
The association of interleukin-6 gene polymorphism and risk of colorectal cancer in Chinese patients
Introduction
Colorectal cancer (CRC) is the third most common cancer worldwide, and a number of 530,000 patients died of CRC each year (1). There are numerous factors influencing the development of CRC, including genetic, environmental, smoking, drinking and the aberrant inflammation of gastrointestinal tract (2). Of note, it is proved that tumor-associated inflammation significantly affected cancer development (3). Among the various inflammation-associated factors, the role of interleukin-6 (IL-6) in CRC tumorigenesis has been well-established.
IL-6 is a pleiotropic inflammatory cytokine which plays a vital role not only in immune and inflammatory response, but also the development of malignancy (4). Moreover, it is reported that the increased production of IL-6 is observed in tumor itself and the serum of CRC patients (5). In addition, IL-6 expression shows a close relation to tumor stage, size, metastasis and survival of CRC patients (6). And these results may result from its role in producing pro-inflammatory cytokines and modulating Th17 and Treg cells in CRC (7). The role of rs1800795, a well-studied IL-6 polymorphism, has been discussed in vast areas (8-21). Our team has conducted a meta-analysis to explore the relationship between IL-6 polymorphisms and CRC risk, and we discovered that IL-6 rs1800795 polymorphism significantly increased the risk of CRC in allele additive model in Europe [odds ratio (OR): 1.07, 95% confidence interval (CI): 1.01, 1.14] (22). However, few studies were carried out to explore the relationship between the IL-6 rs1800795 polymorphism and CRC risk in Chinese population. Hence, the aim of this study is to discover the association between one variant (rs1800795) in IL-6 gene and CRC risk in Chinese population.
Methods
Study population
This is a hospital-based case-control study conducted in Wuxi Traditional Chinese Hospital during October 2015 and July 2017. The inclusion criteria for CRC patients were as follows: positive results of colonoscopy and pathology for colon or rectum tumors. And the control samples should have no gastrointestinal disorders, and these control samples have been subjected to diagnostic colonoscopy and had eligible colonoscopy results with no malignant tumors, adenomatous polyps, or inflammatory ulcers. All subjects were self-reported born in China. The exclusion criteria were: a history of other cancer, HIV, HBV, HCV, or IV drug use, or a mental impairment. We also recorded the information of age, sex, disease stage, smoking, drinking and dietary habit from medical records and pathology reports.
All enrolled subjects were informed and gave written consent to participate in this study according to the Helsinki declaration, and this study protocol was also supported by the ethical committee of Wuxi Traditional Chinese Hospital.
Genotyping
We isolated genomic DNA from 5 mL of peripheral blood leukocytes using the standard phenol-chloroform method. Single nucleotide polymorphism (SNP) genotyping was conducted with predesigned TaqMan assays from Applied Biosystems by minor groove binder probes fluorescently labeled with FAM or VIC (Genesky Biotechnologies Inc., Shanghai, China). The probes and primers used for SNP genotyping are: IL-6 174 G_C (rs1800795), VIC-TCTTGCGATGCTAAA, FAM-TCTTGCCATGCTAAA, Forward primer: TGACGACCTAAGCTGCACTTTTC, Reverse primer: GGGCTGATTGGAAACCTTATTAAGA. Polymerase chain reactions were operated on a Tetrad DNA Engine PCR machine and read in an ABI PRISM 7900 Sequence detection system. In order to keep the quality in genotyping, we repeated the genotyping for three times until to explore the consistent results.
Statistical analysis
The statistical significance of age and body mass index (BMI) was calculated by the Student’s t-test. The χ2 test or Fisher’s exact test was used for univariate analyses of allele and genotype distribution between CRC patients and control subjects. The multivariate analyses of the association between SNPs and CRC risk were evaluated with the OR with 95% CIs by logistic regression, adjusting for age, gender, and BMI. When the minor allele homozygote counts are 14 or more, we calculated in three kinds of logistic regression models (dominant, additive and recessive). When the minor allele homozygote count is less than 14, we only examined in the dominant genetic model. Hardy-Weinberg equilibrium (HWE) was performed to exclude deviations in control subjects by HWE version 1.20 (Columbia University, New York, NY). In addition, the statistical power was measured with the STPLAN 4.3 software under a given sample size and significance level (α=0.05). The results were considered significant when the P values are less than 0.05. All statistical analyses were conducted with SPSS 17.0.
