
LINC00052 suppressed glioma cell proliferation and invasion by downregulating insulin-like growth factor 2
Introduction
Glioma is one of the most general malignant primary intracranial tumors in adults worldwide and brings a huge threat to public health (1). Five cases per 100,000 persons are newly diagnosed of glioma annually. Although great progress has been made in the therapeutic treatment in the past decades, the mortality rate remains high in glioma patients worldwide. The median survival rate of glioma patients is approximately 15 months, which is the poorest five-year survival rate among all cancers (2,3). Therefore, it is extremely urgent to uncover new mechanisms underlying the development of glioma and find out potential therapeutic targets for human glioma.
Noncoding RNA accounts for more than 98% of all the sequences. As one subtype of noncoding RNA, long noncoding RNAs (lncRNAs) participate in a number of cellular processes and pathways during the development of tumorigenesis. For instance, overexpression of lincRNA-p21 represses cell proliferation in gastric cancer and increases cell sensitivity of radiotherapy through regulation of the beta-catenin signaling pathway (4). LncRNA-CCHE1 expression is positively related to the malignancy of colorectal carcinoma via regulation of ERK/COX-2 pathway (5). Downregulation of lncRNA linc-ITGB1 inhibits cell invasion, cell migration and epithelial-mesenchymal transition in non-small cell lung cancer through decreasing Snail expression (6). Activated by ZEB1, lncRNA HCCL5 accelerates cell viability, cell migration, epithelial-mesenchymal transition and the malignancy of hepatocellular carcinoma (7). Moreover, through the modulation of OIP5 expression, lncRNA OIP5-AS1 promotes cell proliferation and inhibits cell apoptosis in bladder cancer (8). However, the role of lncRNA LINC00052 in the progression of glioma remains unexplored. In this study, we found out that the expression level of LINC00052 was remarkably downregulated in glioma samples. Moreover, experiments revealed that LINC00052 depressed cell proliferation, invasion and migration in glioma. Furthermore, we discovered that LINC00052 played its function in glioma by downregulating insulin-like growth factor 2 (IGF2).
Methods
Clinical samples
Human tissues were obtained from 40 glioma patients who underwent surgery at Qingdao municipal hospital. Ten normal brain tissues were collected from craniocerebral trauma or cerebral hernia patients who underwent surgery at Qingdao municipal hospital. All tissues were analyzed by two independent experienced pathologists. All tissues were kept at –80 °C. Written informed consent was offered by every glioma patient before the surgery. This study was ratified as the Ethics Committee of Qingdao municipal hospital required.
Cell culture and lentiviral virus transfection
The glioma cell line U251, U87, T98, SHG44 and U373 was obtained from the Neuroscience Institute of Soochow University. The normal human astrocyte 1800 cell line was obtained from ScienCell™ Research Laboratories (Carlsbad, CA, USA). The culture medium Dulbecco’s Modified Eagle Medium (DMEM; Hyclone, Thermo Fisher Scientific, Waltham, MA, USA) and 10% fetal bovine serum FBS (FBS; Gibco, Invitrogen, Carlsbad, CA, USA) were used to incubate the cells. For transfection, lentiviral virus targeting LINC00052 was compounded and then cloned to pLenti-EF1a-EGFP-F2A-Puro vector (Biosettia Inc., San Diego, CA, USA). LINC00052 lentiviruses (LINC00052) and empty vector were packaged in 293T cells. The entire transfection process was performed by using lipofectamine 2000 (Invitrogen, Carlsbad, CA, USA).
RNA extraction and RT-qPCR
TRIzol reagent (Invitrogen) was utilized to isolate total RNA from tissues and cells for both mRNA analyses. SYBR green (Roche, Switzerland) was conducted to measure the relative expression levels between LINC00052 and IGF2 mRNA. Moreover, the expressions were normalized to the level of β-actin mRNA. RT-qPCR analyses were conducted in triplicate. The primers were used as following: LINC00052, forwards 5'-CCTATCCCTTTCTCTAAGAA-3' and reverse 5'-ACTTCTGCAAAAACGTGCTG-3'; β-actin, forward 5'-GATGGAAATCGTCAGAGGCT-3' and reverse 5'-TGGCACTTAGTTGGAAATGC-3'. The thermal cycle was as follows: 30 sec at 95 °C, 5 sec for 40 cycles at 95 °C, 35 sec at 60 °C. The relative expression was calculated by performing 2−△△CT method.
