
Prognostic significance of artemin in gastric cancer and its role in tumorigenesis
Introduction
Gastric cancer (GC) is one of the most frequent malignancies, accounting for 7% and 9% of annual cancer cases and mortalities, respectively (1). Its prognosis is generally poor with an overall five-year survival rate of 20–45% (2). Although the treatment of GC has made significant advances in recent years, metastasis after radical resection remains a major challenge for GC therapy (3). The underlying molecular mechanism of GC metastasis has not been fully clarified. Accordingly, it is necessary to find new biomarkers to reveal the metastasis mechanisms of GC and possibly provide an effective target for the treatment of GC.
Artemin (ARTN) is a neurotrophic factor, belonging to the glial-cell line-derived neurotrophic factor (GDNF) family of ligands (4). Mounting evidence indicates that ARTN plays a critical role in tumor growth, metastasis, and invasion of various human malignancies, such as pancreatic, esophageal, and mammary carcinoma (5-8). Nevertheless, the biological functions of ARTN in GC remain elusive.
Signal transducer and activator of transcription 3 (STAT3), a member of the STAT protein family, has been considered to be a proto-oncogene and is the convergence point of various signal transduction pathways, which are related to the proliferation, migration and angiogenesis of cancer cells (9). STAT3 can be activated by diverse oncogenic proteins, cytokines and growth factors, such as EGF and IL-6 (10). However, the relationship between ARTN and STAT3 in GC remains unclear.
The current study detected ARTN expression in GC tissues and investigated the relationship between ARTN expression and clinicopathological features of GC. Moreover, we evaluated the effects of ARTN in GC on cell proliferation, DNA synthesis, migration and invasion, and attempted to reveal its mechanism of the influence.
Methods
Patients
We collected gastric adenocarcinoma specimens that were resected in Shandong Cancer Hospital from June 2012 to January 2014. All patients did not undergo radiotherapy and/or chemotherapy before surgery. A total of 140 patients were enrolled, including 45 females and 95 males between 31 and 72 years old, with an average age of 47 years. All patients had complete clinical, pathological, and follow-up information. Additionally, 48 normal gastric mucosae were taken from around the tumors (≥5 cm) of some of these patients and confirmed by postoperative pathology. All 140 GC tissues and 48 normal gastric mucosae were selected for immunochemistry. Tumor histologic types were determined following the criterion provided by the World Health Organization (11). The pathological stage was determined based on the Unified International Gastric Cancer Staging Classification System, as incorporated in the UICC TNM classification manual (4). The patient oncological outcome was calculated from the day of surgery to the date of death or January 2019, and the median follow-up was 38 months (8 to 60 months). The ethics committee of Shandong Cancer Hospital approved the study protocol, and written informed consent was obtained from each patient.
Immunohistochemistry
Immunohistochemistry staining and result evaluation were performed as previously described (12). In brief, the tissue sections were preincubated with the first antibody (ARTN, 1:100 dilution) at 4 °C overnight, and then they were exposed to secondary antibody at room temperature. Finally, the sections were stained with a modified avidin-biotin complex (ABC) technique, and counterstained with the hematoxylin.
Cell culture
The human gastric carcinoma cell lines MGC803, SGC7901 and BGC823 were obtained from the Cell Bank of the Chinese Academy of Sciences (Shanghai, China). These cells were cultured in RPMI-1640 medium (HyClone, Logan, UT, USA) supplemented with 10% fetal bovine serum (FBS, GIBCO, Waltham, MA, USA) in humidified 5% CO2 at 37 °C.
ARTN knockdown by siRNA
Small interfering RNA (siRNA) was designed and synthesized by the GenePharma (Suzhou, China), and the nontargeted control siRNA was used as the negative control (NC). The sense sequences of the siRNA against ARTN and non-targeted control sequence are listed below: ARTN (sense: 5’-UCUTCTUCTAGGCUGACATUCT-3’, anti-sense: 5’-AUUGUCAGGCCUGAAGAGATT-3’) and NC: (sense: 5’-UUCUCCGAACGUGUCACGUTT-3’, anti-sense: 5’-ACGUGACACGUUCGGAGAATT-3’). Briefly, MGC803 cells were cultured until 30–50% confluence and incubated with serum-free 1640 to starve. Then 1.5 µg siRNA and 7.5 µL lipofectamine siRNA transfection reagent were diluted in the serum-free 1640 medium and mixed them. The mixture (siRNA/transfection reagent) was incubated at room temperature for 20 min and added directly onto the cells. After 6-8 h exposure to siRNA, the mixture was removed and continued to culture the MGC803 cell in 1640 containing 5% FBS for another 24 h and used as required.