Results
Characteristics of the study groups
A total of 186 CRC patients and 200 control samples were enrolled in our study, and all samples in our study are all Han ethnic (Table 1). The distribution of the included SNP was in HWE for control subjects (P=0.16). In addition, rs1800795 had a power of 0.264 which is lower than 0.08. There are no significant differences in age (P=0.390), BMI (P=0.052), gender (P=0.465), smoking (P=0.210), drinking (P=0.561) or diet habit (P=0.377). The case group comprises of 115 colon cancer patients and 71 rectum cancer patients. Moreover, there are 33 patients in tumor Node Metastasis (TNM) stage I, 70 patients in TNM stage II, 71 patients in TNM stage III and 12 patients in TNM stage IV.
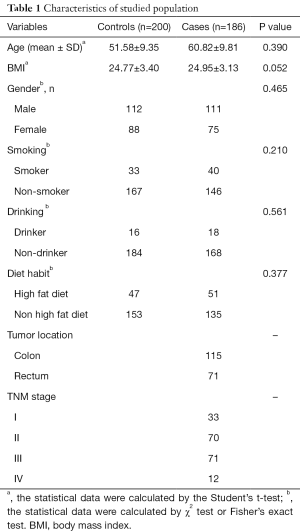
Full table
Univariate analysis
In the univariate analysis of the association between rs1800795 polymorphism and CRC risk, our results showed the significant allele distributions in female (OR: 1.741, 95% CI: 1.037, 2.924, P value: 0.035) and non-smokers (OR: 2.425, 95% CI: 1.626, 3.617, P value: <0.001). Moreover, the significant findings were observed in genotype distribution of non-smokers (P value: <0.001), drinkers (P value: 0.026) and the samples of non-high fat diet (P value: 0.006) (Table 2). In addition, no significant results were seen in the allele or genotype distribution based on CRC kind or TNM stage (Table 3).
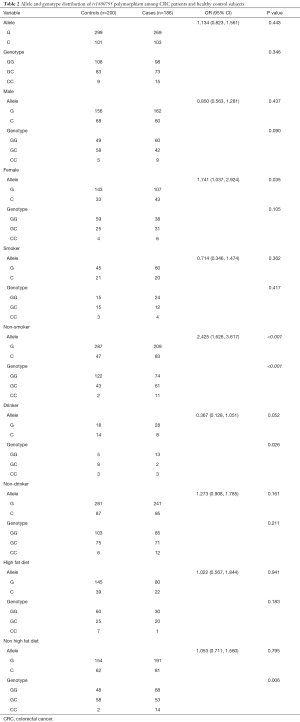
Full table
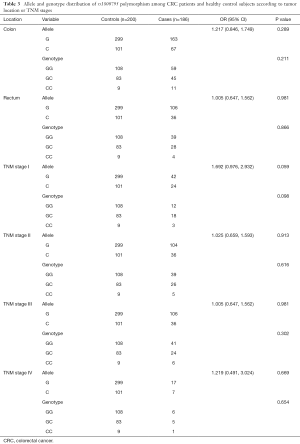
Full table
Multivariate analysis
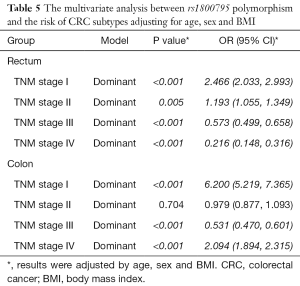
In the multivariate analysis of the association between rs1800795 polymorphism and CRC risk, we adjusted the results for age, sex and BMI. Finally, we did not find any significant association between rs1800795 polymorphism and CRC risk in whole people (OR: 0.966, 95% CI: 0.894, 1.043, P value: 0.374), colon cancer samples (OR: 0.941, 95% CI: 0.866, 1.024, P value: 0.158) or rectum cancer patient (OR: 0.955, 95% CI: 0.871, 1.048, P value: 0.336). Our studies showed significant association between rs1800795 polymorphism and CRC risk in TNM stage I (OR: 4.841, 95% CI: 4.220, 5.554, P value: <0.001) and TNM stage II (OR: 1.138, 95% CI: 1.034, 1.252, P value: 0.008) (Table 4). Moreover, we further explored the associations between SNP and CRC subtypes. Interesting, we found positive correlation between rs1800795 polymorphism and rectum risk in TNM stage I (OR: 2.466, 95% CI: 2.033, 2.993, P value: <0.001) and TNM stage II (OR: 1.193, 95% CI: 1.055, 1.349, P value: 0.005). While negative correlation was reflected in TNM stage III (OR: 0.573, 95% CI: 0.499, 0.658, P value: <0.001) and TNM stage IV (OR: 0.216, 95% CI: 0.148, 0.316, P value: <0.001). In addition, results have shown significant association between rs1800795 polymorphism and colon risk in TNM stage I (OR: 6.200, 95% CI: 5.219, 7.365, P value: <0.001), and TNM stage IV (OR: 2.094, 95% CI: 1.894, 2.315, P value: <0.001) (Table 5).
