
High expression of GNB4 predicts poor prognosis in patients with Helicobacter pylori-positive advanced gastric cancer
Introduction
Gastric cancer (GC) is one of the most common and lethal malignancies worldwide, especially in east Asia (1,2). It remains the fourth most frequent type of cancer and the second leading cause of cancer-related death worldwide (3). In China, both incidence and morbidity rate of GC ranks No.2 of all malignancies according to GLOBOCAN2018 statistics (4). Among the identified high-risk factors contributing to gastric carcinogenesis and progression, Helicobacter pylori (H. pylori) is classified as the definite carcinogen (Group I) and is recognized as the most common etiologic agent of infection-related GC (5). H. pylori plays an unequivocal role in the development of various gastric diseases such as chronic gastritis, gastroduodenal ulcers, gastric mucosa-associated lymphoid tissue (MALT) lymphoma and most importantly, GC (6). It is estimated that approximately 70% of GC burden worldwide is attributable to abnormalities induced by H. pylori (7). Therefore, a large number of studies have been conducted to identify prognostic factors to predict the long-term outcome of H. pylori-positive GC patients. Since virulence factors of H. pylori with high pathogenicity have been considered critically involved in the development of GC, several virulence antigens such as vacuolating toxin A (VacA), cytotoxin-associated gene A (CagA) and H. pylori-related adhesins have been applied in serologic screening to predict the survival rate of GC patients with H. pylori infection (8). Abnormally expressed microRNAs, such as miR-375, miR-490-3p and miR-27a rs895819 variant, have been identified as another major set of prognostic factors for H. pylori-positive GC patients as these identified microRNAs affect the proliferation, differentiation and apoptosis of stomach mucosal epithelial cell infected with H. pylori (9,10). In addition, several determinants in GC development such as vascular endothelial growth factor-A and -C (VEGFA and VEGFC) were shown to be overexpressed in H. pylori-positive GC patients and their clinical significances were investigated (11). However, despite all these efforts to identify only a limited number of GC tissue-based biomarkers with prognostic values for H. pylori-positive GC patients have been reported.
One major strategy to discover reliable prognostic factor is the immunohistochemistry (IHC)-based staining of candidate biomarker on surgically resected tissue, followed by the statistical analysis examining the potential correlation between its expression and clinical significance. For instance, Her2 is identified to promote GC progression and metastasis and its expression status was confirmed to predict the long-term survival rate of stage III/IV GC, which leads to the landmark development of target therapy by Trastuzumab (12,13). Other prominent examples include Bcl-2 (14), E-cadherin (15), EGFR, VEGFR (16), etc. Guanine nucleotide-binding protein beta-4 (GNB4) encodes Gβ subunit 4, which is a key component of heterotrimeric G protein composed of α, β and γ subunit. Once activated by upstream signals, Gβ couples with Gγ subunit and actively mediates downstream signal transduction cascades and regulates cell proliferation, apoptosis, differentiation, motility, etc. (17). Recently, studies have demonstrated the impact of GNB4 on multiple malignancies: in breast cancer, GNB4 plays crucial roles in both tumor growth and chemoresistance (18,19). For patients with urothelial bladder carcinoma, intron-1 haplotypes of GNB4 was shown to be significantly associated with patient overall survival (20). Similar findings were shown in patients with late stage of colorectal cancer (21). Regarding the role of GNB4 in GC, Liu et al. analyzed DNA methylation and gene expression profiling of H. pylori-induced GC patients from multiple datasets of Gene Expression Omnibus (GEO). Notably, GNB4 was among the five identified hub genes of which the hypomethylation and high mRNA expression served as independent unfavorable prognostic factors of H. pylori-induced GC patients (22). Although the clinical significance and prognostic value of GNB4 were not further validated clinically nor functionally, this intriguing study laid the basis for our investigation.
Considering the clinical significance of GNB4 in H. pylori-positive GC patients, in the present study we firstly performed GNB4 expression and survival analysis by analyzing TCGA (The Cancer Genome Atlas) and Genotype-Tissue Expression (GTEx) databases of GC patients with and without H. pylori infection. Next, we evaluated the IHC staining-based expression levels of GNB4 in both early and advanced GC patients infected with H. pylori. The aim of this study was to determine the potential correlation between GNB4 expression and clinicopathological features of H. pylori-positive GC patients. Furthermore, we aimed to evaluate the prognostic value of GNB4 expression for GC patients infected with H. pylori by examining the association between its expression and 5-year OS of these patients.
We present the following article in accordance with the STROBE reporting checklist (available at http://dx.doi.org/10.21037/tcr-19-2914).
Methods
This study was designed to perform multiple statistical analysis not only to investigate the potential correlation between GNB4 expression and clinicopathological parameters of H. pylori-positive GC patients but also to investigate if GNB4 expression predicts the long-term survival of these patients.