Cell counting kit-8 (CCK-8) assay
The CCK-8 assay was used to assess cell proliferation following the manufacturer’s protocol (Dojindo, Kumamoto, Japan). Cells were seeded in 96-well plates at 5×103 cells per well. After 24 h of incubation at 37 °C, 5µL of CCK8 reagent was added into each well. After 2 h of incubation at 37 °C, a spectrophotometer was used to measure the optical density (OD) at the wavelength of 450 nm.
5-bromo-2-deoxyuridine (BrdU) assay
MGC803 cells were seeded in 96-well plates at 5×103 cells per well, and then exposed to serum-free medium for 24 h. According to the experimental group, 90 µL medium and 10 ng/mL BrdU labeling solution to be added each well, and the cells were then cultured for 24 h at 37 °C. The subsequent steps were completed according to the Millipore BrdU proliferation assay kit (Millipore, MA, USA) instructions. Finally, a spectrophotometer was used to detect the OD value at a dual-wavelength of 450/550 nm.
Cell cycle assays
Flow cytometry was used to analyze the cell cycle distribution of GC cells. Cells were cultured and harvested at 70–80% confluence, then centrifuged and suspended with cold PBS, and fixed in 70% ethanol overnight at 4 °C. After the ethanol was removed, the cells were incubated in PBS containing propidium iodide (PI) for 30 min at 37 °C. Subsequently, cellular DNA content was measured by a flow cytometer (BD Biosciences, San Jose, CA, USA).
Cell migration and invasion assays
Cell migration was evaluated using a Transwell chamber assay. Briefly, 2×104 cells (200 µL) suspended in serum-free 1640 medium were seeded into the upper chambers. Simultaneously, 600 µL of 1640 medium containing 10% FBS was added in the lower chamber. After being cultured at 37 °C for 24 h, the cells were fixed with formaldehyde and stained with 1% crystal violet. A wet cotton swab was used to remove cells in the upper part of the filter, and the cells that passed through upper filter were counted under a microscope. Additionally, a 24-well Transwell unit (8 µm pore size) which was coated with 1 mg/mL Matrigel (BD, Franklin Lakes, NJ, USA) was used to detect the cell invasion. The following steps were in accordance with the cell migration assay.
Quantitative real-time PCR
A quantitative real-time PCR (qPCR) assay was performed as previously described (12). Total RNA was reverse transcribed by using oligo-dT primers and Superscript II reverse transcriptase (Invitrogen, Grand Island, NY, USA) according to the manufacturer’s instructions. RT−qPCR was performed using the Roche Cobas 4800 System (Roche, Basel, Switzerland). Relative expression of ARTN was calculated as ΔCt, measured by subtracting the Ct of the GAPDH. ARTN primers used in this study were: forward, 5’-CCGCAGAGTCCCACTAGC-3’ and reverse, 5’-TCCTTCAAATGCTGTCCCTA-3’.
Western blot analysis
Western Blot was performed as previously described (4). PVDF membranes were blocked in 5% non-fat milk and incubated with ARTN (1:1,000), MMP-9 (1:1,000), E-cadherin (1:1,000), STAT3 (1:1,000) and p-STAT3 (Tyr705, 1:1,000; Ser727, 1:1,000) overnight at 4 °C. The blots were then incubated with secondary antibodies and enhanced chemiluminescence reagents. The automatic gel imaging instrument (Bio-Rad, Hercules, CA, USA) was used to scan the film, and the ImageJ software (NIH Image, Bethesda, MD, USA) was used for densitometric analysis.
Statistical analysis
Statistical analyses were performed using the SPSS 22.0 software (SPSS Inc., Chicago, IL, USA). Data are presented as mean ± SD. The categorical data were analyzed by the χ2 test. Patient survival was analyzed by the Kaplan-Meier survival curve and the log-rank tests. Student’s t-tests were used to compare the two independent variables. A P<0.05 was defined as statistically significant.
