
Chronic nicotine exposure affects programmed death-ligand 1 expression and sensitivity to epidermal growth factor receptor-tyrosine kinase inhibitor in lung cancer
Introduction
Somatic mutation of the epidermal growth factor receptor (EGFR) gene is a major oncogenic driver in non-small cell lung cancer (NSCLC) (1). Patients with advanced NSCLC harboring EGFR mutation can receive EGFR-tyrosine kinase inhibitors (TKIs) such as gefitinib, erlotinib, and afatinib with high response rates for first-line treatment (2). However, acquired resistance to EGFR-TKIs in metastatic setting is inevitable, although initial responses are dramatic. In addition, ~20–30% of patients have no good initial clinical response to EGFR-TKIs, although they harbor sensitizing EGFR mutations (3). These primary resistant mechanisms can be explained by de novo resistant mutation, suboptimal drug exposure, failure of apoptosis induction, and other somatic alterations on genes encoding components of major signaling pathways (4). After tumor progression, novel third-generation EGFR-TKIs such as osimertinib and olmutinib have been designed to overcome resistance due to T790M mutation (5,6). Other strategies to overcome different mechanisms are also under development (7,8).
Although EGFR mutations are known to be commonly detected in never or light smokers (9), more than 30% of patients with EGFR mutations are currently smoking at the time of diagnosis according to Japanese data (10). Additionally, Kim et al. (11) have shown that about 40% of ever smokers with EGFR mutant adenocarcinoma have smoking histories of more than 30 pack-years. These smoking histories are independently associated with poor response to EGFR-TKIs in terms of progression-free survival in patients with activating EGFR mutations (10-13). Poor clinical outcomes are mainly due to cross-talks between nicotine-induced nicotinic acetylcholine receptor (nAChR) and EGFR pathway, Src activation, and epithelial mesenchymal transition (EMT) both in vitro and in vivo (14-16).
Recently, immunotherapies based on blocking of immunosuppressive checkpoints have been established as major treatment paradigms for the management of NSCLC. Programmed cell death-1 protein receptor (PD-1) and ligand (PD-L1) pathway is involved in suppressing autoimmunity during T-cell activation, allowing for immune tolerance of PD-L1 expressed cells (17). Multiple solid tumors can adapt these mechanisms and overexpress PD-L1, thereby avoiding immunologic surveillance and promoting cancer growth (17). Given the mechanism of action of anti-PD-1/PD-L1 antibodies, favorable responses have been observed in patients whose tumors express PD-L1 (18,19). Smoking history also has clinical significance for patients with PD-1/PD-L1 blockade (20,21) which is influenced by specific molecular determinants, including somatic mutational burden and genomic instability (22). However, in real practice, activating EGFR mutation is associated with poor clinical response to anti-PD-1/PD-L1 therapy (23). This is explained by low tumoral PD-L1 expression and CD8+ tumor-infiltrating lymphocytes within the tumor microenvironment (24).
Immunotherapies are not routinely considered for the management of patients with activating EGFR mutations (25). Nevertheless, when we consider a plurality of smokers in these population and favorable effect of smoking history on response to checkpoint inhibitors, the biologic influence of nicotine needs to be investigated clinically. Thus, the aim of the present study was to determine the effect of nicotine exposure on PD-L1 expression in EGFR mutant lung cancer cells. Moreover, sensitivity of these cells to EGFR-TKIs and biologic changes regarding EGFR-related pathways upon nicotine exposure were also evaluated.
Methods
Cell culture
Human NSCLC cell line PC9 (EGFR exon 19 deletion) was purchased from Korean Cell Line Bank (Seoul, Korea). Cells were cultured in RPMI-1640 medium (WelGENE, Daegu, Korea) supplemented with 10% fetal bovine serum and 1% penicillin and streptomycin solution (10,000 units/ml penicillin and 10 mg/ml streptomycin) at 37 °C in 5% CO2. They were split regularly before they attained approximately 80% confluence. PC9 cells maintained in the presence of 1 µM nicotine (Sigma-Aldrich, St. Louis, MO, USA) for three months were designated as PC9/N.
Reagents
Gefitinib was purchased from Cell Signaling Technology (Beverly, MA, USA). It was dissolved in DMSO to obtain a stock solution at 100 mM. The final concentration of DMSO in all conditioned media did not exceed 0.1%. Antibodies specific for EGFR, mTOR, AKT, PD-L1, α1-nAchR, β-actin, and horseradish peroxidase-conjugated secondary antibodies were purchased from Cell Signaling Technology (Beverly, MA, USA). Nicotinic acetyl-choline receptor α1 subunit (α1-nAchR) was obtained from Abcam (Cambridge, United Kingdom). To evaluate the influence of gefitinib on EGFR related signaling pathway, cells were stimulated with gefitinib for 48 hrs.