Patients
The study was conducted in accordance with the Declaration of Helsinki (as revised in 2013). The study was approved by the local ethics committee of Ruijin Hospital Affiliated to Shanghai Jiaotong University School of Medicine and informed consent was taken from all the patients. This analysis was conducted on GC specimens from curative surgery performed in Ruijin Hospital Affiliated to Shanghai Jiaotong University School of Medicine between January 2013 and January 2015. In total, a number of 448 H. pylori-infected GC patients with no evidence of distant metastasis received treatment, including 137 EGC patients undergoing standard D2 gastrectomy (implicating the complete removal of group 1 and 2 lymph nodes) alone and 311 AGC patients undergoing the same surgery followed by chemotherapy. The chemotherapy was based on 5-fluorouracil (5-FU) as it was administered alone or combined with other cytotoxic drugs. No distant metastasis such as liver, lung, ovarian, uterus and peritoneum metastasis were assured before proceeding with surgery. None of the patients received neoadjuvant treatment prior to surgery. Clinicopathological features including age, gender, tumor location, Lauren’s classification, tumor differentiation as well as pathological TNM staging (mentioned below “Pathology”) are summarized in Table 1 and Table 2. At the time of data analysis, the death of 89 patients and loss to follow-up of 61 patients were observed and recorded. The follow-up was conducted by consulting their electrical medical record by telephone. Median follow-up time was 52 (range, 3–77) months.
Table 1
| Variable | Number (all patients =137) | GNB4 | P value | |
|---|---|---|---|---|
| GNB4 low expression (n=47) | GNB4 high expression (n=90) | |||
| Age (years) | 0.809 | |||
| <60 | 68 (49.6%) | 24 (35.3%) | 44 (64.7%) | |
| ≥60 | 69 (50.4%) | 23 (33.3%) | 46 (66.7%) | |
| Gender | 0.421 | |||
| Male | 90 (65.7%) | 33 (36.7%) | 57 (63.3%) | |
| Female | 47 (34.3%) | 14 (29.8%) | 33 (70.2%) | |
| Differentiation | 0.411 | |||
| Low | 88 (64.2%) | 28 (31.8%) | 60 (68.2%) | |
| Medium/High | 49 (35.8%) | 19 (38.8%) | 30 (61.2%) | |
| Lauren’s classification | 0.353 | |||
| Diffuse | 51 (37.2%) | 15 (29.4%) | 36 (70.6%) | |
| Intestinal | 86 (62.8%) | 32 (37.2%) | 54 (62.8%) | |
| Site | 0.949 | |||
| Cardia | 9 (6.6%) | 3 (33.3%) | 6 (66.7%) | |
| Non-cardia | 128 (93.4%) | 44 (34.4%) | 84 (65.6%) | |
| Lymph node metastasis | 0.678 | |||
| N0 (negative) | 99 (72.3%) | 35 (35.4%) | 64 (64.6%) | |
| N1–3 (positive) | 38 (27.7%) | 12 (31.6%) | 26 (68.4%) | |
| Stage | 0.633 | |||
| I | 123 (89.8%) | 43 (35.0%) | 80 (65.0%) | |
| II + III | 14 (10.2%) | 4 (28.6%) | 10 (71.4%) | |
Table 2
| Variable | Number (all patients n=311) | GNB4 | P value | |
|---|---|---|---|---|
| GNB4 low expression (n=116) | GNB4 high expression (n=195) | |||
| Age | 0.652 | |||
| <60 | 123 (39.5%) | 44 (35.8%) | 79 (64.2%) | |
| ≥60 | 188 (60.5%) | 72 (38.3%) | 116 (61.7%) | |
| Gender | 0.64 | |||
| Male | 234 (75.2%) | 89 (38.0%) | 145 (62.0%) | |
| Female | 77 (24.8%) | 27 (35.1%) | 50 (64.9%) | |
| Differentiation | 0.312 | |||
| Low | 234 (75.2%) | 91 (38.9%) | 143 (61.1%) | |
| Medium/high | 77 (24.8%) | 25 (32.5%) | 52 (67.5%) | |
| Lauren’s classification | 0.833 | |||
| Diffuse | 81 (26.0%) | 31 (38.3%) | 50 (61.7%) | |
| Intestinal | 230 (74.0%) | 85 (37.0%) | 145 (63.0%) | |
| Site | 0.42 | |||
| Cardia | 83 (26.7%) | 34 (41.0%) | 49 (59.0%) | |
| Non-cardia | 228 (73.3%) | 82 (36.0%) | 146 (64.0%) | |
| Tumor invasion | 0.148 | |||
| T2 | 53 (17.0%) | 26 (49.1%) | 27 (50.9%) | |
| T3 | 81 (26.0%) | 29 (35.8%) | 52 (64.2%) | |
| T4 | 177 (56.9%) | 61 (34.5%) | 116 (65.5%) | |
| Lymph node metastasis | 0.07 | |||
| N0 | 89 (28.6%) | 40 (44.9%) | 49 (55.1%) | |
| N1 | 66 (21.2%) | 25 (37.9%) | 41 (62.1%) | |
| N2 | 71 (22.8%) | 27 (38.0%) | 44 (62.0%) | |
| N3 | 85 (27.3%) | 24 (28.2%) | 61 (71.8%) | |
| Stage | 0.047 | |||
| I | 27 (8.7%) | 14 (51.9%) | 13 (48.1%) | |
| II | 105 (33.8%) | 45 (42.9%) | 60 (57.1%) | |
| III | 179 (57.6%) | 57 (31.8%) | 122 (68.2%) | |
Expression and survival analysis with TCGA, GTEx and GEO database
Data of expression (mRNA, normalized RNAseqV2 RSEM), copy number variation and mutation of GNB4 in GC were retrieved by using the cBioPortal, GEPIA2 and UALCAN, online cancer genomics tools providing visualization, analysis and download of large-scale cancer genomics datasets. Survival analysis were performed to analyze the association between GNB4 mRNA expression level and long-term OS of GC patients with primary data including clinicopathological parameters and status of H. pylori infection retrieved from TCGA database by cBioPortal.