Results
Positive expression of ARTN predicts poor prognosis in GC patients
As shown in Figure 1A, ARTN protein was located in the cytoplasm and cell membrane. In comparison with the adjacent non-tumor normal tissues (20.8%, 10/48), the expression of ARTN in GC tissues (75.7%, 106/140) was significantly higher (P<0.05). We then evaluated the relationship between ARTN expression and the clinicopathological characteristics of GC patients. The detailed data are summarized in Table 1. There were statistically significant differences in lymph node metastasis, depth of invasion, and TNM stage between positive and negative ARTN staining tumors. Nevertheless, the expression of ARTN protein was not correlated with age, gender, degree of GC differentiation, or distant metastasis. Additionally, Kaplan-Meier analysis showed that the survival period of the ARTN positive group was significantly shorter than the negative group (P<0.05; Figure 1B), suggesting that ARTN positive expression predicted a worse prognosis for GC patients.
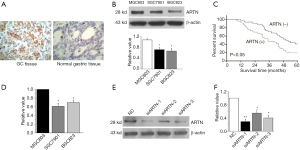
Table 1
| Clinicopathological index | N | ARTN | P | |
|---|---|---|---|---|
| − | + | |||
| Gender | 0.086 | |||
| M | 95 | 19 | 76 | |
| F | 45 | 15 | 30 | |
| Age | 0.321 | |||
| <60 | 72 | 20 | 52 | |
| ≥60 | 68 | 14 | 54 | |
| Differentiation | 0.142 | |||
| Well, moderate | 36 | 12 | 24 | |
| Poor | 104 | 22 | 82 | |
| Invasion | <0.01 | |||
| Mucosae, submucosa | 7 | 5 | 2 | |
| Muscularis | 8 | 5 | 3 | |
| Serosa | 18 | 4 | 14 | |
| Perforation of serosa | 107 | 20 | 87 | |
| Lymph node metastasis | <0.001 | |||
| Yes | 108 | 16 | 92 | |
| No | 32 | 18 | 14 | |
| Distant metastasis | 0.99 | |||
| Yes | 10 | 2 | 8 | |
| No | 130 | 32 | 98 | |
| TNM staging | 0.020 | |||
| I | 8 | 6 | 2 | |
| II | 32 | 7 | 25 | |
| III | 90 | 19 | 71 | |
| IV | 10 | 2 | 8 | |
ARTN expression was related to the presence of lymph node metastasis, depth of invasion and the TNM stage. ARTN, artemin.
ARTN is involved in GC cell proliferation and DNA synthesis in vitro
We investigated ARTN protein expression in 3 GC cell lines (MGC803, SGC7901, and BGC823) using western blot. The expression of ARTN was observed in these cells, and the maximal expression was found in MGC803 (Figure 1C). Meanwhile, we evaluated the expression of ARTN at mRNA level in the above cell lines by qPCR and obtained similar results (Figure 1D). Thence, we chose MGC803 cells as the ARTN-high expression cell for use in the subsequent experiments. Western blot showed that ARTN expression was effectively decreased by transfecting the ARTN siRNA into MGC803 cells, and the siARTN-1 had the highest interference efficiency (Figure 1E).
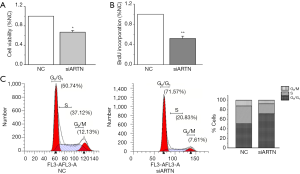
Cell viability was determined by CCK8 assay to demonstrate the effect of ARTN on the MGC803 proliferation. We found that cell viability was decreased by siARTN compared with the NC group (Figure 2A). To assess the population of cells that were actively synthesizing DNA, 5-bromodeoxyuridine (BrdU) incorporation assay was used. Our results showed that siARTN markedly inhibited BrdU incorporation in MGC803 cells (Figure 2B). All these findings indicate that ARTN has a strong promotive role in GC cell proliferation and DNA synthesis (Figure 2A,B).