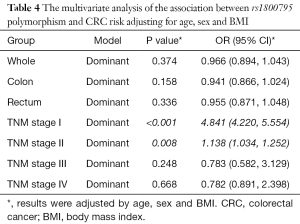
Full table

Full table
Discussion
The SNP rs1800795 is a well-studied SNP, which has been widely discussed in wide districts except for China. To our knowledge, this is the first study to explore the association between rs1800795 polymorphism and CRC risk. Our studies showed the C allele would significant increase the CRC risk in female and non-smokers. And the significantly associations existed in the genotype distribution of rs1800795 polymorphism and CRC risk in non-smokers, drinkers and the samples with non-high fat diet. Our studies showed significant positive relations between rs1800795 polymorphism and CRC risk in TNM stage I and TNM stage II.
IL-6 is an important element of immune and inflammatory system, and it is also a key growth factor for malignancy (4). A recent study reported that the level of IL-6 was significantly higher in CRC tissues compared with noncancerous tissues, furthermore, the level of IL-6 was linked to invasion depth and lymph node metastasis in CRC (23). The team of Hara M discovered that the CRC patients with high serum IL-6 levels would have poorer overall survival and progression-free survival in comparison to those with low serum IL-6 levels, and a Cox proportional hazards regression analysis indicated that the CRC patients with high serum IL-6 levels may have an independent risk factor for a poor outcome (24). In addition, a review survey by Knüpfer and Preiss reported a close association between IL-6 expression and tumor stage, size, metastasis and survival of patients with CRC (6). Beside the findings about the close association between IL-6 and CRC clinically, a number of studies also explored its potential concrete molecular mechanisms. The current findings indicated that IL-6 functions in recruitment of immune cells and modulating Th17 and Treg cells in CRC (7). An increasing number of studies indicated CRC cells secreting IL-6 via STAT3 and JAK phosphorylation to enhance the phagocytic capacity and migration of macrophages in the tumor microenvironment (25-30), and modulating IL-6 would be a promising method to treat CRC (31-35). All these above studies indicated a close relation between IL-6 and CRC.