Pathology
Upon the completion of radical gastrectomy, the fresh specimen was fixed in 10% neutralized formalin followed by paraffin embedding and eosin (H&E) staining for pathological diagnosis. The pathological staging for GC was based on (I) depth of tumor invasion (T), (II) tumor spreading to lymph node tumor invasion (N) and (III) tumor spreading to distant metastasis (M), according to the 8th edition of The American Joint Committee on Cancer (AJCC). Moreover, Lauren’s classification was also recorded to define the gross histological architecture.
Immunohistochemistry
For histological and immunohistochemical analysis, the above mentioned fixed and embedded tumor tissue were made into tissue microarray (TMA). The tissues underwent processes including deparaffinization, hydration and heat-induced epitope retrieval. Next, the sections were incubated with H2O2 for 10 minutes, followed by three times of phosphate-buffered saline (PBS) washing and 30 minutes blocking by blocking solution (HistostainTM kit, Zymed Company, CA, USA). Next, sections were incubated with primary antibodies (30–60 minutes, room temperature) and then with secondary antibody for (10–30 minutes, room temperature in dark), followed by the incubation with avidin-alkaline phosphatase (7 minutes at 40 °C). Then the sections were incubated with the red chromogen at 40 °C for visualization and further counterstained with application of Mayer hematoxylin method. The antibody of GNB4 used in this study is diluted 1:100 (Rabbit polyclonal to GNB4, anti-GNB4 antibody ab223113, Abcam, USA).
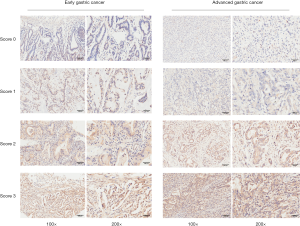
TMA staining was evaluated and categorized as follows: little observation of weak-to-strong complete cell membrane and/or cytoplasm staining in tumor cells as score 0; weak-to-strong complete cell membrane and/or cytoplasm staining in <10% of tumor cells as score 1; weak-to-strong complete cell membrane and/or cytoplasm staining in 10–50% of tumor cells as score 2; strong complete cell membrane and/or cytoplasm staining in ≥50% of tumor cells as score 3. Score 0–1 was recognized as low expression, score of 2–3 was recognized as high expression for subsequent statistical analyses. The evaluation and scoring process was performed by 2 independent pathologists, who were blinded to clinical outcomes.
Statistics
The association of GNB4 expression with clinicopathological parameters of EGC and AGC patients was evaluated by Pearman chi-squared test. Kaplan-Meier survival curves were plotted to perform survival analysis of 5-year overall survival (OS) and comparison of survival rates between groups was further analyzed by long-rank test. OS was defined as the time from the date of surgery to death from any cause or the date of last follow-up. Cases of loss to follow-up were recorded. To determine independent prognostic factors of long-term outcome of GC patients including the subgroup of EGC and AGC patients, univariate and multivariate analysis were conducted using Cox proportional hazards regression model. All statistical analyses were performed two-sided at a significance level of P=0.05 with the application of SPSS 23.0 software (SPSS Inc., Chicago, IL, USA).
Results
We firstly analyzed the copy number variation (CNV), genetic mutation and mRNA expression of GNB4 as well as the potential association between its expression and long-term survival of GC patients with TCGA and GTEx databases. Next, we performed multiple statistical analyses for H. pylori-infected GC patients receiving treatment in Ruijin hospital affiliated to Shanghai Jiaotong University School of Medicine in this study. Patients with initial diagnosis of distant metastasis such as liver, lung, ovarian, uterus and peritoneum metastasis as well as patients with history of neoadjuvant chemotherapy prior to surgery were excluded from this study. Of all 448 eligible patients, 137 (30.6%) EGC patients underwent D2 gastrectomy and the other 311 (69.4%) AGC patients received D2 gastrectomy followed by fluorouracil-based chemotherapy. After a median follow-up time of 52 (range, 3–77) months, death of 6 EGC patients and 83 AGC patients and loss to follow-up of 30 EGC patients and 31 AGC patients were recorded.
Expression analysis of GNB4 from online databases and our collected surgical samples
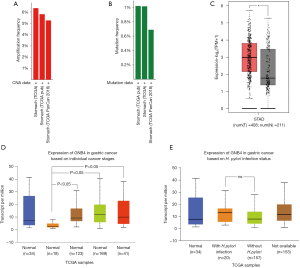
To understand the expression status of GNB4 in GC patients, especially those who were infected with H. pylori, we firstly profiled copy number alteration and genetic mutation of GNB4 with TCGA database using cBioPortal (https://www.cbioportal.org) and demonstrated that genetic amplification and mutation frequency of GNB4 was up to 6% and 1% in GC patients, respectively (Figure 1A,B) (23). Then we performed GNB4 mRNA expression analysis with TCGA and GTEx databases using web-based tools including GEPIA2 (http://gepia2.cancer-pku.cn/#index) and UALCAN (http://ualcan.path.uab.edu/index.html) to analyze cancer-OMICS data. GEPIA2 analysis showed that GNB4 expression was significantly higher in GC tissues compared with normal gastric tissues (P<0.01) (Figure 1C ). Stage-based and H. pylori-infection status-based subgroup analysis enabled by UALCAN revealed that GNB4 expression was significantly lower in GC patients at stage I than other more advanced stages (Figure 1D). Moreover, it was observed that mRNA expression of GNB4 was higher in H. pylori-infected GC patients compared to those without H. pylori-infection, though not significantly (Figure 1E) (24).