ARTN RNA interference induces cell cycle arrest at G1 phase
To evaluate the mechanism of ARTN in promoting cell proliferation, we next focused on whether ARTN could induce cell cycle arrest. Flow cytometry analysis showed that the number of MGC803 cells entering the G2M+S phase in the siARTN group was decreased, while the number of cells in the G1 phase was increased compared with the NC group, suggesting ARTN silencing significantly blocked G1 transition (Figure 2C). This result showed that ARTN knockdown might inhibit cell proliferation by blocking cell cycle progression in GC.
ARTN promotes cell migration and invasion in GC
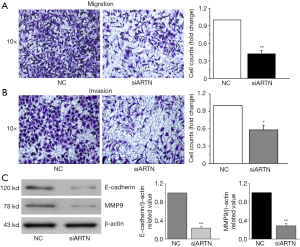
To ascertain the role of ARTN in regulating cell migration and invasion, we performed Transwell chamber assay in the MGC803 cells. After 24 h of cultivation, the number of migrating and invading cells was decreased following the ARTN knockdown in MGC803 cells compared to the NC cells (Figure 3A,B). Previous studies have indicated that E-cadherin and matrix metalloproteinases (MMPs) are involved in cancer cell migration and invasion (13,14). We found that ARTN knockdown down-regulated the expression of MMP9 and E-cadherin proteins in GC cells (Figure 3C). These findings reveal that ARTN might mediate cell migration and invasion via regulating the MMP9 and E-cadherin expression in GC.
ARTN silencing reduced the phosphorylation of STAT3
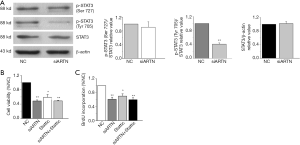
As STAT3 is considered to be a proto-oncogene and is associated with malignant behaviors such as proliferation and migration of cancer cells (9), we detected the expression of both total and activated STAT3 (p-STAT3 Ser727 and Tyr705) in MGC803 cells. Western blot showed that ARTN knockdown inhibited the phosphorylation of STAT3 (Tyr705) in MGC803 cells but had no significant influence on the level of total STAT3 and p-STAT3 (Ser727) protein (Figure 4A). Collectively, these results indicate that ARTN promoted STAT3 phosphorylation at the Tyr705 site, and thus activated the downstream targets of STAT3, which was ultimately involved in the GC progression.
ARTN mediated proliferation of MGC803 via STAT3 signaling pathway
To further ascertain the effect of ARTN and STAT3 signaling pathway in regulating cell proliferation, we treated MGC803 cells with siARTN and the Stattic (1 µM), a STAT3 pathway inhibitor. Stattic suppressed the cell viability in the same way as if it were transfected with siARTN in MGC803. However, cell viability was not further suppressed in MGC803 transfected with siARTN and treated with Stattic compared to the MGC803 transfected with siARTN or treated with Stattic alone (Figure 4B). Similarly, our findings indicated that BrdU incorporation was significantly suppressed by Stattic, whereas the group of transfections with siARTN and treated with Stattic exhibited nearly no change compared to the MGC803 transfected with siARTN or treated with Stattic alone (Figure 4C). These data revealed that STAT3 signaling pathway was implicated in the proliferation of MGC803 cells.
Discussion
In this study, we observed that ARTN was significantly more highly expressed in GC tissue than the normal gastric mucosae, and the positive expression predicted a poor clinical prognosis, which confirmed that ARTN could be used to evaluate the GC prognosis. Subsequently, we discovered that ARTN promoted cell proliferation, DNA synthesis, migration, and invasion capacities in GC in vitro. Further analysis indicated that the influence of ARTN on GC proliferation was predominantly manifested in the impact on cell cycle progression, which was mediated by activating the STAT3 signaling pathway. Moreover, ARTN promoted GC cell migration and invasion by regulating the expression of MMP9 and E-cadherin.
Previous studies have reported that ARTN can be clearly observed in various malignancies, including hepatocellular, pancreatic, and endometrial cancer, etc. (4,6,15). ARTN expression in hepatocellular cancer is related to increased tumor size, rapid recurrence, and short survival (16). To our knowledge, this study was the first to report the relationship between ARTN expression and clinicopathological characteristics and prognosis of GC patients. We also found that ARTN expression was associated with the poor prognosis and aggressive behavior of GC, including lymph node metastasis and tumor invasiveness. On the other hand, ARTN has been shown to stimulate radio- and chemo-resistance by promoting TWIST1-BCL-2-dependent cancer stem cell-like behavior in mammary carcinoma cells (17). Furthermore, ARTN enhanced oncogenicity and invasion of endometrial carcinoma cells utilizing PI3K-AKT signaling pathways (6). In this study, our data showed that migration and invasion of GC cells were decreased after silencing the ARTN. Moreover, the down-regulated expression of ARTN in GC cells significantly suppressed cell proliferation because of attenuated cell cycle progression.