Cell morphological observation
Cells were seeded into 6-well plate at density of 3×105 cells/well. Following 24 hrs of incubation in room air, cells were treated with gefitinib (0, 0.1, or 1 µM) for 48 hrs. They were then stained with Wright and Giemsa. Morphological changes were observed under a phase contrast inverted microscope (original magnification: 200×).
Cell viability assay
Cell viability assay was performed based on the conversion of MTT [3-(4,5-dimethythiazol-2-yl)-2,5-diphenyltetrazolium bromide] to formazan via mitochondrial oxidation. Briefly, 1×104 cells were cultured in 96-well plates (200 µL medium/well). Cells were then treated with gefitinib (0, 0.01, 0.1, 1, 10, and 50 µM) for 48 hrs. MTT solution was added to each well of the plate and incubated in room air for 4 hrs at 37 °C. Formazan crystals were dissolved in acidified and OD value was measured at wavelength of 562 nm.
Reverse transcription (RT)-PCR analysis
To quantify PD-L1 and α1-nAchR mRNA expression levels, total RNAs were extracted from cells using TRIzol reagent™ (Invitrogen, Carlsbad, CA, USA) and cDNA was synthesized using commercially available RT reagent kit (Invitrogen) according to the manufacturer’s instructions. PCR was carried out in reaction mixture containing Taq DNA polymerase, primers, and cDNA. Equal volumes of PCR products were separated by electrophoresis on 2% agarose gels. β-actin was used as an internal control. Quantitative real time PCR (qRT-PCR) was performed using a Rotor-Gene SYBR® Green qPCR Kit Master Mix (Qiagen, Valencia, CA, USA) and carried out in a Real Time PCR Rotor-Gene Q Machine (Qiagen). All experiments were performed in triplicates. The relative expression of PD-L1 was normalized to β-actin expression. Fold change of test gene mRNA expression was determined using the 2-∆∆Ct method (∆Ct = difference in threshold cycles for the test gene, ∆∆Ct = difference in ∆Ct between the control and treatment group with nicotine or gefitinib).
Immunofluorescence
Cells were treated with 0.1 µM gefitinib for 48 hrs. Immunofluorescence staining was then performed according to the manufacturer’s protocol. Briefly, cells were washed with cold PBS, fixed with 4% paraformaldehyde, blocked with blocking buffer (PBS plus 0.1% Tween-20 and 3% goat serum), and incubated with primary antibodies (phospho-EGFR or PD-L1) at 4 °C overnight. After washing with PBS, cells were incubated with fluorochrome-conjugated secondary antibodies (Alexa Fluor 488 anti-rabbit IgG) for 1 hour at room temperature. Cell nuclei were stained with 4′,6-diamidino-2-phenylindole (DAPI) and then microscopy was performed using by LSM800 w/Airyscan confocal laser scanning microscope (Carl Zeiss, Germany).
Western blot analysis
Cells were harvested and suspended in RIPA lysis buffer (Thermo, Hercules, CA, USA) containing a mixture of protease inhibitor and phosphatase inhibitor (GenDEPOT, CA, USA) followed by centrifugation at 13,000 rpm for 30 min at 4 °C. Protein content of the supernatant was determined using a BCA kit (Thermo). Proteins (30 µg per well) were separated by 10% sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS-PAGE) using a 10% PAGE gel with a Tris-glycine-SDS buffer and transferred to polyvinylidene difluoride (PVDF) membranes (Bio-Rad, MA, USA). These membranes were blocked with 5% skim milk (Sigma, St. Louis, MO, USA) at room temperature for 2 hrs followed by incubation with primary antibodies diluted with 3% BSA in Tris buffered saline containing 0.1% Tween 20 (TBS-T) overnight at 4 °C. Membranes were then washed with TBS-T and incubated with HRP-conjugated secondary antibody at room temperature for 2 hrs. Signals were detected using Western Blotting Luminol Reagent (Santa Cruz Biotechnology, Inc., Santa Cruz, CA, USA).
Study population
We consecutively collected 54 patients with pathologically confirmed adenocarcinoma of lung and activating EGFR mutation at diagnosis between January 2016 and December 2017 from Seoul St. Mary’s Hospital, Incheon St. Mary’s Hospital, Bucheon St. Mary’s Hospital, St. Paul’s Hospital, and Uijeongbu St. Mary’s Hospital, Korea. Based on pack-year (PY) defined as average number of cigarettes per day/20× years of smoking, patients were categorized as never smokers (<100 cigarettes in their life-time), light smokers (0< PY <30), or heavy smokers (PY ≥30). Genotying of EGFR was done using peptide nucleic acid (PNA)-mediated PCR clamping method such as PNAClamp TM EGFR Mutation Detection Kit (PANAGENE, Inc., Daejeon, Korea) using real-time PCR. Activating EGFR mutation was defined as exon 19 deletion or exon 21 point mutation. PD-L1 immunohistochemistry testing was performing using PD-L1 clone 22C3 pharmDx kit and Dako automated Link 48 platform (Dako, Carpenteria, CA, USA). PD-L1 tumor proportion score (TPS) was calculated as the percentage of at least 100 viable tumor cells with complete or partial membrane staining.