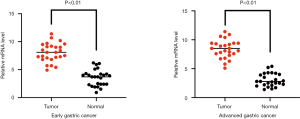
To further validate if GNB4 was differentially expressed between tumor and non-tumor tissue of GC patients with H. pylori infection, we firstly performed quantitative real time-PCR (qRT-PCR) to examine GNB4 mRNA expression level in 50 paired cancerous and non-cancerous tissues of H. pylori-positive GC patients including 25 EGC and 25 AGC patients. As shown in Figure 2, GNB4 mRNA was significantly higher in cancerous tissues of both EGC and AGC compared to their matching controls. To further clarify GNB4 expression status in H. pylori-positive GC tissues, we performed IHC study of GNB4 to evaluate its expression in the surgical samples of 137 EGC and 311 AGC patients with H. pylori infection. Figure 3 demonstrates the representative immunostaining of GNB4 scored as 0, 1, 2 and 3.
Survival analysis of GNB4 from TCGA and GTEx databases
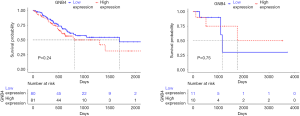
Previous study conducted bioinformatic analysis for GSE62254 dataset from GTEx database and identified high expression of GNB4 as an independent unfavorable prognostic factor for H. pylori-induced GC patients (P=0.031) (22). To validate their findings and further investigate the clinical significance of GNB4 overexpression in GC, we conducted survival analysis of GNB4 expression with TCGA cohort. Survival analysis of a total number of 161 GC patients without H. pylori infection revealed no significant association between GNB4 expression and OS of these patients (P=0.24) (Figure 4). Regarding GC patient with H. pylori infection analysis validity was limited by the low number of patients (n=21) (Figure 4), addressing the importance of further investigation on large-scale number of GC patients with H.pylori infection.
Associations of GNB4 expression with clinicopathological characteristics and 5-year OS of H. pylori-positive EGC patients
The clinical parameters and their associations with GNB4 expression of H. pylori-positive EGC patients were detailed in Table 1. There was no statistical association between GNB4 expression and any clinicopathological parameter. In the survival analysis using Kaplan-Meier method, the association between GNB4 expression and long-term survival of H. pylori-positive EGC patients (survival duration ranging from 15 to 77 months) was insignificant (P=0.982). Both univariate and multivariate Cox proportional hazards model analyses demonstrated that tumor invasion of lymph node (P=0.048, HR 7.25 and 95% CI: 1.271–101.742, multivariate analysis) and pathological TNM staging (P=0.047, HR 13.58 and 95% CI: 1.03–178.95, multivariate analysis) are independent prognostic factors of long-term outcome of EGC patients with H. pylori infection (Table 3).
Table 3
| Variable | Univariate analysis | Multivariate analysis | |||||
|---|---|---|---|---|---|---|---|
| 95% CI | HR | P value | 95% CI | HR | P value | ||
| Age | |||||||
| <60 | 1.0 | 1.0 | |||||
| ≥60 | 0.35–10.39 | 1.90 | 0.457 | 0.34–16.44 | 2.37 | 0.383 | |
| Gender | |||||||
| Male | 1.0 | 1.0 | |||||
| Female | 0.36–8.87 | 1.79 | 0.476 | 0.41–22.29 | 3.01 | 0.281 | |
| Differentiation | |||||||
| Low | 1.0 | 1.0 | |||||
| Medium/high | 0.04–3.14 | 0.37 | 0.359 | 0.05–5.22 | 0.49 | 0.553 | |
| Lauren’s classification | |||||||
| Diffuse | 1.0 | 1.0 | |||||
| Intestinal | 0.31–22.37 | 2.61 | 0.381 | 0.25–46.83 | 3.42 | 0.357 | |
| Tumor site | |||||||
| Cardia | 1.0 | 1.0 | |||||
| Non-cardia | 0.04–2.75 | 0.32 | 0.300 | 0.01–2.96 | 1.33 | 0.273 | |
| Lymph node invasion | |||||||
| Negative | 1.0 | 1.0 | |||||
| Positive | 1.65–120.86 | 14.11 | 0.016* | 1.27–101.74 | 7.25 | 0.048* | |
| TNM staging (II–III) | |||||||
| I | 1.0 | 1.0 | |||||
| II+III | 3.86–116.03 | 21.16 | 0.000* | 1.03–178.95 | 13.58 | 0.047* | |
| GNB4 expression | |||||||
| Low expression | 1.0 | 1.0 | |||||
| High expression | 0.19–5.57 | 1.02 | 0.982 | 0.16–5.71 | 0.95 | 0.955 | |
*, statistical significant (P<0.05).