As a transcription factor, STAT3 has been reported to promote cell growth and metastasis in several malignancies (10,18,19). It has two key phosphorylation sites: tyrosine phosphorylation site (p-STAT3Tyr705) and the serine phosphorylation site (p-STAT3Ser727). The Tyr705 phosphorylation event could regulate the expression of target genes by binding specific promoter sequences, and the serine 727- phosphorylated STAT3 helps maintain its transcriptional activity (20). STAT3 had a vital role in stromal cells which were recruited to the tumor microenvironment to enhance the cancer progression (21-24). Notably, the activation of STAT3 is also an effective immune checkpoint for various anti-tumor immune responses (21,23). Our results showed that siARTN decreased the expression of phospho-STAT3 (Tyr 705). Cell viability and BrdU incorporation were suppressed after blocking the STAT3 pathway with Stattic. Therefore, our data suggest that STAT3 was involved in ARTN-mediated GC cell proliferation and DNA synthesis.
In conclusion, this study confirms that, as a potential carcinogen, ARTN has a markedly higher expression in GC tissue. Furthermore, ARTN promoted cell proliferation and DNA synthesis via activating a STAT3 signaling pathway in GC, while the effects of ARTN on migration and invasion of GC cells involved MMP-9 and E-cadherin. Our study offers a novel view that ARTN participates in regulating the progression of GC, and this finding might result in a promotion of a new anticancer therapy to GC treatment.
Acknowledgments
Funding: This study was supported by
Footnote
Conflicts of Interest: All authors have completed the ICMJE uniform disclosure form (available at http://dx.doi.org/10.21037/tcr.2019.11.13). The authors have no conflicts of interest to declare.
Ethical Statement: The authors are accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved. The study was conducted in accordance with the Declaration of Helsinki (as revised in 2013). This article was approved by an independent ethical committee review board at Shandong Cancer Hospital (Shandong Cancer Hospital Ethical Committee) and written informed consent was obtained from each patient.
Open Access Statement: This is an Open Access article distributed in accordance with the Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License (CC BY-NC-ND 4.0), which permits the non-commercial replication and distribution of the article with the strict proviso that no changes or edits are made and the original work is properly cited (including links to both the formal publication through the relevant DOI and the license). See: https://creativecommons.org/licenses/by-nc-nd/4.0/.
References
- Torre LA, Bray F, Siegel RL, et al. Global cancer statistics, 2012. CA Cancer J Clin 2015;65:87-108. [Crossref] [PubMed]
- Pasechnikov V, Chukov S, Fedorov E, et al. Gastric cancer: prevention, screening and early diagnosis. World J Gastroenterol 2014;20:13842-62. [Crossref] [PubMed]
- Rau B, Brandl A, Thuss-Patience P, et al. The efficacy of treatment options for patients with gastric cancer and peritoneal metastasis. Gastric Cancer 2019;22:1226-37. [Crossref] [PubMed]
- Gao L, Bo H, Wang Y, et al. Neurotrophic Factor Artemin Promotes Invasiveness and Neurotrophic Function of Pancreatic Adenocarcinoma In Vivo and In Vitro. Pancreas 2015;44:134-43. [Crossref] [PubMed]
- Ceyhan GO, Giese NA, Erkan M, et al. The neurotrophic factor artemin promotes pancreatic cancer invasion. Ann Surg 2006;244:274-81. [Crossref] [PubMed]
