
EGFR mutation is positively correlated with C-Met protein expression: a study of 446 resected lung adenocarcinoma
Introduction
Non-small-cell lung cancer (NSCLC) is a leading cause of cancer-related death world-wide. The molecular classification of lung adenocarcinoma and corresponding target therapy have effectively improved the survival status of lung cancer (1). Tyrosine kinase inhibitor (TKI) is the specific targeted therapy, which is the personalized medicine revolution in the treatment of advanced-stage NSCLC, the biomarker-defined indications for targeted therapy include epidermal growth factor receptor (EGFR)-mutant, ALK-rearranged (ALK+), ROS1-rearranged or BRAFV600E-mutant (2). In recent years, along with the development of immunotherapy, such as anti-programmed cell death 1 (PD-1), tumor mutational burden (TMB), tumor-infiltrating lymphocytes (TILs), which are also the treatment landscape of NSCLC (3). Among many molecular subtypes of lung adenocarcinoma, EGFR-mutated lung adenocarcinoma accounts for more than 50% in Chinese patients (4). EGFR is one of the four members of ErbB family of tyrosine kinase receptors. Activation of EGFR leads to regulating cellular proliferation, differentiation, and survival (5). EGFR has been identified as an oncogenic driver of NSCLC, especially EGFR mutations and its inhibition with specific TKIs have an important impact on tumor (6). Researches on the efficacy and resistance mechanism of EGFR tyrosine kinase inhibitors (EGFR-TKIs) have facilitated the continuing renewal of a series of targeting drugs. However, virtually all advanced lung cancer patients with EGFR mutation will inevitably acquire resistance or have primary resistance to EGFR-TKIs. Among these, mesenchymal-epithelial transition factor (C-Met) amplification or exon 14 splice mutation is one of the factors (7). C-Met is a multifunctional transmembrane tyrosine kinase and acts as a receptor for hepatocyte growth factor (8). It has been shown that there is a significant correlation between C-Met amplification and C-Met protein expression (9). In the patients with stage IV or treated with erlotinib after failure of first-line chemotherapy, EGFR mutation was correlated with progression-free survival (PFS) and response rate of erlotinib. While C-Met amplification of gene copy number did not show a significant correlation with erlotinib or PFS (10). According to our clinical observation, advanced patient with high C-Met protein expression alone could benefit from crizotinib (11). Nevertheless, in recent years, researchers have paid more attention to mechanisms of EGFR T790M-mutant and Met amplifications that responsible for acquired resistance to EGFR-TKIs, while little is known about the high expression of C-Met protein with primary EGFR mutation and its significance in lung adenocarcinomas.
In this study, we collected curatively resected lung adenocarcinoma specimens without preoperative neoadjuvant chemotherapy for analysis of EGFR mutation and C-Met protein expression, with the aim of exploring C-Met protein expression and its correlation of EGFR mutation in lung adenocarcinoma.
We present the following article in accordance with the STROBE reporting checklist (available at http://dx.doi.org/10.21037/tcr-20-2202).
Methods
Case selection
All pathologically diagnosed pulmonary adenocarcinoma surgical resection specimens were consecutively collected from the Department of Pathology of National Cancer Center/Cancer Hospital, Chinese Academy of Medical Sciences from 2013 to 2015. In total, 446 cases were enrolled into the analysis, among which 439 underwent curative intent surgical resection, and the other seven underwent palliative surgery. None of the patients received preoperative adjuvant chemotherapy. Clinicopathological data were extracted from medical archives, including age, gender, and AJCC 7th pathological stage and so on.
Histological review
All 446 archived pathological reports and slides were reviewed by two senior pathologists (HL and LY), according to the 2015 World Health Organization classification of lung tumors (12). Differentiation degree was also included as one of the observing variables which was defined as mild, moderate and poorly degree according to extent of difference between tumor tissue and normal lung tissue. The inclusion criteria for enrolled cases are: (I) underwent surgical resection from 2013 to 2015, (II) the pathological diagnosis of adenocarcinoma, (III) not under preoperative chemoradiotherapy. The exclusion criteria documents of clinicopathological is incompletely.