Statistical analysis
The differences in the levels of cell viability between the two study groups in the presence of individual doses of gefitinib were compared using the Mann-Whitney test. The Fisher exact test was applied to compare the high PD-L1 TPS status (≥50%) among different burdens of smoking dosage. The cell viability levels were summarized using means and standard errors (SEM), and a P value <0.05 was used to define statistical significance. All analyses were performed using Graph-Pad Prism version 5.00 for Windows (GraphPad Software, CA, USA).
Results
Morphological changes and growth inhibition by chronic nicotine exposure in EGFR sensitive cells
PC9 cells were exposed to 1 µM nicotine for 3 months and then treated with gefitinib (0, 0.01, 0.1, 1, 10, and 50 µM) for 48 hrs. Figure 1 shows nicotine-induced cellular growth promotion and nonadherent clustered appearance on culture plate under a phase contrast inverted microscopy. Gefitinib treatment decreased cellular proliferation of PC9 and PC9/N cells in a concentration-dependent manner. After treatment with gefitinib, PC9 cells showed more pleomorphic appearance with atypical nuclei and more detached cells shrunken with condensed nuclei than PC9/N cells.
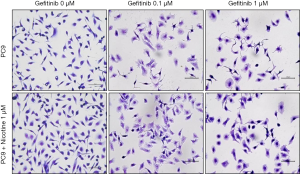
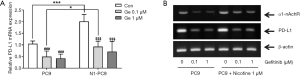
Figure 2 shows effects of gefitinib on the growth of PC9 and PC9/N cells determined by MTT assay. Gefitinib at concentration of 0.01 µM significantly suppressed the growth of PC9 cells (23.3%±3.0%, **P<0.01 compared to PC9 + gefitinib 0 µM), but not PC9/N cells. There was a statistical difference in cell viability between the two cells at concentration of 0.01 µM of gefitinib (P<0.05). DMSO treated cells showed no significant effect on cell morphology or viability (data not shown). Cellular growth was inhibited by treatment with gefitinib in a concentration-dependent manner. Gefitinib significantly suppressed the growth of both cell lines consistent with morphological changes, however, PC9 cells were significantly more sensitive to gefitinib than PC9/N cells.
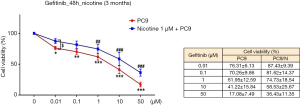

Effects on PD-L1 and nAchR gene expression by chronic nicotine exposure after treatment with gefitinib in EGFR sensitive cells
mRNA expression levels of PD-L1 and α1-nAchR in cells after treatment with gefitinib were measured by PCR (Figure 3). Quantitative reverse transcription (qRT)-PCR results showed that the relative gene mRNA expression of PD-L1 in PC9 and PC9/N cells was significantly decreased after treatment with gefitinib at 0.1 or 1 µM. The expression level of PD-L1 mRNA tended to be slightly higher in PC9/N cells compared to that in PC9 cells after treatment with gefitinib (0.1 or 1 µM) (Figure 3A). Reverse transcription (RT)-PCR results also revealed that α1-nAchR and PD-L1 mRNA expression levels were increased by nicotine exposure but decreased after treatment with gefitinib (Figure 3B). Similar results in their protein expression levels were also observed by Western blot analysis (Figure 4).
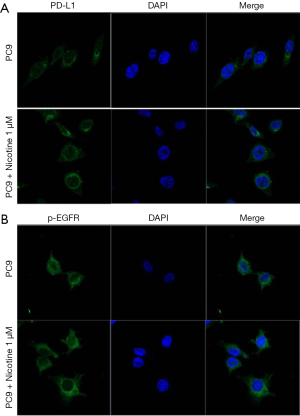
The localization of PD-L1 and p-EGFR in PC9 and PC9/N cells was determined by immunofluorescence staining counterstained with DAPI. Higher expression levels of PD-L1 and p-EGFR on membranes of EGFR sensitive cells after nicotine exposure were observed (red fluorescence, Figure 5).