Cell proliferation assay
Following the protocol (Dojindo Molecular Technologies, Inc.), cell proliferation of these treated cells in 96-well plates was monitored by CCK8 assay every 24 h. Spectrophotometer (Thermo Scientific, Rockford, IL, USA) was utilized to measure the absorbance at 450 nm.
Wound healing assay
Cells, transferred into 6-well plates, were cultured in DMEM medium overnight. After scratched with a plastic tip, cells were cultured in serum-free DMEM. After 48 h, wound closure was viewed. Each assay was repeated for three times independently.
In vitro invasion assay
These treated cells were transformed to the top of Matrigel-coated invasion chambers (24-well insert, 8-lm pore size; BD Biosciences) with 200 µL serum-free DMEM. And the bottom chamber was added with DMEM and FBS as a chemoattractant. After being incubated for 48 h, non-invading cells were removed from the inner part of the insert by cotton swab. After being fixed in 4% formaldehyde, cells on the lower membrane surface were stained with 0.1% crystal violet. The microscope was used to manually count invading cells in three randomly-chosen fields and take pictures.
Western blot analysis
Anti-β-actin (Santa Cruz Biotechnology, Santa Cruz, CA, USA) and anti-IGF2 (Abcam, Tokyo, Japan) were used as the primary antibodies. After separated with 12% SDS-PAGE, the protein samples were transferred onto polyvinylidene fluoride (PVDF) membrane. The primary antibodies were utilized to incubate the membranes at 4 °C for the whole night. Furthermore, after washed, membranes were incubated with goat anti-rabbit secondary antibody (ProSci, Poway, CA, USA) for 2 h. ECL western blotting detection reagents (Pierce antibodies; Thermo Fisher Scientific) was then used. Chemiluminescent film was applied for assessment of protein expression with Image J software.
Statistical analysis
SPSS 13.0 (SPSS Inc., Chicago, IL, USA) was utilized to conduct the statistical analysis. Two-tailed Student’s t-test was performed to analyze the significance. When P<0.05, the data were considered statistically significant.
Results
LINC00052 in glioma tissues and cells
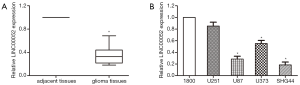
First, RT-qPCR was conducted for detecting LINC00052 expression in 40 patients’ tissues and 4 glioma cells. As the result, LINC00052 was significantly downregulated in 40 glioma patients’ tissue samples than that in 10 normal brain tissues (glioma tissues 1.13±0.63; normal brain tissues 4.86±0.93; Figure 1A). Besides, LINC00052 level was significantly lower in glioma cells than that in normal human astrocyte 1800 cell line (Figure 1B).

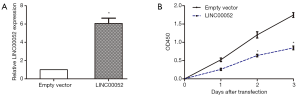
Overexpression of LINC00052 inhibits cell proliferation
According to LINC00052 expression in glioma cells, we chose SHG44 glioma cells for overexpression of LINC00052. The LINC00052 lentivirus (LINC00052) and the empty vector were synthetized and transduced into SHG44 cells. Then the LINC00052 expression was determined by RT-qPCR (Figure 2A). Moreover, the outcome of CCK8 assay suggested that cell proliferation of glioma cells was repressed after LINC00052 was overexpressed (Figure 2B).
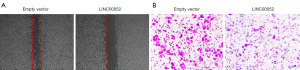
Overexpression of LINC00052 represses cell migration and invasion
Overexpression of LINC00052 inhibited glioma cell migration through wound healing assay (Figure 3A). Furthermore, cell invasion of glioma cells was inhibited after LINC00052 was overexpressed through transwell assay (Figure 3B).
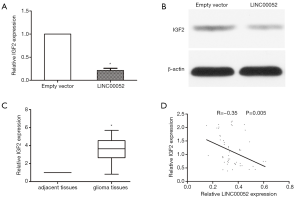
LINC00052 inhibits glioma tumorigenesis via suppressing IGF2
IGF2 mRNA expression was downregulated in glioma cells transfected with LINC00052 lentivirus through RT-qPCR (Figure 4A). IGF2 protein expression level was downregulated in glioma cells after transfected with LINC00052 lentivirus through western blot analysis (Figure 4B). To explore the interaction between LINC00052 and IGF2, the expression level of IGF2 was detected in tumor tissues. IGF2 expression increased obviously in glioma tissues when compared with that in normal brain tissues (Figure 4C). The expression of IGF2 is negatively correlated to LINC00052 expression in glioma tissues through linear correlation analysis (Figure 4D).