- Pandey V, Qian PX, Kang J, et al. Artemin stimulates oncogenicity and invasiveness of human endometrial carcinoma cells. Endocrinology 2010;151:909-20. [Crossref] [PubMed]
- Li S, Li Z, Guo F, et al. miR-223 regulates migration and invasion by targeting Artemin in human esophageal carcinoma. J Biomed Sci 2011;18:24. [Crossref] [PubMed]
- Banerjee A, Wu ZS, Qian P, et al. ARTEMIN synergizes with TWIST1 to promote metastasis and poor survival outcome in patients with ER negative mammary carcinoma. Breast Cancer Res 2011;13:R112. [Crossref] [PubMed]
- Yu H, Lee H, Herrmann A, et al. Revisiting STAT3 signalling in cancer: new and unexpected biological functions. Nat Rev Cancer 2014;14:736-46. [Crossref] [PubMed]
- Liu X, Chen B, You W, et al. The membrane bile acid receptor TGR5 drives cell growth and migration via activation of the JAK2/STAT3 signaling pathway in non-small cell lung cancer. Cancer Letters 2018;412:194-207. [Crossref] [PubMed]
- Baloh RH, Gorodinsky A, Golden JP, et al. GFRalpha3 is an orphan member of the GDNF/neurturin/persephin receptor family. Proc Natl Acad Sci U S A 1998;95:5801-6. [Crossref] [PubMed]
- Guo J, Fan KX, Xie LI, et al. Effect and prognostic significance of the KAI1 gene in human gastric carcinoma. Oncol Lett 2015;10:2035-42. [Crossref] [PubMed]
- Shih YW, Lee YC, Wu PF, et al. Plumbagin inhibits invasion and migration of liver cancer HepG2 cells by decreasing productions of matrix metalloproteinase-2 and urokinase- plasminogen activator. Hepatol Res 2009;39:998-1009. [Crossref] [PubMed]
- Wijnhoven BP, Dinjens WN, Pignatelli M. E-cadherin-catenin cell-cell adhesion complex and human cancer. Br J Surg 2000;87:992-1005. [Crossref] [PubMed]
- Kang J, Qian PX, Pandey V, et al. Artemin is estrogen regulated and mediates antiestrogen resistance in mammary carcinoma. Oncogene 2010;29:3228-40. [Crossref] [PubMed]
- Hezam K, Jiang J, Sun F, et al. Artemin promotes oncogenicity, metastasis and drug resistance in cancer cells. Rev Neurosci 2018;29:93-8. [Crossref] [PubMed]
- Banerjee A, Qian P, Wu ZS, et al. Artemin stimulates radio- and chemo-resistance by promoting TWIST1-BCL-2-dependent cancer stem cell-like behavior in mammary carcinoma cells. J Biol Chem 2012;287:42502-15. [Crossref] [PubMed]
- Yang L, Ma C, Zhang L, et al. 15-Lipoxygenase-2/15(S)-hydroxyeicosatetraenoic acid regulates cell proliferation and metastasis via the STAT3 pathway in lung adenocarcinoma. Prostaglandins Other Lipid Mediat 2018;138:31-40. [Crossref] [PubMed]
- Tang Q, Zheng F, Wu J, et al. Combination of Solamargine and Metformin Strengthens IGFBP1 Gene Expression Through Inactivation of Stat3 and Reciprocal Interaction Between FOXO3a and SP1. Cell Physiol Biochem 2017;43:2310-26. [Crossref] [PubMed]
- Ouedraogo ZG, Biau J, Kemeny JL, et al. Role of STAT3 in Genesis and Progression of Human Malignant Gliomas. Mol Neurobiol 2017;54:5780-97. [Crossref] [PubMed]
- Zhang L, Alizadeh D, Van Handel M, et al. Stat3 inhibition activates tumor macrophages and abrogates glioma growth in mice. Glia 2009;57:1458-67. [Crossref] [PubMed]
- Herrmann A, Kortylewski M, Kujawski M, et al. Targeting Stat3 in the myeloid compartment drastically improves the in vivo antitumor functions of adoptively transferred T cells. Cancer Res 2010;70:7455-64. [Crossref] [PubMed]
- Yu H, Kortylewski M, Pardoll D. Crosstalk between cancer and immune cells: role of STAT3 in the tumour microenvironment. Nat Rev Immunol 2007;7:41-51. [Crossref] [PubMed]
- Wang L, Yi T, Kortylewski M, et al. IL-17 can promote tumor growth through an IL-6-Stat3 signaling pathway. J Exp Med 2009;206:1457-64. [Crossref] [PubMed]