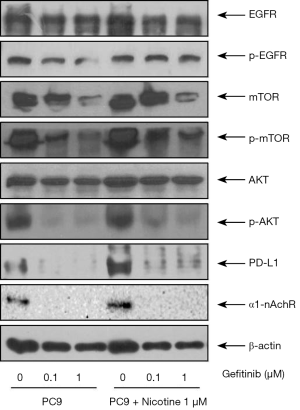
Changes of EGFR-related protein expression by chronic nicotine exposure after treatment with gefitinib
Figure 4 shows protein expression levels of EGFR, nAchR, and related pathway molecules after gefitinib treatment in PC9 and PC9/N cells. Nicotine exposure upregulated protein expressions of p-EGFR, p-mTOR, p-AKT, PD-L1, and α1-nAchR in EGFR sensitive cells. The phosphorylation of EGFR, mTOR and AKT in the PC9/N cell lines was decreased by gefitinib to a lesser extent than that observed in PC9 cells. PD-L1 expression was also decreased by gefitinib to a lesser extent than that observed in PC9 cells. These results indicate that nicotine induced mTOR-Akt activation is involved in the effect of nicotine on cell survival and cell proliferation following gefitinib treatment.
The overall PD-L1 TPS according to burden of smoking dosage
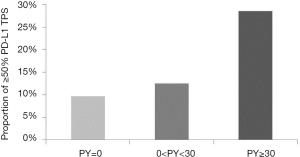
A total of 54 eligible patients with lung adenocarcinoma harboring activating EGFR mutation were included in the present study (Table 1). Their median age at diagnosis was 62 years (range, 39–88 years). Thirty-two (59.3%) of 54 patients were males. Thirty-one (57.4%) were never smokers while seven (13.0%) had heavy smoking history (≥30 PY). Most (92.6%) of these patients had advanced diseases (clinical stage III or IV). With regard to EGFR mutation status, thirty-three (61.1%) had exon 19 deletion mutation and twenty-one (38.9%) showed exon 21 point mutation. According to levels of PD-L1 TPS, seven (13.0%) patients had ≥50% TPS and twenty (37.0%) patients revealed 0% PD-L1 TPS expression. On analysis according to burden of smoking dosage, heavy smokers (≥30 PY) showed 28.5% of ≥50% PD-L1 TPS while light smoker and never smokers had 12.5% and 9.7% of ≥50% PD-L1 TPS, respectively (Figure 6). However, these results were not statistically significant (P=0.628).
Table 1
| Factors (n=54) | Frequency (%) |
|---|---|
| Median age, range | 62 [39–88]* |
| Sex | |
| Male | 32 (59.3%) |
| Female | 22 (40.7%) |
| Smoking history | |
| Never smoker | 31 (57.4%) |
| Light smoker (<30 PY) | 16 (29.6%) |
| Heavy smoker (≥30 PY) | 7 (13.0%) |
| Clinical stage | |
| I–II | 4 (7.4%) |
| III–IV | 50 (92.6%) |
| EGFR mutation | |
| Exon19 deletion | 33 (61.1%) |
| Exon 21 point mutation | 21 (38.9%) |
| PD-L1 TPS in tumor tissues | |
| 0% | 20 (37.0%) |
| 1–49% | 27 (50.0%) |
| ≥50% | 7 (13.0%) |
Data are presented as *median [range] for age and number (percentage). PY, pack-year; EGFR, epidermal growth factor receptor; PD-L1, programmed death ligand 1; TPS, tumor proportion score.
Discussion
Our study showed that chronic nicotine exposure could increase the expression of PD-L1 in EGFR mutant lung cancer cells. In addition, nicotine exposure induced nAChR expression and upregulated EGFR downstream pathways such as Akt and mTOR. These biologic changes were associated with reduced sensitivity to EGFR-TKIs, thus conferring nicotine-related resistant mechanisms in EGFR mutant cancer cells.
Nicotine, the major addictive component of tobacco smoke, can induce cell-cycle progression, migration, invasion, and evasion of apoptosis, thus contributing to the progression and metastasis of tumor initiated by tobacco carcinogens (26). These effects occur mainly through the binding of nicotine to cell-surface receptors followed by their activation (especially nAChR, and β-adrenergic receptors to a certain extent) (27). Pallai et al. (28) have shown than nicotine can bind to nAChR and induce EGF in NSCLC, resulting in transactivation of EGFR and activation of mitogenic and antiapoptotic pathways. In EGFR mutant cells, chronic nicotine exposure can induce EGFR-TKI resistance via cross-talk between α1-nAChR and EGFR in PC9 cells (16), compatible with our data. In addition, nicotine can activate PI3K/Akt-related pathway and mediate chemotherapy-induced apoptosis in a dose-dependent and time-dependent manner in vitro (29). Akt inhibitor can cause reduction in chemotherapy-induced apoptosis and block anti-apoptotic effect caused by nicotine in A549 cells (30). These findings indicate that nicotine-induced activation of EGFR downstream pathways can mediate EGFR-TKI resistance (31).