Based on the marked association between IL-6 and CRC, many researchers turned to reveal the relation between IL-6 polymorphism and CRC risk. To date, several IL-6 SNPs have been explored their association and CRC risk (8-21). Of note, rs1800795 is the most well studies SNP. Rs1800795 is located in promoter region of the gene (-174 n) and it is responsible for binding of certain transcriptional factors affecting transcription, RNA elongation, splicing, or maturation (36-39). Therefore, mutations in rs1800795 may also influence the expression of IL-6. To date, rs1800795 has been discussed its role in Spain (8,21), USA (9,13,17,20), Denmark (10,12), Greece (11), Sweden (14), France (15), Czech (16), Croatia (18) and Canada (19). However, no studies have been explored in China. In this study, this is the first study to explore the association between rs1800795 polymorphism and CRC risk in Chinese. Our studies showed the C allele would significantly increase the CRC risk in female and non-smokers. And the significant associations existed in the genotype distribution of rs1800795 polymorphism and CRC risk in non-smokers, drinkers and the samples with non-high fat diet. These studies indicated the distributions of allele and genotype in specific groups are significantly different, which indicated screening the rs1800795 polymorphism in specific groups would be a promising method to predict the risk of CRC. In addition, our studies showed significant positive relations between rs1800795 polymorphism and CRC risk in TNM stage I and TNM stage II in multivariate analysis after adjusting for the results for age, sex and BMI, these results indicated that rs1800795 polymorphism may be a useful marker for CRC prognosis.
However, there are some limitations in our study. First, the size of enrolled samples is relatively small. Relatively small sample size is more likely to be lack of sufficient statistical power to verify the true positive findings if the result is negative. While when the result is positive, the outcome would be changed by chance due to the small sample size; then, further studies needed to be conducted to explore the role of rs1800795 mutations in CRC development and progression.
This study is the first report of rs1800795 polymorphisms in IL-6 gene among CRC patients from China. However, more large size of population needed to verify these results, and further studies needed to be conducted to explore its potential mechanism.
Acknowledgments
Funding: This work was supported by youth project of Wuxi city health and family planning commission (Q201619).
Footnote
Conflicts of Interest: All authors have completed the ICMJE uniform disclosure form (available at http://dx.doi.org/10.21037/tcr.2018.03.37). The authors have no conflicts of interest to declare.
Ethical Statement: The authors are accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved. All enrolled subjects were informed and gave written consent to participate in this study according to the Helsinki declaration (as revised in 2013), and this study protocol was also supported by the ethical committee of Wuxi Traditional Chinese Hospital (approval ID: 2016122003).
Open Access Statement: This is an Open Access article distributed in accordance with the Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License (CC BY-NC-ND 4.0), which permits the non-commercial replication and distribution of the article with the strict proviso that no changes or edits are made and the original work is properly cited (including links to both the formal publication through the relevant DOI and the license). See: https://creativecommons.org/licenses/by-nc-nd/4.0/.
References
- Gao S, Tibiche C, Zou J, et al. Identification and Construction of Combinatory Cancer Hallmark-Based Gene Signature Sets to Predict Recurrence and Chemotherapy Benefit in Stage II Colorectal Cancer. JAMA Oncol 2016;2:37-45. [Crossref] [PubMed]
