2-weighted MRI was completed in breath hold using a respiratory-triggered fast relaxation fast spin echo sequence. The imaging parameters for T2-weighted MRI were as follows: TR/TE: 12,857 ms/64.196 ms; FOV: 350×227 mm2; 5-mm slice thickness. In the present study, MRI scanning was conducted on the same day as 4DCT, 5–6 hours later.
All images, including 4DCT and MRI, were exported into MIM version 6.5.6, (MIM Software Inc., Cleveland, OH, USA) for analysis.
Target delineation
GTVs and liver contours were delineated on all images and IGTVs acquired by merging ten GTVs at all phases of 4DCT images. The GTVs on T2-weighted MRI images were deformably registered to each phase of 4DCT images and labeled –DR. The IGTVDR was obtained by merging the ten GTVsDR after DR. The volume difference among the GTVs, and between IGTV and IGTVDR were compared. All GTVs and liver contours were delineated by the same radiation oncologist, using the same window width and level on 4DCT images (width: 400 HU, level: 40 HU).
T2-weighted MRI and 4DCT DR and accuracy measurements
The contours on T2-weighted MRI were propagated to all 4DCT images using deformable image registration provided by our treatment planning system (MIM). The deformation image registration technique was based on the Gauss mixture model. In this study, T2-weighted MRI images were selected as the target image and ten phases of 4DCT images used as the reference images. The target image was registered to different 4DCT phase images on the basis of application of automatic deformation, and Reg Refine used to lock the local interest area. Tracking liver volume as the organ of interest was used to assess the registration effect.
To check the accuracy of automatic DR, two methods were utilized. Liver contours were delineated on all images and liver volume calculated on every phase image of 4DCT(V-LIVERCT00–V-LIVERCT90)and on T2-weighted MRI images (V-LIVERT2w MRI). P-Liver, defined as (V-LIVERT2w MRI/V-LIVERCT)*100, was used to evaluate the accuracy of DR. Additionally, the maximum displacement on three dimensional directions of two specific anatomical landmarks, the bifurcation point in the portal vein and the celiac trunk, was also contoured for both 4DCT and MRI to confirm accuracy of DR. The left to right distance was labeled X, the cranial to caudal distance Y and the anterior to posterior distance labeled Z. Then, the Reg Reveal and Reg Refine were used to minimize the registration error manually.
Statistical analysis
The data were analyzed using the SPSS 17.0 software package (IBM, Chicago, IL, USA). The difference of GTVs and IGTVs before DR and after DR was determined by a paired t-test. Data are presented with as the mean () ± standard deviation(s). A value of P<0.05 was considered statistically significant.
Results
T2-weighted MRI and 4DCT DR and accuracy
The mean liver volume as determined by 4DCT and T2-weighted MRI was 1,987.03±989.87 cm3 (1,065.86–4,144.06 cm3) and 2,039.69±988.21 cm3 (1,321.99–4,238.36 cm3), respectively. The P-LIVER was 115.4%±13.8%. Automatic deformation registration was used. The maximal observed displacements of the bifurcation point in the portal vein were 3.6±1.2 mm along the X-axes, 5.0±1.4 mm along the Y-axes, 4.2±0.9 mm along the Z-axes, whereas the celiac trunk demonstrated maximal displacements of 2.8±2.3 mm along the X-axes, 3.0±1.7 mm on the Y-axes, and 2.5±3.4 mm on the Z-axes. After using Reg Refine registration tool, the maximal observed displacements of the bifurcation point in the portal vein were 0.3±0.8 mm along the X-axes, 0.8±1.8 mm along the Y-axes, 0.5±1.5 mm along the Z-axes, whereas the celiac trunk demonstrated maximal displacements of 0.1±1.0 mm along the X-axes, 0.7±1.2 mm on the Y-axes, 0.6±2.0 mm on the Z-axes (as show in Table 2). The Reg Reveal and Reg Refine were used to minimize the registration error manually within 1 mm.

Full table
One patient is described as an example. Automatic deformation registration was used. When automatic deformation registration was used, the maximal observed displacements of the bifurcation point in the portal vein was 2.5, 1.0 and 2.0 mm along X-axes, Y-axes and Z-axes. The maximal observed displacements of the celiac trunk were 3.0, 1.8 and 1.0 mm along X-axes, Y-axes, Z-axes. After using the Reg Refine registration tool, the maximal observed displacements of the bifurcation point in the portal vein was 0.5, 0.5 and 1.0 mm along the X-axes, Y-axes and Z-axes. The maximal observed displacements of the celiac trunk were 0.5, 0.6 and 1.0 mm along X-axes, Y-axes, Z-axes (Figure 1).