Potential therapeutic effect of nAChR antagonists has been experimentally observed in smoking-related malignancies, especially lung cancer (32). Nicotine via nAChR activation can induce invasion, migration, and EMT known to involve MEK/ERK signaling pathway in NSCLC (33,34). Although several novel compounds have recently been developed to overcome acquired EGFR-TKI resistance, little is known about the intrinsic resistance. T790M is a frequent reported mutation of EGFR gene. It has been detected in more than 50% of patients with acquired resistance (3). The major signaling pathways targeted by PD-1 are PI3K-Akt and Ras-MET-ERK pathways (35). PD-L1 expression in tumor cells is markedly increased after acquiring resistance to gefitinib in a subset of patients (36). Gefitinib-resistant sublines (PC9GR1, PC9GR2) that had no acquired T790M mutation showed increased PD-L1 expression with activation in MEK and ERK pathways and EMT phenotype. PD-L1 expression was mediated by PI3K-Akt and MET-ERK signaling pathways in EGFR mutant cell lines (37). PD-L1 upregulation promoted EMT and accelerated carcinogenesis in skin epithelial cells (38). Hata et al. (39) have demonstrated that T790M status is correlated with lower PD-L1 expression in patients with re-biopsied EGFR mutant tumors. Therefore, our results demonstrating increased PD-L1 expression via nAChR-related EGFR resistance is compatible with high PD-L1 expression with EGFR resistance mechanism by alternative bypass signaling activation, not T790M mutation.
PD-L1 expression and its biologic effect in EGFR mutant lung cancer cells are inconclusive. Previous studies have shown that EGFR activation is involved in the regulation of PD-L1 expression in cell lines (40,41) and that high PD-L1 overexpression is associated with activating EGFR mutation in EGFR mutant cells through surgically resected tumors (42). However, conflicting data have also shown that PD-L1 tumor proportion score ≥50% seldom overlaps with the presence of activating EGFR mutation in lung adenocarcinoma (43,44). According to a meta-analysis, EGFR mutation is significantly correlated with high PD-L1 expression. Such discrepancies in results might be due to heterogeneous study populations and variable definitions of PD-L1 expression (45).
Higher PD-L1 expression level was correlated with primary resistance to EGFR-TKIs in advanced EGFR mutant lung cancer (46). In addition, tumor mutation burden (TMB) is known as one of the predictive biomarkers in immunotherapy, and there exists an association between smoking history and high TMB in NSCLC patients (47). Interestingly, there was no difference in TMB based on smoking status in patients with driver mutations (48,49). These reported findings are similar with our results that PD-L1 TPS does not exhibit significant difference according to smoking history (never, light, heavy smokers).
Uninflamed tumor microenvironment with immunological tolerance and weak immunogenicity has been suggested to explain the inferior response of EGFR mutant NSCLC to anti-PD-1/PD-L1 therapy (50). Clinical responses to PD-1 blockade are associated with PD-L1 expression on tumor-resident cells induced by pre-existing CD8+ T cells in “adaptive immune resistance” infiltrating neoplastic tissue (51,52). Even if a tumor expresses PD-L1, anticancer immunosurveillance is unlikely to be reinstated if a tumor is devoid of cytotoxic T cell. Another possible reason is that tumor mutational loads in EGFR mutant-type tumors are lower than those of wild-type tumors (53). Putative biomarkers that can predict the response to anti-PD-1/PD-L1 immunotherapy including tumoral PD-L1 expression, CD8+ T-cell infiltration, and tumor mutational burden have been developed (54).
Although the role of anti-PD-1/PD-L1 therapy in TKI naïve EGFR mutant patients is currently unclear (23), clinical trials in settings of smoking-related intrinsic resistance and TKI failure without T790M are clinically challenging. A retrospective study has shown that smokers with EGFR mutant tumor can predict favorable clinical benefit provided by anti-PD-1/PD-L1 therapy (55), supporting our findings. Recent strategies of immunotherapy combined with cytotoxic chemotherapy, other immunotherapeutic agents, and radiotherapy are promising. They have potential to be applied in EGFR mutant patients. The present data did not show other biomarkers rather than PD-L1 expression. In addition, this study had limitation in that in vitro experiment was performed. Therefore, the efficacy of anti-PD-1/PD-L1 therapy needs to be tested through in vivo study.
In conclusion, chronic nicotine exposure can increase PD-L1 expression related to intrinsic resistance to EGFR-TKI in NSCLC patients harboring activating EGFR mutation. Considering the clinical importance of inevitable EGFR resistance, further studies regarding the role of anti-PD-1/PD-L1 treatment are needed, especially in EGFR mutant smokers.
Acknowledgments
Funding: This work was supported by
Footnote
Provenance and Peer Review: This article was commissioned by the Guest Editors (Hui-Yi Lin, Tung-Sung Tseng) for the series “Population Science in Cancer” published in Translational Cancer Research. The article has undergone external peer review.
Conflicts of Interest: All authors have completed the ICMJE uniform disclosure form (available at http://dx.doi.org/10.21037/tcr.2019.05.02). The series “Population Science in Cancer” was commissioned by the editorial office without any funding or sponsorship. The authors have no other conflicts of interest to declare.