Discussion
LncRNAs have been reported to participate in the regulation of glioma development. For example, lncRNA ATB promotes cell migration and cell invasion in glioma by suppressing miR-2043p (9). LncRNA HOXD-AS2 facilitates the progression of glioma by regulating cell cycle and may be a potential diagnostic biomarker and therapeutic target for glioma (10). Silence of lncRNA OIP5-AS1 upregulates miR-410 and further inhibits cell proliferation, cell migration and promotes cell apoptosis in glioma by blocking the Wnt-7b/beta-catenin pathway (11). In addition, lncRNA MEG3 depresses cell proliferation and cell invasion, and induces autophagy in glioma by regulating Sirt7 and PI3K/AKT/mTOR pathway (12).
Recent researches have indicated that LINC00052 plays an important role in tumorigenesis in several cancers. For instance, LINC00052 depresses cell migration and cell invasion in hepatocellular carcinoma by upregulation of EPB41L3 (13). LINC00052 acts as an oncogene in gastric cancer by promoting cell proliferation and cell metastasis through activation of Wnt/beta-Catenin signaling pathway (14). In addition, upregulation of LINC00052 enhances the progression of breast cancer by HER3-mediated downstream signaling (15). Our study showed that the expression of LINC00052 was decreased in both glioma tissue and cells. Furthermore, after LINC00052 was overexpressed, the glioma cell growth, migration and invasion was suppressed. These data indicate that LINC00052 functions as a tumor suppressor and inhibits the development and metastasis of glioma.
IGF2, as a member of the IGF/insulin signaling pathway, possesses the capacities of anti-apoptosis and mitosis which is widely identified to be involved in modulation of tumor development and metastasis. For example, overexpression of IGF2 is remarkably correlated with the sensitivity of colorectal cancer tumor to BI 885578 (16). Curcumin functions as a tumor suppressor in the development of urothelial tumor through depressing IGF2 expression and IGF2-mediated AKT/mTOR pathway (17). LncRNA 91H enhances aggressive phenotype of breast cancer cells and upregulates the expression of H19/IGF2 (18). Moreover, high expression of IGF-2 takes part in angiogenesis in invasive bladder cancer and is associated with poor prognosis of patient (19). In our experiment, IGF2 was downregulated after LINC00052 was overexpressed in vitro. What’s more, IGF2 expression was remarkably upregulated in glioma samples compared with that in normal brain tissues. Moreover, negative correlation was discovered between IGF2 and LINC00052 expression in glioma tissues. The results above reveal that LINC00052 may realize its function in glioma progression via downregulating IGF2.
Conclusions
Our research suggests that LINC00052 is vital in the carcinogenesis of glioma and can be served as a promising biomarker for glioma. The mechanism between LINC00052 and IGF2 in glioma needs our further research.
Acknowledgments
Funding: None.
Footnote
Conflicts of Interest: All authors have completed the ICMJE uniform disclosure form (available at http://dx.doi.org/10.21037/tcr.2019.07.55). The authors have no conflicts of interest to declare.
Ethical Statement: The authors are accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved. Written informed consent was offered by every glioma patient before the surgery. This study was ratified as the Ethics Committee of Qingdao municipal hospital required. The study was conducted in accordance with the Declaration of Helsinki (as revised in 2013).
Open Access Statement: This is an Open Access article distributed in accordance with the Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License (CC BY-NC-ND 4.0), which permits the non-commercial replication and distribution of the article with the strict proviso that no changes or edits are made and the original work is properly cited (including links to both the formal publication through the relevant DOI and the license). See: https://creativecommons.org/licenses/by-nc-nd/4.0/.
