
B4GALNT1 enhances cell proliferation and growth in oral squamous cell carcinoma via p38 and JNK MAPK pathway
Introduction
Head and neck cancer ranks the sixth among the most common malignant tumor around the world (1). It is also reported that oral squamous cell carcinoma (OSCC) occupies almost 90% of the cancer found in head and neck region (2). The prognosis of the cancer is highly associated with the stages, differentiation as well as the lymph node metastasis of the tumor (3-5). Although the diagnostic and therapeutic measures including surgical resection, radiotherapy, chemotherapy and comprehensive sequence therapy have been proven remarkable measures to achieve better life quality recently, the morbidity and mortality of OSCC remain high and the overall 5-year survival rate for OSCC patients is reported to be relatively low, which rendering OSCC still a global concern of cancer-related death (3,6).
Beta-1,4-N-Acetyl-Galactosaminyltransferase 1 (B4GALNT1) is an enzyme encoded by the B4GALNT (7,8) in human cells. B4GALNT1 has been found to widely express in various organs among which the central nervous system contains B4GALNT1 much more than the other parts of the body (9). There are researches confirming that normal brain tissues and some malignant transformed cells can be observed to highly express B4GALNT1, such as malignant melanoma, neuroblastoma and the adult T cell leukemia (10-12). It is reported that B4GALNT1 takes part in the normal function of nervous systems and the development of some neurogenic diseases (12). However, the role B4GALNT1 plays in the development and progression of tumorigenesis in head and neck region has not been reported yet. Recently, we have discovered B4GALNT1 was highly expressed in OSCC cells and therefore this study aimed to investigate whether B4GALNT1 has any relationship with the tumorigenesis of OSCC and to try to explore the possible regulation mechanism behind it.
Methods
Patients and cell lines
We used Dulbecco’s modified Eagle’s medium (DMEM) which contained 10% fetal bovine serum (FBS) to culture human OSCC cell line CAL-27. Another OSCC cell line SCC-25 was properly cultured in the DMEM and Ham’s F12 medium mixed in the ratio of 1:1 with 10% FBS.
We collected 30 oral cancer samples and 10 non-neoplastic oral mucosa samples from the Fifth Affiliated Hospital of Sun Yat-sen University from September 2015 to September 2018. Quantitative real-time PCR was carried out to check the expression of B4GALNT1 in these samples and the relationship between its expression and clinicopathological data was analyzed.
All of the analyses were performed in accordance with the principle of relevant guidelines and regulations. Besides, Informed consent was obtained from the participants enrolled.
The Cancer Genome Atlas (TCGA) gene expression data
B4GALNT1 mRNA levels in 15 OSCC specimens and paired normal adjacent mucosal tissues were investigated and analyzed. The level 3 RNA-Seq and pertinent clinical data of HNSCC patients were obtained from TCGA database (https://tcga-data.nci.nih.gov/tcga/).
B4GALNT1 shRNA Design and Lentivirus Construction
Small interfering RNA (shRNA) precisely targeting B4GALNT1 was created and lentivirus was constructed to effectively express B4GALNT1 shRNA. shRNA targeting B4GALNT1 (target sequence as TACTCTGAAAGCAAGCAAGAA) was designed while a scramble sequence (TTCTCCGAACGTGTC ACGT) was selected and used as a negative control (NC). After that, we synthesized, annealed and inserted the stem-loop DNA oligonucleotides into the lentiviral vector GV115 which carries green fluorescent protein (GFP) gene. We chose Lentivector Expression Systems (GeneChem, Shanghai, China) to create lentivirus and transfected an oral squamous cell line called CAL-27. In the meantime, quantitative real-time PCR (qPCR) was performed to examine the knockdown efficacy of B4GALNT1.