Ethical Statement: The authors are accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved. The study was conducted in accordance with the Declaration of Helsinki (as revised in 2013). The study was approved by the ethics committee at each center (XC14OIMI0070). Informed consent was obtained from all patients.
Open Access Statement: This is an Open Access article distributed in accordance with the Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License (CC BY-NC-ND 4.0), which permits the non-commercial replication and distribution of the article with the strict proviso that no changes or edits are made and the original work is properly cited (including links to both the formal publication through the relevant DOI and the license). See: https://creativecommons.org/licenses/by-nc-nd/4.0/.
References
- Pao W, Girard N. New driver mutations in non-small-cell lung cancer. Lancet Oncol 2011;12:175-80. [Crossref] [PubMed]
- Tan DS, Yom SS, Tsao MS, et al. The International Association for the Study of Lung Cancer Consensus Statement on Optimizing Management of EGFR Mutation-Positive Non-Small Cell Lung Cancer: Status in 2016. J Thorac Oncol 2016;11:946-63. [Crossref] [PubMed]
- Wang J, Wang B, Chu H, et al. Intrinsic resistance to EGFR tyrosine kinase inhibitors in advanced non-small-cell lung cancer with activating EGFR mutations. Onco Targets Ther 2016;9:3711-26. [Crossref] [PubMed]
- Ohashi K, Maruvka YE, Michor F, et al. Epidermal growth factor receptor tyrosine kinase inhibitor-resistant disease. J Clin Oncol 2013;31:1070-80. [Crossref] [PubMed]
- Jänne PA, Yang JC, Kim DW, et al. AZD9291 in EGFR inhibitor-resistant non-small-cell lung cancer. N Engl J Med 2015;372:1689-99. [Crossref] [PubMed]
- Walter AO, Sjin RT, Haringsma HJ, et al. Discovery of a mutant-selective covalent inhibitor of EGFR that overcomes T790M-mediated resistance in NSCLC. Cancer Discov 2013;3:1404-15. [Crossref] [PubMed]
- Liao BC, Lin CC, Lee JH, et al. Optimal management of EGFR-mutant non-small cell lung cancer with disease progression on first-line tyrosine kinase inhibitor therapy. Lung Cancer 2017;110:7-13. [Crossref] [PubMed]
- Liu S, He Y, Jiang T, et al. EGFR-TKIs plus chemotherapy demonstrated superior efficacy than EGFR-TKIs alone as first-line setting in advanced NSCLC patients with EGFR mutation and BIM deletion polymorphism. Lung Cancer 2018;120:82-7. [Crossref] [PubMed]
- Rosell R, Moran T, Queralt C, et al. Screening for epidermal growth factor receptor mutations in lung cancer. N Engl J Med 2009;361:958-67. [Crossref] [PubMed]
- Nishinarita N, Igawa S, Kasajima M, et al. Smoking History as a Predictor of Epidermal Growth Factor Receptor Tyrosine Kinase Inhibitors in Patients with Non-Small Cell Lung Cancer Harboring EGFR Mutations. Oncology 2018;95:109-15. [Crossref] [PubMed]
- Kim MH, Kim HR, Cho BC, et al. Impact of cigarette smoking on response to epidermal growth factor receptor (EGFR)-tyrosine kinase inhibitors in lung adenocarcinoma with activating EGFR mutations. Lung Cancer 2014;84:196-202. [Crossref] [PubMed]
- Zhang P, Nie X, Bie Z, et al. Impact of heavy smoking on the benefits from first-line EGFR-TKI therapy in patients with advanced lung adenocarcinoma. Medicine (Baltimore) 2018;97:e0006. [Crossref] [PubMed]
- Hasegawa Y, Ando M, Maemondo M, et al. The role of smoking status on the progression-free survival of non-small cell lung cancer patients harboring activating epidermal growth factor receptor (EGFR) mutations receiving first-line EGFR tyrosine kinase inhibitor versus platinum doublet chemotherapy: a meta-analysis of prospective randomized trials. Oncologist 2015;20:307-15. [Crossref] [PubMed]
- Li H, Wang S, Takayama K, et al. Nicotine induces resistance to erlotinib via cross-talk between alpha 1 nAChR and EGFR in the non-small cell lung cancer xenograft model. Lung Cancer 2015;88:1-8. [Crossref] [PubMed]