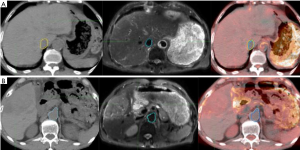
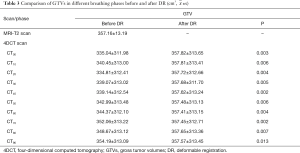
Comparison of GTVs before and after DR
GTVs obtained from 4DCT images were increased by an average of 4.23% (P<0.05) after DR, with GTVs following DR of each phase consistent with T2-weighted MRI image basic volume (Table 3, Figures 2,3). After DR all 4DCT image phases GTV were larger than DR before (except CT90, P<0.05).
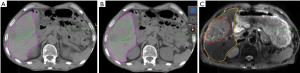
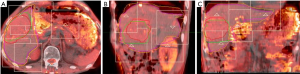
Comparison of IGTVs before and after DR
The IGTV before DR was 383.89±342.53 cm3, with a 9.67% increase observed following DR (421.02±382.13 cm3), indicating that a significant increase in IGTV volume occurred after DR (P=0.018) (as shown in Table 4).
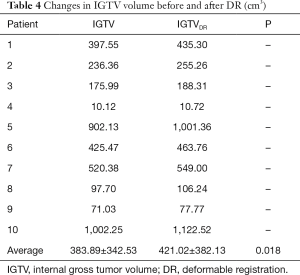
Full table
Discussion
The tumor target can move and deform significantly during respiration, which is a critical parameter for precision radiotherapy in order to accurately delineate the target lesions for treatment. To combat this, 4DCT is increasingly used in radiation oncology to account for the effect of breathing on organ and tumor position in the thorax and abdomen. Accordingly, the use of 4DCT may minimize the PTV, resulting in improvements in organs at risk sparing due to more precise target localization (5).
CT imaging is used in radiation dose calculations as it provides electronic density information, but it is unable to precisely identify the boundary of HCC lesions. However, this defect can be offset by use of T2-weighted MRI. Nonetheless, CT and MRI imaging are associated with a number of limitations. T2-weighted MRI is performed in deep inspiration, and thus differences in liver shape may occur. Additionally, the MRI aperture is small and does not allow space for a position-fixing device, thus the MRI scanning position is not completely consistent with CT. Furthermore, MRI and CT scan are generated at different times, potentially resulting in differences in location and filling degree of internal organs. Given these limitations, this study investigated DR technology to combine 4DCT and T2-weighted MRI images for use in radiotherapy planning in the treatment of HCC.
Deformable image registration can achieve the corresponding points to space position and anatomical position exactly, or at least all the points of diagnostic significance in order to match between two given images through a space algorithm (6). It has been reported that DR technology improves the accuracy of the target sketch and cumulative dose assessment (7-10). This study acquired DR on different sequence images from 4DCT, and made targets on the T2-weighted MRI image deformation to each phase image of 4DCT. Use of 4DCT provided target movement information, and improved the precision of the target sketch. Our results indicate that the delineation of GTVs size and scope based on T2-weighted MRI was greater than CT scan images. Although DR is associated with increased processing time, relative to the artificial sketch based on MRI images with each phase image of 4DCT, it may significantly increase the precision outline of target.
DR was performed using the VoxAlign algorithm incorporated with MIM. The VoxAlign algorithm is a constrained, intensity-based, free form DIR algorithm. Reg Reveal and Reg Refine were implemented recently in MIM as tools that allow the user to view and refine DR. Pukala et al. (11) tested the algorithm between five commercial deformable image registration algorithms (MIM, Velocity, RayStation, Pinnacle, and Eclipse) for head and neck cancer patients and determined that the target registration error (TRE) for an individual DR using the MIM algorithm under similar conditions was less than 1 mm for these structures. Furthermore, Annemarie et al. (12) used this algorithm to assess liver tumor motion. Thus, these reports support the accuracy and reliability of organ registration using MIM.
To assess the accuracy of image registration, we adopted three indicators: overlap degree of the liver, the displacement of the bifurcation point in the portal vein and celiac axis. Our results demonstrated that maximal displacement occurred in the cranial to caudal direction, in keeping with previously published reports (12,13). We used DR technology to analyze the application of 4DCT and MRI images in the delineation of the liver target. Our results revealed that the delineation of GTV size and range based on MRI is significantly greater than 4DCT images. GTV increased by an average of 4.23% (P<0.05) and IGTV volume increased by an average of 9.67% (P<0.05), suggesting that MRI may be a good supplement to CT image information. This not only improves the accuracy of HCC target identification using 4DCT but also may be beneficial in cases where there is a lot of exit noise on the original 4DCT image and failure to identify tumor boundaries.