Cell proliferation was detected by Celigo cell counting and MTT assay
Digested with 0.25% Trypsin (Thermo Fisher Scientific), cells transfected with shCtrl or shB4GALNT1 were seeded in 96-well plates at a density of 2000 cells/well. We did cell counts via the Celigo imaging cytometer (Nexcelom Bioscience, Lawrence, MA) was used to investigate viable cells once a day for consecutive five days and cell growth curves were created accordingly. 3-(4,5-dimethylthiazol-2- yl)-2,5-diphenyltetrazolium bromide (MTT, Sigma, USA) assay was performed to analyze the cell count. Five groups of Cal-27 cells were seeded into 96 well plate with the density of 2,000 cells per well. On the consecutive 5 days afterwards, one plate on each day was removed to be added 20 µL of MTT (5 mg/mL). Incubating at 37 °C for 4 hours, absorbance at 490 nm was measured on the microplate reader after the removal of MTT.
Flow Cytometry Analysis
CAL-27 cells transfected with either shB4GALNT1 or shCtrl were detected by flow cytometry analysis (FCM). After achieving 70% confluence, cells were harvested and mixed with binding buffer and staining buffer afterward. 100ul cell suspension was stained with 5ul Annexin V-APC (eBioscience, San Diego, US) and finally subjected to flow cytometry.
RNA Isolation and Quantitative Real-Time PCR
Total RNA extraction from CAL-27 cells by trizol reagent (Invitrogen) and Quantitative Real-Time PCR was performed as previously described. The housekeeping gene GAPDH was used as an internal control. 2−ΔΔ CT method was used to quantify the relative gene expression. In term of the mRNA expression of B4GALNT1 in OSCC tissue, the median levels of B4GALNT1 were used as cutoff values for dividing high or low expression.
Western blotting
In the western blotting, RIPA lysis buffer was used to extract total proteins from cultured cells. 20 µg of protein samples were subjected to 10% sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE) before transferred to the polyvinylidene difluoride (PVDF) membrane. Then we sealed the membrane with 5% skimmed milk and incubate it overnight with primary antibodies: anti-JNK (CST, 1:500), anti-p-JNK (CST, 1:500), anti-p38 (CST, 1:1,000), anti-p-p38(CST, 1:1,000) and anti-GAPDH(Santa-Cruz, 1:500). Afterwards, the peroxidase- labeled secondary antibody (anti-rabbit IgG, 1:5,000, Cell Signal, USA) was used to incubate the PVDF membrane for 2 hours at room temperature. Finally, the protein bands were visualized by Pierce™ ECL Plus Western Blotting Substrate (Thermo, USA).
Cell cycle analysis
Cell cycle distribution was detected by flow cytometry analysis using staining solution [40×propidium iodide (PI; 2 mg/mL, Sigma-Aldrich® Co.LLC., St. Louis, US): 100×RNase A (10 mg/mL, Fermentas®, Vilnius, Lithuania): 1 × D-Hanks =25:10:1,000]. The fluorescence of DNA-bound PI in cells was measured with FACS Calibur™ Flow Cytometer (BD Biosciences, Franklin Lakes, US), and the cell populations in different phases of the cell cycle were analyzed with ModFit 3.0 software.
Statistics analysis
All statistical analyses were performed using SPSS 18.0 statistical software (SPSS Inc., Chicago, IL, USA). Statistical differences were determined by the two-tailed student’s t-test between two groups for quantitative data, and chi-square test for qualitative data; a P value <0.05 was considered statistically significant.
Results
Overexpression of B4GALNT1 was found in oral cancer tissues as well as OSCC cell lines
Analysis into B4GALNT1 expression of oral cancer tissues and normal oral mucosa samples from 15 OSCC patients containing RNA sequencing results in TCGA revealed that B4GALNT1 was expressed higher in oral cancer tissues than in normal oral paratumor mucosa tissues (P<0.01) (Figure 1A). Similarly, we found significantly higher expression of B4GALNT1 in OSCC samples comparing with normal oral mucosa samples. Our data suggested that high B4GALNT1 expression was associated with advanced T stage (P<0.05) (Table 1). Also, we detected expression of B4GALNT1 in two OSCC cell lines: CAL-27 and SCC-25, whose quantitative PCR results showed that both OSCC cell lines had higher B4GALNT1 expression than the internal control (Figure 1B).
