
ONECUT2 as a new therapeutic target in androgen receptor-indifferent prostate cancer
Kaochar and Mitsiades (1) summarized in this journal the findings from two recent studies, one from our group (2), that identified the atypical homeobox protein ONECUT2 (HNF6β) as a driver of lethal prostate cancer (2,3). ONECUT2 had been identified previously as a transcriptional regulator of the prostate cancer-associated long non-coding RNA (lncRNA) PCaT1 (4), however the finding that this transcription factor (TF) plays a prominent role in a subset of aggressive prostate cancers was still unexpected. Rotinen et al. employed an integrative computational analysis of TF activity in 260 castration-resistant prostate cancers (CRPC) (2). In a theoretical CRPC TF network, ONECUT2 appeared as a prominent node that was closely associated with other known prostate cancer master regulators, including androgen receptor (AR) and EZH2. Experimental studies and analysis of human datasets demonstrated that ONECUT2 suppresses the AR signaling axis through several mechanisms, a somewhat paradoxical result given the central role played by the AR, even in CRPC. ONECUT2 was also shown to be a driver of a neural gene expression program and to be upstream of neuroendocrine master regulators, such as the placental gene PEG10. Guo et al. identified ONECUT2 as a mediator of the hypoxia response in prostate cancer and showed it to be a driver of neuroendocrine differentiation (3). Although the two studies addressed the question of the role of ONECUT2 from distinct perspectives and approaches, they are in close agreement with respect to their conclusions of the importance of this TF in aggressive prostate cancer.
Rotinen et al. identified a novel compound, CSRM617, using in silico modeling and chemical library screening, and showed it suppressed the growth of human CRPC cell lines as well as established metastases generated by intra-cardiac injection of 22Rv1 cells. Guo et al. further linked hypoxia as a therapeutic target in the transdifferentiation from adenocarcinoma to neuroendocrine prostate cancer by using a hypoxia-activated prodrug, TH-302, to suppress growth of patient-derived xenografts. This transdifferentiation to neuroendocrine variants can occur in response to AR-targeted therapy and is a mechanism whereby treatment-insensitive disease emerges following therapy with AR signaling inhibitors (ARSIs). Consequently, ONECUT2 is a targetable protein in human prostate cancer, likely with multiple approaches and strategies. The ONECUT2 network is also of great interest because other components within this network will perhaps be found to be targetable as well.
It is clear from both studies that ONECUT2 plays an important role in only a subset of lethal, or potentially lethal, prostate cancers. Which cancers are these, and how can this information be translated into a therapeutic strategy against this target? The 22Rv1 model, in which the CSRM617 agent was shown to be effective in vivo, is an aggressive human CRPC cell line that expresses AR. This model is therefore distinct from CRPC models which do not express AR, such as PC-3, where ONECUT2 was also shown in both the Rotinen et al. and Guo et al. studies to be active. AR-negative prostate cancer models are thought not to be particularly physiologically relevant, because AR expression is generally retained in the tumor, even in the case of CRPC bone and visceral metastases. The de novo small cell/neuroendocrine prostate cancer (SCNC) variant, which makes up a tiny fraction of lethal prostate cancers, is typically AR-negative. However, it is clear that ONECUT2 activity emerges in AR-positive tumors, including in patients that have not undergone treatment with ARSIs.
The biology of ONECUT2 suggests it will be possible to develop companion diagnostics that will identify patients who might benefit from targeted therapy. Rotinen et al. showed that subcellular localization of ONECUT2 multiplexed with the AR identifies cells within the tumor where the two proteins are differentially localized to the nucleus vs. the cytoplasm. The quantitative immunolabeling study of ONECUT2/AR subcellular localization is consistent with the transcriptome and laboratory studies, which demonstrated that ONECUT2 can oppose the AR through multiple mechanisms, because nuclear localization of the two TFs was found to be inversely related. Single-cell transcriptome analyses would clarify and possibly strengthen this point. A ONECUT2 activity signature was applied to a range of specimen types, including needle biopsies, suggesting that gene expression profiling may be a means of identifying patients where ONECUT2 is highly active, including prior to therapy. Because gene expression profiling can now be applied to circulating tumor cells (5), a ONECUT2 companion diagnostic may be possible as a blood test, a scenario where it would do the most clinical benefit.