Associations between GNB4 expression and clinicopathological characteristics of H. pylori-positive AGC patients
Regarding AGC patients with H. pylori infection, GNB4 expression was not correlated with clinical parameters including patient age, gender, tumor differentiation, Lauren’s classification, tumor site, tumor invasion depth and tumor invasion of lymph node. Notably however, significant association was observed between GNB4 expression and pathological stage of H. pylori-positive AGC patients (P=0.047, HR 0.53 and 95% CI: 0.31–0.90) (Table 2).
Expression of GNB4 and 5-year OS of H. pylori-positive AGC patients
Survival analysis for H. pylori-positive AGC patients (survival duration ranging from 3 to 77 months) was performed to investigate the potential association between GNB4 expression and long-term outcome of these patients. Remarkably, Kaplan-Meier plot demonstrated that high GNB4 expression correlated with a worse long-term OS of AGC patients (P<0.01) (Figure 5A). We further performed subgroup analysis based on the patient’s pathological stage. Among all 311 H. pylori-positive AGC patients, the number of patients at stage I, II and III was 27, 105 and 179, respectively. Stage-based survival analyses further demonstrated that GNB4 expression was significantly associated with 5-year OS of stage III (P=0.043) rather than stage I (P=0.932) or stage II (P=0.05) AGC patients (Figure 5B,C,D). Moreover, univariate and multivariate Cox proportional hazards model analyses were conducted to evaluate the prognostic value of GNB4 expression in H. pylori-positive AGC patients. As shown in univariate analysis, tumor invasion depth (P=0.001), lymph node metastasis (P<0.001), pathological stage (P<0.001) as well as GNB4 high expression (P=0.002) were significantly correlated with 5-year OS of H. pylori-positive AGC patients; Multivariate analysis further identified that both lymph node metastasis (P=0.013) and GNB4 high expression (P=0.020) were significantly correlated with 5-year OS of these patients (Table 4), indicating GNB4 high expression as an independent prognostic factor of long-term survival of AGC patients infected with H. pylori.
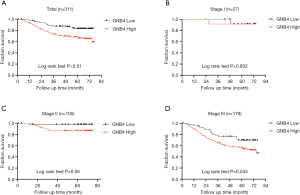
Table 4
| Variable | Univariate analysis | Multivariate analysis | |||||
|---|---|---|---|---|---|---|---|
| 95% CI | HR | P value | 95% CI | HR | P value | ||
| Age (years) | |||||||
| <60 | 1.0 | 1.0 | |||||
| ≥60 | 0.64–1.55 | 1.0 | 0.985 | 0.71–1.81 | 1.13 | 0.607 | |
| Gender | |||||||
| Male | 1.0 | 1.0 | |||||
| Female | 0.86–2.18 | 1.37 | 0.192 | 0.80–2.15 | 1.31 | 0.284 | |
| Differentiation | |||||||
| Low | 1.0 | 1.0 | |||||
| Medium/High | 0.33–1.03 | 0.58 | 0.063 | 0.47–1.57 | 0.86 | 0.617 | |
| Lauren’s classification | |||||||
| Diffuse | 1.0 | 1.0 | |||||
| Intestinal | 0.48–1.25 | 0.77 | 0.291 | 0.53–1.45 | 0.61 | 0.877 | |
| Tumor site | |||||||
| Cardia | 1.0 | 1.0 | |||||
| Non-cardia | 0.57–1.48 | 0.91 | 0.714 | 0.47–1.27 | 0.77 | 0.309 | |
| Tumor invasion | 0.001* | 0.194 | |||||
| T2 | 1.0 | 1.0 | |||||
| T3 | 0.81–6.04 | 2.21 | 0.121 | 0.21–4.55 | 0.98 | 0.981 | |
| T4 | 1.71–10.57 | 4.25 | 0.002* | 0.35–7.96 | 1.67 | 0.520 | |
| Lymph node invasion | 0.000* | 0.013* | |||||
| N0 | 1.0 | 1.0 | |||||
| N1 | 1.26–8.56 | 3.29 | 0.015* | 0.65–11.62 | 2.76 | 0.167 | |
| N2 | 1.82–11.30 | 4.54 | 0.001* | 0.66–14.30 | 3.08 | 0.152 | |
| N3 | 4.25–23.48 | 9.98 | 0.000* | 1.31–26.54 | 5.89 | 0.021* | |
| TNM staging (II–III) | 0.000* | 0.380 | |||||
| I | 1.0 | 1.0 | |||||
| II | 0.26–5.48 | 1.18 | 0.830 | 0.05–4.73 | 0.50 | 0.545 | |
| III | 1.61–26.75 | 6.56 | 0.009* | 0.05–16.66 | 0.88 | 0.934 | |
| GNB4 expression | |||||||
| Low expression | 1.0 | 1.0 | |||||
| High expression | 0.26–0.74 | 0.44 | 0.002* | 0.31–0.90 | 0.53 | 0.020* | |
*, statistical significant (P<0.05).
Discussion
In the present study, we aimed to investigate the potential association between GNB4 expression and clinicopathological features of H. pylori-positive GC patients. Both bioinformatic analysis from online dataset and statistical analysis from our own cohorts demonstrated that high level of GNB4 expression correlated with pathological stage as well as poor prognosis of H. pylori-positive AGC patients, indicating the potential impact of GNB4 in the progression of GC induced by H. pylori.