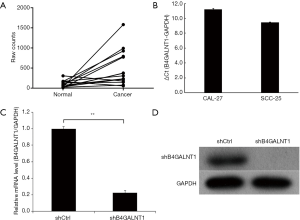
Table 1
| Characteristics | B4GALNT1 | |||
|---|---|---|---|---|
| n | High | Low | P value | |
| Age, years | 0.282 | |||
| <50 | 14 | 6 | 8 | |
| ≥50 | 16 | 10 | 6 | |
| Sex | 0.510 | |||
| Male | 19 | 11 | 8 | |
| Female | 11 | 5 | 6 | |
| Clinical stage | 0.961 | |||
| Low (I–II) | 13 | 7 | 6 | |
| High (III–IV) | 17 | 9 | 8 | |
| T stage | 0.028 | |||
| T1–T2 | 15 | 5 | 10 | |
| T3–T4 | 15 | 11 | 4 | |
| Cervical lymph node metastasis | 0.811 | |||
| Yes | 9 | 4 | 5 | |
| No | 21 | 12 | 9 | |
| Pathological typing | 0.431 | |||
| Highly differentiated | 17 | 8 | 9 | |
| Moderate-low differentiation | 13 | 8 | 5 | |
P-values were calculated by the chi-squared test. B4GALNT1, Beta-1,4-N-Acetyl-Galactosaminyltransferase 1.
B4GALNT1 shRNA specifically suppressed B4GALNT1 expression
After shRNA infection to CAL-27 cell lines was completed, we performed quantitative real-time PCR (qPCR) to detect relative mRNA level and western blotting to examine the protein level of expression. The results showed that shRNA significantly inhibited mRNA and protein expression of B4GALNT1 in CAL-27 three days after shRNA infection compared to shCtrl group (P<0.01) (Figure 1C,D). The knockdown efficacy was counted to be 77.6%.
B4GALNT1 shRNA inhibited CAL-27 cells proliferation and suppressed the growth of tumor cells
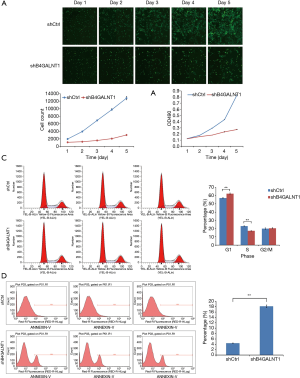
We cultured and counted by Celigo the CAL-27 cells every day for the 5 consecutive days after the shB4GALNT1 and shCtrl transfection were completed. According to the results, a growth curve was generated, which showed the growth of cells in shB4GALNT1 group was significantly inhibited compared to the shCtrl group (Figure 2A). Also, MTT assay was performed and the results revealed the similar difference in cell growth between two groups. (Figure 2B). Therefore, from the similar results of Celigo test and MTT assay, proliferation was significantly inhibited in cells transfected with shB4GALNT1 compared to that in shCtrl group.
G1-phase cell cycle arrest was induced by B4GALNT1 knockdown
We performed flow cytometry analysis (FCM) to examine whether B4GALNT1 knockdown would have any influence in CAL-27 cell cycle and the results showed cell cycle percentage in G1 phase was significantly climbed to 62.4%±0.77% in B4GALNT1 knockdown group from 57.11%±0.79% in control group (P<0.01) while cell percentage in S phase presented a significant drop to 17.19%±0.23% in B4GALNT1-shRNA-treated group from 22.98%±0.41% in control group (P<0.01). There was no notable change in cell cycle percentage in G2/M phase (20.41%±0.69% in knockdown group vs. 19.91%±0.83% in control group, P>0.05) (Figure 2C).
B4GALNT1 shRNA induced cell apoptosis in CAL-27 cell lines compared to the control group
Flow cytometry analysis showed that a notable increase in proportion of apoptotic cells was detected in shB4GALNT1 (4.42%±0.09%) compared to the shCtrl group (18.16%±0.46%) (P<0.01) (Figure 2D). This result suggested that B4GALNT1 knockdown may participate in the progress of cell apoptosis in CAL-27 cells.