- Hacker UT, Wolf J, Wendtner CM. Recent results of research on cancer of the colon, gastric cancer, sarcoma and bronchial carcinoma. Dtsch Med Wochenschr 2011;136:179-81. [Crossref] [PubMed]
- Wang SW, Hu J, Guo QH, et al. AZD1480, a JAK inhibitor, inhibits cell growth and survival of colorectal cancer via modulating the JAK2/STAT3 signaling pathway. Oncol Rep 2014;32:1991-8. [Crossref] [PubMed]
- Candido J, Hagemann T. Cancer-related inflammation. J Clin Immunol 2013;33:S79-84. [Crossref] [PubMed]
- Chung YC, Chang YF. Serum interleukin-6 levels reflect the disease status of colorectal cancer. J Surg Oncol 2003;83:222-6. [Crossref] [PubMed]
- Knüpfer H, Preiss R. Serum interleukin-6 levels in colorectal cancer patients--a summary of published results. Int J Colorectal Dis 2010;25:135-40. [Crossref] [PubMed]
- Jones SA, Richards PJ, Scheller J, et al. IL-6 transsignaling: the in vivo consequences. J Interferon Cytokine Res 2005;25:241-53. [Crossref] [PubMed]
- Landi S, Moreno V, Gioia-Patricola L, et al. Association of common polymorphisms in inflammatory genes interleukin (IL)6, IL8, tumor necrosis factor alpha, NFKB1, and peroxisome proliferator-activated receptor gamma with colorectal cancer. Cancer Res 2003;63:3560-6. [PubMed]
- Gunter MJ, Canzian F, Landi S, et al. Inflammation-related gene polymorphisms and colorectal adenoma. Cancer Epidemiol Biomarkers Prev 2006;15:1126-31. [Crossref] [PubMed]
- Gaustadnes M, Orntoft TF, Jensen JL, et al. Validation of the use of DNA pools and primer extension in association studies of sporadic colorectal cancer for selection of candidate SNPs. Hum Mutat 2006;27:187-94. [Crossref] [PubMed]
- Theodoropoulos G, Papaconstantinou I, Felekouras E, et al. Relation between common polymorphisms in genes related to inflammatory response and colorectal cancer. World J Gastroenterol 2006;12:5037-43. [Crossref] [PubMed]
- Vogel U, Christensen J, Dybdahl M, et al. Prospective study of interaction between alcohol, NSAID use and polymorphisms in genes involved in the inflammatory response in relation to risk of colorectal cancer. Mutat Res 2007;624:88-100. [Crossref] [PubMed]
- Slattery ML, Wolff RK, Herrick JS, et al. IL6 genotypes and colon and rectal cancer. Cancer Causes Control 2007;18:1095-105. [Crossref] [PubMed]
- Wilkening S, Tavelin B, Canzian F, et al. Interleukin promoter polymorphisms and prognosis in colorectal cancer. Carcinogenesis 2008;29:1202-6. [Crossref] [PubMed]
- Kury S, Buecher B, Robiou-du-Pont S, et al. Low-penetrance alleles predisposing to sporadic colorectal cancers: a French case-controlled genetic association study. BMC Cancer 2008;8:326. [Crossref] [PubMed]
- Vasku A, Vokurka J, Bienertova-Vasku J. Obesity-related genes variability in Czech patients with sporadic colorectal cancer: preliminary results. Int J Colorectal Dis 2009;24:289-94. [Crossref] [PubMed]
- Tsilidis KK, Helzlsouer KJ, Smith MW, et al. Association of common polymorphisms in IL10, and in other genes related to inflammatory response and obesity with colorectal cancer. Cancer Causes Control 2009;20:1739-51. [Crossref] [PubMed]
- Cacev T, Jokic M, Loncar B, et al. Interleukin-6-174 G/C polymorphism is not associated with IL-6 expression and susceptibility to sporadic colon cancer. DNA Cell Biol 2010;29:177-82. [Crossref] [PubMed]
- Hawken SJ, Greenwood CM, Hudson TJ, et al. The utility and predictive value of combinations of low penetrance genes for screening and risk prediction of colorectal cancer. Hum Genet 2010;128:89-101. [Crossref] [PubMed]
- Ognjanovic S, Yamamoto J, Saltzman B, et al. Serum CRP and IL-6, genetic variants and risk of colorectal adenoma in a multiethnic population. Cancer Causes Control 2010;21:1131-8. [Crossref] [PubMed]