H. pylori infection plays key roles in the initiation and development of GC (25). Eradication of H. pylori reduces incidence of GC and complementarily improves the prognosis of H. pylori-positive GC patients though curative surgery remains the major treatment of this lethal malignancy (26). Although molecular mechanism underlying H. pylori-induced cancer development has not been fully clarified, several studies showed that both the hypermethylation of tumor suppressor genes and the hypomethylation of oncogene induced by H. pylori infection significantly influenced the process of GC initiation and development (27,28). Liu et al. comprehensively analyzed datasets of four gene methylation profiling microarrays and one gene expression profiling microarray of H. pylori-induced GC patients and identified GNB4 as one of both the most differentially hypomethylated and overexpressed genes. Both the hypomethylation and high expression of GNB4 were significantly correlated with a worse OS of H. pylori-induced GC patients, indicating the potential clinical and biological significance of GNB4 in this group of GC patients (22). Considering the significance of oncogene overexpression which could be induced by hypomethylation in the process of carcinogenesis and progression, it is speculated that the H. pylori infection induced hypomethylation of GNB4 which leads to its overexpression, which partially underlies our findings that high expression of GNB4 is significantly correlated with more advanced pathological stage as well as lower overall survival of AGC patients. On the other hand, studies repeatedly showed that H. pylori contributed to GC progression via aberrant activation of key signaling pathways such as mitogen-activated protein kinase (MAPK) cascade (29). For instance, Liu et al investigated the role of key effectors in MAPK pathway in the progression of H. pylori-induced GC. They not only observed the correlation between H.pylori infection and MAPK expression but also proved that H. pylori promoted GC aggressiveness by up-regulating the expression of Heparanase, a key modulator of issue remodeling and cell migration, via the activation of MAPK cascade (30). Ding et al. demonstrated that specific inhibition of key effectors of MAPK pathway (such as Mek1/2, p38, JNK and etc.) increased H. pylori-induced cell cycle G1 arrest, enhanced cell apoptosis and decreased cell proliferation (31). Notably, in breast cancer high expression of GNB4 is suggested to promote the proliferation, migration as well as the chemoresistance of tumor cell via activation of the Erk1/2, a key effector in the MAPK signaling pathway (18). Although the role of GNB4 in GC progression remains to be clarified, these results suggest that overexpression of GNB4 modulated by H. pylori may be involved in GC development, reinforcing our findings of the clinical significance of GNB4 in H. pylori-positive AGC patients.
G-protein (also known as guanine nucleotide-binding protein) is a heterotrimer composed of Gα, Gβ and Gγ subunits. This heterotrimeric protein plays crucial role in signal transduction between epithelial receptors to intracellular effectors (32): upstream signals trigger G protein conformational change and activate Gα by exchanging its bound GDP to GTP. Activated Gα dissociates from the other two subunits which then form the Gβγ dimmer. Both the activated GTP-bound Gα and Gβγ dimmer stimulate distinctive downstream cascades such as mitogen-activated protein kinase (MAPK) and PI3k-Akt pathways which essentially regulate normal and cancerous cell survival, proliferation, apoptosis and cell cycle, etc. (33). Studies of downstream signaling of Gβγ dimer demonstrate its regulation of multiple key effectors including GIRK1/2, PLCβ1/2/3, Raf-1 and Ras exchange factor (34). Of note, it is shown that H. pylori stimulates VEGFA expression to promote GC progression and MAPK-mediated activation of key transcription factors SP1 and SP2 underlies this process (35).
GNB4, the encoding gene of Gβ subunit 4, is one of the five Gβ-encoding genes which is recognized as one major contributor to the progression of multiple malignancies: Wang et al. showed that overexpression of GNB4 in two chemoresistant breast cancer cell lines significantly upregulated the activities of key effectors controlling cell cycle and apoptosis whereas DNA methyltransferase 3B (DNMT3B)-mediated epigenetic silencing of GNB4 expression inhibited cell proliferation and induced cell cycle arrest (18). Umar et al. further explored the potential clinical significance of GNB4 expression in breast cancer with the application of nano-LC-FTICR mass spectrometry. They performed comparative proteomics on laser capture microdissection-derived breast tumor cells and identified GNB4 as a biomarker associated with tamoxifen therapy resistance in recurrent breast cancer (19). Moreover, Riemann et al. performed comprehensive screening of the promoter, intron 1 and exon 1 of GNB4 for single nucleotide polymorphisms (SNPs) and characterized distinct haplotype blocks in urothelial bladder carcinoma and colorectal carcinoma, respectively. Notably, intron-1 haplotype block 1 of GNB4 independently predicted the progression and survival of patients with urothelial bladder cancer (20), whereas in colorectal carcinoma it was block 2 of the same haplotype of GNB4 that significantly correlated with the long-term survival of patients at pathological III and IV stage of CRC (21). With respect to the role of GNB4 in GC, the identification of GNB4 hypomethylation and overexpression as independent predictors of poor prognosis of H. pylori-induced GC patients inspired us to further investigate its clinical significance via immunostaining-based strategy. In the present study, we firstly conducted GNB4 expression and survival analysis with TCGA and GTEx datasets. GNB4 expression in GC patients at pathological stage I was significantly lower than patients at more advanced stages in UALCAN, which was in line with our findings that GNB4 expression was significantly correlated with H. pylori-positive GC patients at advanced stage rather than early stage. To further validate if high mRNA expression of GNB4 independently predicted the survival rate of GC patients with and without H. pylori infection, we retrieved clinical data from TCGA cohort and performed survival analysis for 161 GC patients without and 21 GC patients with H. pylori infection. However, the limited number of H. pylori-positive GC patient restricted the quality of clinical-relevant statistical analysis, addressing the necessity of GNB4 immunostaining on large number of surgical specimens from GC patients with their H. pylori infection status validated. As shown above, our statistical analysis not only revealed the significant correlation between GNB4 overexpression and pathological stage of AGC patients but also identified GNB4 high expression as an independent unfavorable prognostic factor for H. pylori-positive AGC patients. On the contrary, little clinical significance of GNB4 expression was observed in GC patients at early stage, which might be explained by the relatively limited number of EGC patients enrolled in the study. Moreover, as we merely focused on the H. pylori-positive GC patients, no investigation was conducted to clarify the clinical significance of GNB4 expression in GC patients without H. pylori infection. These limitations highlight the importance of further large-scale study with H. pylori-positive and -negative patients included.