Western blotting of protein expression
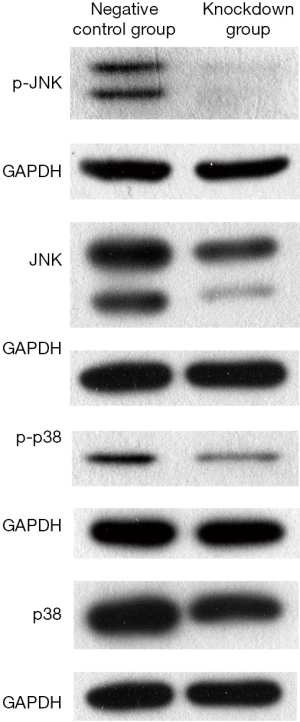
The results of western blotting revealed the expression of JNK, p-JNK, p38 and p-p38 in CAL-27 cell lines was all significantly inhibited in B4GALNT1 knockdown group compared to the negative control group after silencing B4GALNT1 expression. GAPDH was chosen as internal control (Figure 3).
Discussion
B4GALNT1 is an enzyme encoded by B4GALNT1 (8). In recent years, a mounting number of researches have shown that B4GALNT1 plays an essential role in the development and normal function of the central nervous system. There are studies associating the mutation of B4GALNT1 with some biochemical reactions like loss of enzyme activity and ganglioside biosynthesis (12,13), and with some systemic diseases such as hereditary spastic paraplegias, neuroblastoma and type 1 diabetes mellitus (14-18). Yang et al. (19) reported that B4GALNT1 was highly expressed in clear cell renal cell carcinoma (ccRCC) and found that spread of ccRCC may associated with the complement system, cholesterol metabolism and the calcium pathways. However, there are limited researches investigating the relationship between the expression of B4GALNT1 and the development of tumor in head and neck region. Pondering this question, here our group compared and discovered that 15 OSCC tissues extracted from TCGA database presented significantly higher expression of B4GALNT1 compared with the paired normal tissue from para-cancerous area. Further investigation on the cell lines experiment confirmed a much higher expression of B4GALNT1in CAL-27 and SCC-25 than that of the internal control. All of these discoveries indicated that a higher expression of B4GALNT1 may act a potential role in the development of OSCC. And to date, specific regulation mechanism has not been reported yet.
To our knowledge, the result of present cell function experiments revealed that cell count presented a remarkable decrease in B4GALNT1 shRNA group compared with the control group, which suggested that proliferation of tumor cell was greatly inhibited after knockdown of B4GALNT1. It was indicated, in other words, that B4GALNT1 was highly associated with the ability of cell proliferation in CAL-27 cell lines. Moreover, the increased proportion of apoptotic cells in flow cytometry analysis suggested that cell apoptosis in OSCC cell lines can be triggered by knockdown of B4GALNT1 via lentivirus-mediated shRNA system. In addition, there is so far few studies investigating whether silencing B4GALNT1 can impact cell cycle. Our results revealed the knockdown of B4GALNT1 resulted in cell cycle arrest at G1 phase when synthesis of RNA and ribosome necessary for next phase is the most biochemical reaction in the cells. However, decreasing proportion of S phase in CAL-27 cell lines when synthesis of DNA and essential enzymes for DNA duplication is completed in cells suggests that the proliferation and development of OSCC may be impacted after the knockdown of B4GALNT1, which from another aspect supports the positive relationship between B4GALNT1 and tumorigenesis of OSCC.
The Mitogen Activated Protein Kinase (MAPK) pathway has been proven to actively participate in the development and progression of various cancers (20). MAPK family includes proteins such as ERK, JNK and p38, which have been discovered to play essential roles in the migration of different cell types (21,22). In previous studies, there exists researches showing p38 signaling pathway in breast cancer cells is a key regulation pathway in the suppression of tumor metastasis (23,24). Zheng et al. has demonstrated that inhibiting the phosphorylation of p-JNK and p-p38 can repress the migration of cervical cancer cells (25). In present study, we discovered that the expression of JNK and p38 after the knockdown of B4GALNT1, no matter phosphorylated or dephosphorylated, were both significantly decreased compared to the negative control, which we consider MAPK a potential regulation mechanism of signaling pathway between B4GALNT1 and the tumorigenesis of OSCC. B4GALNT1 may regulate the cell proliferation, apoptosis and cell cycle to affect the development and progression of OSCC by regulating the expression of downstream genes and activating JNK and p38 MAPK signaling pathway. However, detailed molecular mechanism by which B4GALNT1 regulates JNK and p38 signaling pathway has not been reported yet. Thus, our study has provided a new prospective of treatment strategy for OSCC patients and deeper research on the specific molecular mechanism remains to be carried out in future experiments.