- Li D, Zhang L, Zhou J, et al. Cigarette smoke extract exposure induces EGFR-TKI resistance in EGFR-mutated NSCLC via mediating Src activation and EMT. Lung Cancer 2016;93:35-42. [Crossref] [PubMed]
- Wang S, Takayama K, Tanaka K, et al. Nicotine induces resistance to epidermal growth factor receptor tyrosine kinase inhibitor by alpha1 nicotinic acetylcholine receptor-mediated activation in PC9 cells. J Thorac Oncol 2013;8:719-25. [Crossref] [PubMed]
- Pardoll DM. The blockade of immune checkpoints in cancer immunotherapy. Nat Rev Cancer 2012;12:252-64. [Crossref] [PubMed]
- Garon EB, Rizvi NA, Hui R, et al. Pembrolizumab for the treatment of non-small-cell lung cancer. N Engl J Med 2015;372:2018-28. [Crossref] [PubMed]
- Topalian SL, Hodi FS, Brahmer JR, et al. Safety, activity, and immune correlates of anti-PD-1 antibody in cancer. N Engl J Med 2012;366:2443-54. [Crossref] [PubMed]
- Herbst RS, Soria JC, Kowanetz M, et al. Predictive correlates of response to the anti-PD-L1 antibody MPDL3280A in cancer patients. Nature 2014;515:563-7. [Crossref] [PubMed]
- Govindan R, Ding L, Griffith M, et al. Genomic landscape of non-small cell lung cancer in smokers and never-smokers. Cell 2012;150:1121-34. [Crossref] [PubMed]
- Vogelstein B, Papadopoulos N, Velculescu VE, et al. Cancer genome landscapes. Science 2013;339:1546-58. [Crossref] [PubMed]
- Lisberg A, Cummings A, Goldman JW, et al. A Phase II Study of Pembrolizumab in EGFR-mutant, PD-L1+, Tyrosine Kinase Inhibitor (TKI) Naive Patients with Advanced NSCLC. J Thorac Oncol 2018;13:1138-45. [Crossref] [PubMed]
- Gainor JF, Shaw AT, Sequist LV, et al. EGFR Mutations and ALK Rearrangements Are Associated with Low Response Rates to PD-1 Pathway Blockade in Non-Small Cell Lung Cancer: A Retrospective Analysis. Clin Cancer Res 2016;22:4585-93. [Crossref] [PubMed]
- Reck M, Rodriguez-Abreu D, Robinson AG, et al. Pembrolizumab versus Chemotherapy for PD-L1-Positive Non-Small-Cell Lung Cancer. N Engl J Med 2016;375:1823-33. [Crossref] [PubMed]
- Schaal C, Chellappan SP. Nicotine-mediated cell proliferation and tumor progression in smoking-related cancers. Mol Cancer Res 2014;12:14-23. [Crossref] [PubMed]
- Singh S, Pillai S, Chellappan S. Nicotinic acetylcholine receptor signaling in tumor growth and metastasis. J Oncol 2011;2011:456743. [Crossref] [PubMed]
- Pillai S, Rizwani W, Li X, et al. ID1 facilitates the growth and metastasis of non-small cell lung cancer in response to nicotinic acetylcholine receptor and epidermal growth factor receptor signaling. Mol Cell Biol 2011;31:3052-67. [Crossref] [PubMed]
- Czyżykowski R, Polowinczak-Przybylek J, Potemski P. Nicotine-induced resistance of non-small cell lung cancer to treatment--possible mechanisms. Postepy Hig Med Dosw (Online) 2016;70:186-93. [Crossref] [PubMed]
- Zhang J, Kamdar O, Le W, et al. Nicotine induces resistance to chemotherapy by modulating mitochondrial signaling in lung cancer. Am J Respir Cell Mol Biol 2009;40:135-46. [Crossref] [PubMed]
- Yoneyama R, Aoshiba K, Furukawa K, et al. Nicotine enhances hepatocyte growth factor-mediated lung cancer cell migration by activating the alpha7 nicotine acetylcholine receptor and phosphoinositide kinase-3-dependent pathway. Oncol Lett 2016;11:673-7. [Crossref] [PubMed]
- Cesario A, Russo P, Nastrucci C, et al. Is alpha7-nAChR a possible target for lung cancer and malignant pleural mesothelioma treatment? Curr Drug Targets 2012;13:688-94. [Crossref] [PubMed]
- Zhang C, Ding XP, Zhao QN, et al. Role of alpha7-nicotinic acetylcholine receptor in nicotine-induced invasion and epithelial-to-mesenchymal transition in human non-small cell lung cancer cells. Oncotarget 2016;7:59199-208. [PubMed]
- Paleari L, Catassi A, Ciarlo M, et al. Role of alpha7-nicotinic acetylcholine receptor in human non-small cell lung cancer proliferation. Cell Prolif 2008;41:936-59. [Crossref] [PubMed]
- Boussiotis VA. Molecular and Biochemical Aspects of the PD-1 Checkpoint Pathway. N Engl J Med 2016;375:1767-78. [Crossref] [PubMed]