In conclusion, we have identified high expression of GNB4 as a potentially significant biomarker for the poor prognosis in AGC rather than EGC patients infected with H. pylori. Further studies are required to investigate the potential influence of GNB4 and its possible underlying mechanisms in H. pylori-positive GC progression.
Acknowledgments
We thank the Shanghai Institute of Digestive Surgery in Ruijin hospital affiliated to Shanghai Jiaotong University School of Medicine for their generous help.
Funding: This work was supported by
Footnote
Reporting Checklist: The authors have completed the STROBE reporting checklist. Available at: http://dx.doi.org/10.21037/tcr-19-2914
Data Sharing Statement: Available at http://dx.doi.org/10.21037/tcr-19-2914
Conflicts of Interest: All authors have completed the ICMJE uniform disclosure form (available at http://dx.doi.org/10.21037/tcr-19-2914). The authors have no conflicts of interest to declare.
Ethical Statement: The authors are accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved. The study was conducted in accordance with the Declaration of Helsinki (as revised in 2013). The study was approved by the local ethics committee of Ruijin hospital affiliated to Shanghai Jiaotong University School of Medicine [(2018)Linlunshen No. 151] and informed consent was taken from all the patients.
Data Availability Statement: The Gene Expression Omnibus (GEO) data and TCGA data referenced during the study are available in a public repository from the GEO website (https://www.ncbi.nlm.nih.gov/geo/), GEPIA website (
Open Access Statement: This is an Open Access article distributed in accordance with the Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License (CC BY-NC-ND 4.0), which permits the non-commercial replication and distribution of the article with the strict proviso that no changes or edits are made and the original work is properly cited (including links to both the formal publication through the relevant DOI and the license). See: https://creativecommons.org/licenses/by-nc-nd/4.0/.
References
- Van Cutsem E, Sagaert X, Topal B, et al. Gastric cancer. Lancet 2016;388:2654-64. [Crossref] [PubMed]
- Kim HJ, Oh SC. Novel Systemic Therapies for Advanced Gastric Cancer. J Gastric Cancer 2018;18:1-19. [Crossref] [PubMed]
- Hamashima C. Current issues and future perspectives of gastric cancer screening. World J Gastroenterol 2014;20:13767-74. [Crossref] [PubMed]
- Bray F, Ferlay J, Soerjomataram I, et al. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin 2018;68:394-424. [Crossref] [PubMed]
- Wroblewski LE, Peek RM Jr, Wilson KT. Helicobacter pylori and gastric cancer: factors that modulate disease risk. Clin Microbiol Rev 2010;23:713-39. [Crossref] [PubMed]
- Camilo V, Sugiyama T, Touati E. Pathogenesis of Helicobacter pylori infection. Helicobacter 2017;22 Suppl 1.