In conclusion, within the limitation of the study, the present research confirmed B4GALNT1 was found to overexpress in OSCC and further cell function experiments proved that B4GALNT1 enhanced the proliferation and suppressed the apoptosis of oral cancer cells. All of these phenotypes of cancer cells might be mediated through JNK, p38 MAPK pathway. However, detailed regulation mechanism on molecular aspect remained unknown, which provided a new possibility to perform deeper researches in our future experiments.
Acknowledgments
Funding: None.
Footnote
Conflicts of Interest: All authors have completed the ICMJE uniform disclosure form (available at http://dx.doi.org/10.21037/tcr.2020.03.73). The authors have no conflicts of interest to declare.
Ethical Statement: The authors are accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved. The study was approved by the institutional ethics board of our hospital (No. [2019]K149-1). Informed consent was obtained from the participants enrolled. The study was conducted in accordance with the Declaration of Helsinki (as revised in 2013).
Open Access Statement: This is an Open Access article distributed in accordance with the Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License (CC BY-NC-ND 4.0), which permits the non-commercial replication and distribution of the article with the strict proviso that no changes or edits are made and the original work is properly cited (including links to both the formal publication through the relevant DOI and the license). See: https://creativecommons.org/licenses/by-nc-nd/4.0/.
References
- Tandon P, Dadhich A, Saluja H, et al. The prevalence of squamous cell carcinoma in different sites of oral cavity at our Rural Health Care Centre in Loni, Maharashtra - a retrospective 10-year study. Contemp Oncol (Pozn) 2017;21:178-83. [Crossref] [PubMed]
- Hosni A, Huang SH, Chiu K, et al. Predictors of Early Recurrence Prior to Planned Postoperative Radiation Therapy for Oral Cavity Squamous Cell Carcinoma and Outcomes Following Salvage Intensified Radiation Therapy. Int J Radiat Oncol Biol Phys 2019;103:363-73. [Crossref] [PubMed]
- Wang Q, Zhi Y, Ren W, et al. Suppression of OSCC malignancy by oral glands derived-PIP identified by iTRAQ combined with 2D LC-MS/MS. J Cell Physiol 2019; [Crossref] [PubMed]
- Michalek J, Brychtova S, Pink R, et al. Prognostic and predictive markers for perineural and bone invasion of oral squamous cell carcinoma. Biomed Pap Med Fac Univ Palacky Olomouc Czech Repub 2019;163:302-8. [Crossref] [PubMed]
- Druy AE, Shorikov EV, Tsaur GA, et al. Prospective investigation of applicability and the prognostic significance of bone marrow involvement in patients with neuroblastoma detected by quantitative reverse transcription PCR. Pediatr Blood Cancer 2018;65:e27354. [Crossref] [PubMed]
- Liu T, David M, Ellis O, et al. Treatment for oral squamous cell carcinoma: Impact of surgeon volume on survival. Oral Oncol 2019;96:60-5. [Crossref] [PubMed]
- Nagata Y, Yamashiro S, Yodoi J, et al. Expression cloning of beta 1,4 N-acetylgalactosaminyltransferase cDNAs that determine the expression of GM2 and GD2 gangliosides. J Biol Chem 1992;267:12082-9. [PubMed]
- Miyazaki T, Miyashita R, Nakamura S, et al. Biochemical characterization and mutational analysis of silkworm Bombyx mori beta-1,4-N-acetylgalactosaminyltransferase and insight into the substrate specificity of beta-1,4-galactosyltransferase family enzymes. Insect Biochem Mol Biol 2019;115:103254. [Crossref] [PubMed]