- Han JJ, Kim DW, Koh J, et al. Change in PD-L1 Expression After Acquiring Resistance to Gefitinib in EGFR-Mutant Non-Small-Cell Lung Cancer. Clin Lung Cancer 2016;17:263-70.e2. [Crossref] [PubMed]
- Ota K, Azuma K, Kawahara A, et al. Induction of PD-L1 Expression by the EML4-ALK Oncoprotein and Downstream Signaling Pathways in Non-Small Cell Lung Cancer. Clin Cancer Res 2015;21:4014-21. [Crossref] [PubMed]
- Cao Y, Zhang L, Kamimura Y, et al. B7-H1 overexpression regulates epithelial-mesenchymal transition and accelerates carcinogenesis in skin. Cancer Res 2011;71:1235-43. [Crossref] [PubMed]
- Hata A, Katakami N, Nanjo S, et al. Programmed death-ligand 1 expression and T790M status in EGFR-mutant non-small cell lung cancer. Lung Cancer 2017;111:182-9. [Crossref] [PubMed]
- Chen N, Fang W, Zhan J, et al. Upregulation of PD-L1 by EGFR Activation Mediates the Immune Escape in EGFR-Driven NSCLC: Implication for Optional Immune Targeted Therapy for NSCLC Patients with EGFR Mutation. J Thorac Oncol 2015;10:910-23. [Crossref] [PubMed]
- Zhang N, Zeng Y, Du W, et al. The EGFR pathway is involved in the regulation of PD-L1 expression via the IL-6/JAK/STAT3 signaling pathway in EGFR-mutated non-small cell lung cancer. Int J Oncol 2016;49:1360-8. [Crossref] [PubMed]
- Azuma K, Ota K, Kawahara A, et al. Association of PD-L1 overexpression with activating EGFR mutations in surgically resected nonsmall-cell lung cancer. Ann Oncol 2014;25:1935-40. [Crossref] [PubMed]
- Takada K, Toyokawa G, Tagawa T, et al. PD-L1 expression according to the EGFR status in primary lung adenocarcinoma. Lung Cancer 2018;116:1-6. [Crossref] [PubMed]
- Rangachari D, VanderLaan PA, Shea M, et al. Correlation between Classic Driver Oncogene Mutations in EGFR, ALK, or ROS1 and 22C3-PD-L1 >/=50% Expression in Lung Adenocarcinoma. J Thorac Oncol 2017;12:878-83. [Crossref] [PubMed]
- Zhang M, Li G, Wang Y, et al. PD-L1 expression in lung cancer and its correlation with driver mutations: a meta-analysis. Sci Rep 2017;7:10255. [Crossref] [PubMed]
- Hsu KH, Huang YH, Tseng JS, et al. High PD-L1 expression correlates with primary resistance to EGFR-TKIs in treatment naive advanced EGFR-mutant lung adenocarcinoma patients. Lung Cancer 2019;127:37-43. [Crossref] [PubMed]
- Rizvi NA, Hellmann MD, Snyder A, et al. Cancer immunology. Mutational landscape determines sensitivity to PD-1 blockade in non-small cell lung cancer. Science 2015;348:124-8. [Crossref] [PubMed]
- Nagahashi M, Sato S, Yuza K, et al. Common driver mutations and smoking history affect tumor mutation burden in lung adenocarcinoma. J Surg Res 2018;230:181-5. [Crossref] [PubMed]
- Offin M, Rizvi H, Tenet M, et al. Tumor Mutation Burden and Efficacy of EGFR-Tyrosine Kinase Inhibitors in Patients with EGFR-Mutant Lung Cancers. Clin Cancer Res 2019;25:1063-9. [Crossref] [PubMed]
- Dong ZY, Zhang JT, Liu SY, et al. EGFR mutation correlates with uninflamed phenotype and weak immunogenicity, causing impaired response to PD-1 blockade in non-small cell lung cancer. Oncoimmunology 2017;6:e1356145. [Crossref] [PubMed]
- Ribas A, Shin DS, Zaretsky J, et al. PD-1 Blockade Expands Intratumoral Memory T Cells. Cancer Immunol Res 2016;4:194-203. [Crossref] [PubMed]
- Daud AI, Loo K, Pauli ML, et al. Tumor immune profiling predicts response to anti-PD-1 therapy in human melanoma. J Clin Invest 2016;126:3447-52. [Crossref] [PubMed]
- Soo RA, Lim SM, Syn NL, et al. Immune checkpoint inhibitors in epidermal growth factor receptor mutant non-small cell lung cancer: Current controversies and future directions. Lung Cancer 2018;115:12-20. [Crossref] [PubMed]
- Topalian SL, Taube JM, Anders RA, et al. Mechanism-driven biomarkers to guide immune checkpoint blockade in cancer therapy. Nat Rev Cancer 2016;16:275-87. [Crossref] [PubMed]
- Garassino MC, Gelibter AJ, Grossi F, et al. Italian Nivolumab Expanded Access Program in Nonsquamous Non-Small Cell Lung Cancer Patients: Results in Never-Smokers and EGFR-Mutant Patients. J Thorac Oncol 2018;13:1146-55. [Crossref] [PubMed]