- Amieva M, Peek RM Jr. Pathobiology of Helicobacter pylori-Induced Gastric Cancer. Gastroenterology 2016;150:64-78. [Crossref] [PubMed]
- Su YL, Huang HL, Huang BS, et al. Combination of OipA, BabA, and SabA as candidate biomarkers for predicting Helicobacter pylori-related gastric cancer. Sci Rep 2016;6:36442. [Crossref] [PubMed]
- Zou D, Xu L, Li H, et al. Role of abnormal microRNA expression in Helicobacter pylori associated gastric cancer. Crit Rev Microbiol 2019;45:239-51. [Crossref] [PubMed]
- Lee SW, Park KC, Kim JG, et al. Dysregulation of MicroRNA-196b-5p and MicroRNA-375 in Gastric Cancer. J Gastric Cancer 2016;16:221-9. [Crossref] [PubMed]
- Taghizadeh S, Sankian M, Ajami A, et al. Expression levels of vascular endothelial growth factors a and C in patients with peptic ulcers and gastric cancer. J Gastric Cancer 2014;14:196-203. [Crossref] [PubMed]
- Kataoka Y, Okabe H, Yoshizawa A, et al. HER2 expression and its clinicopathological features in resectable gastric cancer. Gastric Cancer 2013;16:84-93. [Crossref] [PubMed]
- Bao W, Fu H J, Xie Q S, et al. HER2 interacts with CD44 to up-regulate CXCR4 via epigenetic silencing of microRNA-139 in gastric cancer cells. Gastroenterology 2011;141:2076-87.e6. [Crossref] [PubMed]
- Tsamandas AC, Kardamakis D, Tsiamalos P, et al. The potential role of Bcl-2 expression, apoptosis and cell proliferation (Ki-67 expression) in cases of gastric carcinoma and correlation with classic prognostic factors and patient outcome. Anticancer Res 2009;29:703-9. [PubMed]
- Li T, Chen J, Liu QL, et al. Meta-analysis: E-cadherin immunoexpression as a potential prognosis biomarker related to gastric cancer metastasis in Asian patients. Eur Rev Med Pharmacol Sci 2014;18:2693-703. [PubMed]
- Lieto E, Ferraraccio F, Orditura M, et al. Expression of vascular endothelial growth factor (VEGF) and epidermal growth factor receptor (EGFR) is an independent prognostic indicator of worse outcome in gastric cancer patients. Ann Surg Oncol 2008;15:69-79. [Crossref] [PubMed]
- Hurowitz EH, Melnyk JM, Chen YJ, et al. Genomic characterization of the human heterotrimeric G protein alpha, beta, and gamma subunit genes. DNA Res 2000;7:111-20. [Crossref] [PubMed]
- Wang B, Li D, Rodriguez-Juarez R, et al. A suppressive role of guanine nucleotide-binding protein subunit beta-4 inhibited by DNA methylation in the growth of anti-estrogen resistant breast cancer cells. BMC Cancer 2018;18:817. [Crossref] [PubMed]
- Umar A, Kang H, Timmermans A M, et al. Identification of a putative protein profile associated with tamoxifen therapy resistance in breast cancer. Mol Cell Proteomics 2009;8:1278-94. [Crossref] [PubMed]
- Riemann K, Struwe H, Eisenhardt A, et al. Characterization of intron-1 haplotypes of the G protein beta 4 subunit gene--association with survival and progression in patients with urothelial bladder carcinoma. Pharmacogenet Genomics 2008;18:999-1008. [Crossref] [PubMed]
- Riemann K, Struwe H, Alakus H, et al. Association of GNB4 intron-1 haplotypes with survival in patients with UICC stage III and IV colorectal carcinoma. Anticancer Res 2009;29:1271-4. [PubMed]
- Liu D, Ma X, Yang F, et al. Discovery and validation of methylated-differentially expressed genes in Helicobacter pylori-induced gastric cancer. Cancer Gene Ther 2020;27:473-85. [Crossref] [PubMed]
- Gao J, Aksoy B A, Dogrusoz U, et al. Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci Signal 2013;6:pl1. [Crossref] [PubMed]
- Chandrashekar DS, Bashel B, Balasubramanya SAH, et al. UALCAN: A Portal for Facilitating Tumor Subgroup Gene Expression and Survival Analyses. Neoplasia 2017;19:649-58. [Crossref] [PubMed]
- Cooke CL, Torres J, Solnick JV. Biomarkers of Helicobacter pylori-associated gastric cancer. Gut Microbes 2013;4:532-40. [Crossref] [PubMed]
- Take S, Mizuno M, Ishiki K, et al. The long-term risk of gastric cancer after the successful eradication of Helicobacter pylori. J Gastroenterol 2011;46:318-24. [Crossref] [PubMed]
- Compare D, Rocco A, Liguori E, et al. Global DNA hypomethylation is an early event in Helicobacter pylori-related gastric carcinogenesis. J Clin Pathol 2011;64:677-82. [Crossref] [PubMed]
- Maeda M, Moro H, Ushijima T. Mechanisms for the induction of gastric cancer by Helicobacter pylori infection: aberrant DNA methylation pathway. Gastric Cancer 2017;20:8-15. [Crossref] [PubMed]
- Polk DB, Peek RM. Helicobacter pylori: gastric cancer and beyond. Nature Reviews Cancer 2010;10:403-14. [Crossref] [PubMed]
- Liu LP, Sheng XP, Shuai TK, et al. Helicobacter pylori promotes invasion and metastasis of gastric cancer by enhancing heparanase expression. World J Gastroenterol 2018;24:4565-77. [Crossref] [PubMed]
- Ding SZ, Smith MF Jr, Goldberg JB. Helicobacter pylori and mitogen-activated protein kinases regulate the cell cycle, proliferation and apoptosis in gastric epithelial cells. J Gastroenterol Hepatol 2008;23:e67-78. [Crossref] [PubMed]
- Rens-Domiano S, Hamm HE. Structural and functional relationships of heterotrimeric G-proteins. FASEB J 1995;9:1059-66. [Crossref] [PubMed]
- Goldsmith ZG, Dhanasekaran DN. G protein regulation of MAPK networks. Oncogene 2007;26:3122-42. [Crossref] [PubMed]
- Smrcka AV. Molecular targeting of Galpha and Gbetagamma subunits: a potential approach for cancer therapeutics. Trends Pharmacol Sci 2013;34:290-8. [Crossref] [PubMed]
- Strowski MZ, Cramer T, Schafer G, et al. Helicobacter pylori stimulates host vascular endothelial growth factor-A (vegf-A) gene expression via MEK/ERK-dependent activation of Sp1 and Sp3. FASEB J 2004;18:218-20. [Crossref] [PubMed]