Our study is limited by a relatively small number of patients. In addition, MRI scans were performed in the absence of breath control devices under an independent state of deep inspiration. Further studies should address these limitations in MRI-4DCT image DR.
Conclusions
Here, we demonstrated the IGTVs obtained from combination of MRI and 4DCT images may better determine the scope and the trajectory of the tumor target, and thus, improve target delineation accuracy for use in radiation treatment of HCC.
Acknowledgments
Funding: This work is supported by Natural Science Foundation of China (No.81272699, No.81472811), Natural Science Foundation of Shandong Province (ZR2016HM41), Projects of Medical and Health Technology Development Program in Shandong Province (2016WS0553), Project funded by China Postdoctoral Science Foundation (2017M610430).
Footnote
Conflicts of Interest: All authors have completed the ICMJE uniform disclosure form (available at http://dx.doi.org/10.21037/tcr.2018.01.20). The authors have no conflicts of interest to declare.
Ethical Statement: The authors are accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved. The study was conducted in accordance with the Declaration of Helsinki (as revised in 2013). The study was approved by the Research Ethics Board of the Shandong Cancer Hospital, with all patients providing written informed consent at enrollment.
Open Access Statement: This is an Open Access article distributed in accordance with the Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License (CC BY-NC-ND 4.0), which permits the non-commercial replication and distribution of the article with the strict proviso that no changes or edits are made and the original work is properly cited (including links to both the formal publication through the relevant DOI and the license). See: https://creativecommons.org/licenses/by-nc-nd/4.0/.
References
- Eccles C, Brock KK, Bissonnette JP, et al. Reproducibility of liver position using active breathing coordinator for liver cancer radiotherapy. Int J Radiat Oncol Biol Phys 2006;64:751-9. [Crossref] [PubMed]
- Wang L, Hayes S, Paskalev K, et al. Dosimetric comparison of stereotactic body radiotherapy using 4D CT and multiphase CT images for treatment planning of lung cancer: evaluation of the impact on daily dose coverage. Radiother Oncol 2009;91:314-24. [Crossref] [PubMed]
- Kierans AS, Elazzazi M, Braga L, et al. Thermoablative treatments for malignant liver lesions: 10-year experience of MRI appearances of treatment response. AJR Am J Roentgenol 2010;194:523-9. [Crossref] [PubMed]
- Mahmoud BEMH, Elkholy S, Nabeel M, et al. Role of MRI in the assessment of treatment response after radiofrequency and microwave ablation therapy for hepatocellular carcinoma. The Egyptian Journal of Radiology and Nuclear Medicine 2016;47:377-85. [Crossref]
- Korreman S, Persson G, Nygaard D, et al. Respiration-correlated image guidance is the most important radiotherapy motion management strategy for most lung cancer patients. Int J Radiat Oncol Biol Phys 2012;83:1338-43. [Crossref] [PubMed]
- Li W, Huang Y, Tian X, et al. The experimental research on the frameless registration based on the digital subtraction angiography. Sheng Wu Yi Xue Gong Cheng Xue Za Zhi 2007;24:23-5, 44.
- Brock KK, Lee M, Eccles CL, et al. Deformable registration and dose accumulation to investigate marginal liver cancer recurrences. Int J Radiat Oncol Biol Phys 2008;72:S538. [Crossref]
- Wijesooriya K, Weiss E, Dong L, et al. Quantifying the accuracy of automated structure segmentation in 4DCT images using a deformable image registration algorithm. Med Phys 2008;35:1251-60. [Crossref] [PubMed]
- Jung SH, Yoon SM, Park SH, et al. Four-dimensional dose evaluation using deformable image registration in radiotherapy for liver cancer. Med Phys 2013;40:011706 [Crossref] [PubMed]
- Velec M, Moseley JL, Eccles CL, et al. Effect of breathing motion on radiotherapy dose accumulation in the abdomen using deformable registration. Int J Radiat Oncol Biol Phys 2011;80:265-72. [Crossref] [PubMed]
- Pukala J, Johnson PB, Shah AP, et al. Benchmarking of five commercial deformable image registration algorithms for head and neck patients. J Appl Clin Med Phys 2016;17:25-40. [Crossref] [PubMed]
- Annemarie TF, Smith A, Yin Lingshu, et al. Comparative assessment of liver tumor motion using cine-magnetic resonance imaging versus 4-dimensional computed tomography. Int J Radiat Oncol Biol Phys 2015;91:1034-40. [Crossref] [PubMed]
- JI Yu, Kim JS, Park HC, et al. Evaluation of anatomical landmark position differences between respiration-gated MRI and four-dimensional CT for radiation therapy in patients with hepatocellular carcinoma. Br J Radiol 2013;86:20120221 [Crossref] [PubMed]