- Abuli A, Fernandez-Rozadilla C, Alonso-Espinaco V, et al. Case-control study for colorectal cancer genetic susceptibility in EPICOLON: previously identified variants and mucins. BMC Cancer 2011;11:339. [Crossref] [PubMed]
- Wang S, Zhang W. Genetic variants in IL-6/JAK/STAT3 pathway and the risk of CRC. Tumour Biol 2016;37:6561-9. [Crossref] [PubMed]
- Zeng J, Tang ZH, Liu S, et al. Clinicopathological significance of overexpression of interleukin-6 in colorectal cancer. World J Gastroenterol 2017;23:1780-6. [Crossref] [PubMed]
- Hara M, Nagasaki T, Shiga K, et al. High serum levels of interleukin-6 in patients with advanced or metastatic colorectal cancer: the effect on the outcome and the response to chemotherapy plus bevacizumab. Surg Today 2017;47:483-9. [Crossref] [PubMed]
- Yeh KY, Wu TH, Wu TL. Colorectal cancer cell-derived interleukin-6 enhances the phagocytic capacity and migration of THP-1 cells. Cytokine 2016;79:82-9. [Crossref] [PubMed]
- Holmer R, Watzig GH, Tiwari S, et al. Interleukin-6 trans-signaling increases the expression of carcinoembryonic antigen-related cell adhesion molecules 5 and 6 in colorectal cancer cells. BMC Cancer 2015;15:975. [Crossref] [PubMed]
- Rasanen K, Lehtinen E, Nokelainen K, et al. Interleukin-6 increases expression of serine protease inhibitor Kazal type 1 through STAT3 in colorectal adenocarcinoma. Mol Carcinog 2016;55:2010-23. [Crossref] [PubMed]
- Lin J, Li Q, Chen H, et al. Hedyotis diffusa Willd. extract suppresses proliferation and induces apoptosis via IL-6-inducible STAT3 pathway inactivation in human colorectal cancer cells. Oncol Lett 2015;9:1962-70. [Crossref] [PubMed]
- Patel SA, Bhambra U, Charalambous MP, et al. Interleukin-6 mediated upregulation of CYP1B1 and CYP2E1 in colorectal cancer involves DNA methylation, miR27b and STAT3. Br J Cancer 2014;111:2287-96. [Crossref] [PubMed]
- De Simone V, Franze E, Ronchetti G, et al. Th17-type cytokines, IL-6 and TNF-alpha synergistically activate STAT3 and NF-kB to promote colorectal cancer cell growth. Oncogene 2015;34:3493-503. [Crossref] [PubMed]
- Park KI, Kim DG, Lee BH, et al. Fermented Herbal Formulas KIOM-MA128 Ameliorate IL-6-Induced Intestinal Barrier Dysfunction in Colon Cancer Cell Line. Mediators Inflamm 2016;2016:6189590 [PubMed]
- Han S, Jeong AJ, Yang H, et al. Ginsenoside 20(S)-Rh2 exerts anti-cancer activity through targeting IL-6-induced JAK2/STAT3 pathway in human colorectal cancer cells. J Ethnopharmacol 2016;194:83-90. [Crossref] [PubMed]
- Wang L, Zhao M, Guo C, et al. PDCD4 Deficiency Aggravated Colitis and Colitis-associated Colorectal Cancer Via Promoting IL-6/STAT3 Pathway in Mice. Inflamm Bowel Dis 2016;22:1107-18. [Crossref] [PubMed]
- Rodriguez JA, Huerta-Yepez S, Law IK, et al. Diminished expression of CRHR2 in human colon cancer promotes tumor growth and EMT via persistent IL-6/Stat3 signaling. Cell Mol Gastroenterol Hepatol 2015;1:610-30. [Crossref] [PubMed]
- Saadatdoust Z, Pandurangan AK, Ananda Sadagopan SK, et al. Dietary cocoa inhibits colitis associated cancer: a crucial involvement of the IL-6/STAT3 pathway. J Nutr Biochem 2015;26:1547-58. [Crossref] [PubMed]
- Giacopelli F, Rosatto N, Divizia MT, et al. The first intron of the human osteopontin gene contains a C/EBP-beta-responsive enhancer. Gene Expr 2003;11:95-104. [Crossref] [PubMed]
- Hugo H, Cures A, Suraweera N, et al. Mutations in the MYB intron I regulatory sequence increase transcription in colon cancers. Genes Chromosomes Cancer 2006;45:1143-54. [Crossref] [PubMed]
- Baralle M, Pastor T, Bussani E, et al. Influence of Friedreich ataxia GAA noncoding repeat expansions on pre-mRNA processing. Am J Hum Genet 2008;83:77-88. [Crossref] [PubMed]
- Buratti E, Brindisi A, Pagani F, et al. Nuclear factor TDP-43 binds to the polymorphic TG repeats in CFTR intron 8 and causes skipping of exon 9: a functional link with disease penetrance. Am J Hum Genet 2004;74:1322-5. [Crossref] [PubMed]