Other than hypoxia, it is not clear how ONECUT2 is activated. Model systems of lineage plasticity show that ONECUT2 mRNA levels are increased in response to TP53 and RB1 inactivation (6) (Figure 1). These common inactivating mutations in aggressive cancers appear to alter the epigenomic landscape broadly, thereby creating a permissive chromatin state where developmental regulators normally silenced, or expressed at low levels, can be upregulated and assume critical roles. The emergence of SCNC transcriptome programs was recently identified as a convergent process occurring in a range of epithelial cancers and associated with particularly poor prognosis (7). Why epithelial cancers of distinct ontogenetic origin should adopt this singular strategy is not clear, however it could be a consequence of the reawakening of an early developmental process associated with cell migration, such as formation of the neural crest. It has been noted by us and others that ONECUT2 may be a viable target in other malignancies. High expression of the ONECUT2 mRNA is particularly notable in cholangiocarcinoma, liver hepatocellular carcinoma, and pancreatic adenocarcinoma, based on publicly available data. There has been limited study of this protein and its potential functional partners in cancer. Consequently, the prostate cancer studies may pave the way to productive research in other areas of cancer biology and translational medicine.
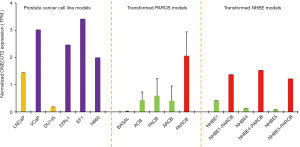
The relationship between ONECUT2 and the AR remains somewhat mysterious. Although the two proteins appear to be antagonists in many respects, the possibility that they might function cooperatively at certain genomic loci should not be ruled out. In the treatment-emergent variants of CRPC, AR is still expressed, co-existing with an emerging SCNC transcriptional program (8). ONECUT2 and AR can reside together in a multi-protein complex. How such complexes form, what their biological roles are, and under what conditions they are most active are exciting topics for future research.
Acknowledgments
Funding:
Footnote
Provenance and Peer Review: This article was commissioned and reviewed by the Section Editor Dr. Peng Zhang (Department of Urology, Zhongnan Hospital of Wuhan University, Wuhan, China).
Conflicts of Interest: All authors have completed the ICMJE uniform disclosure form (available at http://dx.doi.org/10.21037/tcr.2019.10.15). The authors have no conflicts of interest to declare.
Ethical Statement: The authors are accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved.
Open Access Statement: This is an Open Access article distributed in accordance with the Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License (CC BY-NC-ND 4.0), which permits the non-commercial replication and distribution of the article with the strict proviso that no changes or edits are made and the original work is properly cited (including links to both the formal publication through the relevant DOI and the license). See: https://creativecommons.org/licenses/by-nc-nd/4.0/.
References
- Kaochar S, Mitsiades N. Multimodal action of ONECUT2 in driving neuroendocrine prostate cancer. Transl Cancer Res 2019;8:S198-203. [Crossref] [PubMed]
- Rotinen M, You S, Yang J, et al. ONECUT2 is a targetable master regulator of lethal prostate cancer that suppresses the androgen axis. Nat Med 2018;24:1887-98. [Crossref] [PubMed]
- Guo H, Ci X, Ahmed M, et al. ONECUT2 is a driver of neuroendocrine prostate cancer. Nat Commun 2019;10:278. [Crossref] [PubMed]
- Guo H, Ahmed M, Zhang F, et al. Modulation of long noncoding RNAs by risk SNPs underlying genetic predispositions to prostate cancer. Nat Genet 2016;48:1142-50. [Crossref] [PubMed]
- Jan YJ, Yoon J, Chen JF, et al. A circulating tumor cell-RNA assay for assessment of androgen receptor signaling inhibitor sensitivity in metastatic castration-resistant prostate cancer. Theranostics 2019;9:2812-26. [Crossref] [PubMed]
- Park JW, Lee JK, Sheu KM, et al. Reprogramming normal human epithelial tissues to a common, lethal neuroendocrine cancer lineage. Science 2018;362:91-5. [Crossref] [PubMed]
- Balanis NG, Sheu KM, Esedebe FN, et al. Pan-cancer convergence to a small-cell neuroendocrine phenotype that shares susceptibilities with hematological malignancies. Cancer Cell 2019;36:17-34.e7. [Crossref] [PubMed]
- Aggarwal R, Huang J, Alumkal JJ, et al. Clinical and genomic characterization of treatment-emergent small-cell neuroendocrine prostate cancer: a multi-institutional prospective study. J Clin Oncol 2018;36:2492-503. [Crossref] [PubMed]