- Schnaar RL. Gangliosides of the Vertebrate Nervous System. Journal of Molecular Biology 2016;428:3325-36. [Crossref] [PubMed]
- Furukawa K, Soejima H, Niikawa N, et al. Genomic organization and chromosomal assignment of the human beta1, 4-N-acetylgalactosaminyltransferase gene. Identification of multiple transcription units. J Biol Chem 1996;271:20836-44. [Crossref] [PubMed]
- Yamaguchi T, Yamauchi Y, Furukawa K, et al. Expression of B4GALNT1, an essential glycosyltransferase for the synthesis of complex gangliosides, suppresses BACE1 degradation and modulates APP processing. Sci Rep 2016;6:34505. [Crossref] [PubMed]
- Li TA, Schnaar RL. Congenital Disorders of Ganglioside Biosynthesis. Prog Mol Biol Transl Sci 2018;156:63-82. [Crossref] [PubMed]
- Liang L, Weng J, You Y, et al. Role of Noxa in proliferation, apoptosis, and autophagy in human adenoid cystic carcinoma. J Oral Pathol Med 2019;48:52-9. [Crossref] [PubMed]
- Schwarzmann G. Labeled gangliosides: their synthesis and use in biological studies. FEBS Lett 2018;592:3992-4006. [Crossref] [PubMed]
- Trinchera M, Parini R, Indellicato R, et al. Diseases of ganglioside biosynthesis: An expanding group of congenital disorders of glycosylation. Mol Genet Metab 2018;124:230-7. [Crossref] [PubMed]
- Bhuiyan RH, Ohmi Y, Ohkawa Y, et al. Loss of Enzyme Activity in Mutated B4GALNT1 Gene Products in Patients with Hereditary Spastic Paraplegia Results in Relatively Mild Neurological Disorders: Similarity with Phenotypes of B4galnt1 Knockout Mice. Neuroscience 2019;397:94-106. [Crossref] [PubMed]
- Travaglini L, Aiello C, Stregapede F, et al. The impact of next-generation sequencing on the diagnosis of pediatric-onset hereditary spastic paraplegias: new genotype-phenotype correlations for rare HSP-related genes. Neurogenetics 2018;19:111-21. [Crossref] [PubMed]
- Wakil SM, Monies DM, Ramzan K, et al. Novel B4GALNT1 mutations in a complicated form of hereditary spastic paraplegia. Clin Genet 2014;86:500-1. [Crossref] [PubMed]
- Yang H, Li W, Lv Y, et al. Exploring the mechanism of clear cell renal cell carcinoma metastasis and key genes based on multi-tool joint analysis. Gene 2019;720:144103. [Crossref] [PubMed]
- Dad R, Malik U, Javed A, et al. Structural annotation of Beta-1,4-N-acetyl galactosaminyltransferase 1 (B4GALNT1) causing Hereditary Spastic Paraplegia 26. Gene 2017;626:258-63. [Crossref] [PubMed]
- Ban Z, He J, Tang Z, et al. LRG1 enhances the migration of thyroid carcinoma cells through promotion of the epithelialmesenchymal transition by activating MAPK/p38 signaling. Oncol Rep 2019;41:3270-80. [PubMed]
- Li L, Zhang J, Zhang Q, et al. High Glucose Suppresses Keratinocyte Migration Through the Inhibition of p38 MAPK/Autophagy Pathway. Front Physiol 2019;10:24. [Crossref] [PubMed]
- Ge D, Gao J, Han L, et al. Novel effects of sphingosylphosphorylcholine on the apoptosis of breast cancer via autophagy/AKT/p38 and JNK signaling. J Cell Physiol 2019;234:11451-62. [Crossref] [PubMed]
- Wada M, Canals D, Adada M, et al. P38 delta MAPK promotes breast cancer progression and lung metastasis by enhancing cell proliferation and cell detachment. Oncogene 2017;36:6649-57. [Crossref] [PubMed]
- Zheng HY, Shen FJ, Tong YQ, et al. PP2A Inhibits Cervical Cancer Cell Migration by Dephosphorylation of p-JNK, p-p38 and the p-ERK/MAPK Signaling Pathway. Curr Med Sci 2018;38:115-23. [Crossref] [PubMed]