References
- Sathornsumetee S, Rich JN. New treatment strategies for malignant gliomas. Expert Rev Anticancer Ther 2006;6:1087-104. [Crossref] [PubMed]
- Penas-Prado M, Gilbert MR. Molecularly targeted therapies for malignant gliomas: advances and challenges. Expert Rev Anticancer Ther 2007;7:641-61. [Crossref] [PubMed]
- Kalpathy-Cramer J, Gerstner ER, Emblem KE, et al. Advanced magnetic resonance imaging of the physical processes in human glioblastoma. Cancer Res 2014;74:4622-37. [Crossref] [PubMed]
- Chen L, Yuan D, Yang Y, et al. LincRNA-p21 enhances the sensitivity of radiotherapy for gastric cancer by targeting the beta-catenin signaling pathway. J Cell Biochem 2019;120:6178-87. [Crossref] [PubMed]
- Gaballah HH, Gaber RA, Elrashidy MA, et al. Expression of long non-coding RNA CCHE1 in colorectal carcinoma: correlations with clinicopathological features and ERK/COX-2 pathway. Mol Biol Rep 2019;46:657-67. [Crossref] [PubMed]
- Guo L, Sun C, Xu S, et al. Knockdown of long non-coding RNA linc-ITGB1 inhibits cancer stemness and epithelial-mesenchymal transition by reducing the expression of Snail in non-small cell lung cancer. Thorac Cancer 2019;10:128-36. [Crossref] [PubMed]
- Peng L, Jiang B, Yuan X, et al. Super-enhancer-associated long non-coding RNA HCCL5 is activated by ZEB1 and promotes the malignancy of hepatocellular carcinoma. Cancer Res 2019;79:572-84. [Crossref] [PubMed]
- Wang Y, Shi F, Xia Y, et al. LncRNA OIP5-AS1 predicts poor prognosis and regulates cell proliferation and apoptosis in bladder cancer. J Cell Biochem 2018; [Epub ahead of print]. [PubMed]
- Bian EB, Chen EF, Xu YD, et al. Exosomal lncRNAATB activates astrocytes that promote glioma cell invasion. Int J Oncol 2019;54:713-21. [PubMed]
- Qi Y, Wang Z, Wu F, et al. Long noncoding RNA HOXD-AS2 regulates cell cycle to promote glioma progression. J Cell Biochem 2018; [Epub ahead of print]. [PubMed]
- Sun WL, Kang T, Wang YY, et al. Long Noncoding RNA OIP5-AS1 targets Wnt-7b to affect glioma progression via modulation of miR-410. Biosci Rep 2019;BSR20180395. [Crossref] [PubMed]
- Xu DH, Chi GN, Zhao CH, et al. Long noncoding RNA MEG3 inhibits proliferation and migration but induces autophagy by regulation of Sirt7 and PI3K/AKT/mTOR pathway in glioma cells. J Cell Biochem 2018; [Epub ahead of print]. [PubMed]
- Zhu L, Yang N, Chen J, et al. LINC00052 upregulates EPB41L3 to inhibit migration and invasion of hepatocellular carcinoma by binding miR-452-5p. Oncotarget 2017;8:63724-37. [PubMed]
- Shan Y, Ying R, Jia Z, et al. LINC00052 Promotes Gastric Cancer Cell Proliferation and Metastasis via Activating the Wnt/beta-Catenin Signaling Pathway. Oncol Res 2017;25:1589-99. [Crossref] [PubMed]
- Salameh A, Fan X, Choi BK, et al. HER3 and LINC00052 interplay promotes tumor growth in breast cancer. Oncotarget 2017;8:6526-39. [Crossref] [PubMed]
- Sanderson MP, Hofmann MH, Garin-Chesa P, et al. The IGF1R/INSR Inhibitor BI 885578 Selectively Inhibits Growth of IGF2-Overexpressing Colorectal Cancer Tumors and Potentiates the Efficacy of Anti-VEGF Therapy. Mol Cancer Ther 2017;16:2223-33. [Crossref] [PubMed]
- Tian B, Zhao Y, Liang T, et al. Curcumin inhibits urothelial tumor development by suppressing IGF2 and IGF2-mediated PI3K/AKT/mTOR signaling pathway. J Drug Target 2017;25:626-36. [Crossref] [PubMed]
- Vennin C, Spruyt N, Robin YM, et al. The long non-coding RNA 91H increases aggressive phenotype of breast cancer cells and up-regulates H19/IGF2 expression through epigenetic modifications. Cancer Lett 2017;385:198-206. [Crossref] [PubMed]
- Roudnicky F, Dieterich LC, Poyet C, et al. High expression of insulin receptor on tumour-associated blood vessels in invasive bladder cancer predicts poor overall and progression-free survival. J Pathol 2017;242:193-205. [Crossref] [PubMed]