Experimental methods & positive interpretation standard
Pre experimental preparation
All surgical specimens were routinely fixed in 10% formalin for about 24 h at 10 times the volume of the tissue liquid, and then embedded in paraffin. Consecutive 4-µm-thick sections and wax roll were prepared for immunohistochemical staining and real-time PCR (RT-PCR), respectively.
C-Met protein expression was determined by immunohistochemistry (IHC). All staining steps were completed on the fully automatic Roche immunohistochemical instruments (Roche Diagnostics, Shanghai, China) according to the recommended standard protocols. C-Met protein was localized primarily in the cytoplasm (Figure 1). According to the manufacture’s scoring algorithm, intensity was scored according to a four-tier system: including negative (0), no staining or less than 5% staining; weakly positive (1+), 5–25% tumor cells stained; moderately positive (2+), 25–50% tumor cells stained; strongly positive (3+), >50% tumor cells stained. Negative quality control sections were first evaluated to remain unstained before evaluation for immunostaining on every case. For statistical analysis, negative or low expression was defined as 0 and 1+, and high expression was defined as 2+ and 3+.
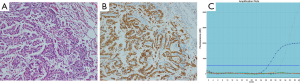
Genomic DNA was obtained using the QIAamp DNA Mini Tissue kit (Qiagen, Germany). EGFR (exon 18-21) mutation tests were performed using a RT-PCR assay (ACCB, Beijing, China), together with the Stratagene Mx3000P (Agilent Technologies Inc., Santa Clara, CA, USA).
Statistical analysis
Independent Chi-square test was used to compare frequency of clinicopathological characters between C-Met high expression and low expression groups, and between EGFR mutant and wild type groups. Spearman correlation analysis was made between EGFR mutation and C-Met protein expression. The statistical analyses were conducted using SPSS version 24.0 software. Statistical significance was set as P<0.05 (two side).
Ethical statement
The study was conducted in accordance with the Declaration of Helsinki (as revised in 2013). The study was approved by Ethics Committee of National Cancer Center/Cancer Hospital, Chinese Academy of Medical Sciences and Peking Union Medical College (No. 20/234-2430) and informed consent was taken from all the patients.
Results
Patient characteristics
The 446 lung adenocarcinomas included 181 males (40.58%) and 265 females (59.42%), with a mean age of 57 y. According to the 7th AJCC/UICC staging system, all enrolled patients were classified as stage I (n=161), stage II (n=83), stage III (n=188) and stage IV (n=14), respectively. According to 2015 WHO histological classification, they were adenocarcinoma in situ (0.22%, 1/446), minimally invasive adenocarcinoma (0.22%, 1/446), lepidic adenocarcinoma (6.28%, 28/446), acinar adenocarcinoma (53.14%, 237/446), papillary adenocarcinoma (17.26%, 77/446), solid adenocarcinoma (15.92%, 71/446), micropapillary (1.35%, 6/446), mucinous adenocarcinoma (3.81%, 17/446), adenocarcinoma combined squamous carcinoma (1.12%, 5/446), adenocarcinoma combined small cell carcinoma (0.22%, 1/446), enteric adenocarcinoma (0.22%, 1/446) and carcinosarcoma (0.22%, 1/446). As for differentiation degree, they were classified as mild (n=63), moderate (n=318), and poorly ones (n=65). Other clinicopathological features were listed in Table 1.
Table 1
| Clinicopathological features | Case number (n=446) | Percentage (%) |
|---|---|---|
| Age | 56.99±10.01 | |
| Gender | ||
| Male | 181 | 40.58 |
| Female | 265 | 59.42 |
| Pathological stage | ||
| Early (I–II) | 244 | 54.71 |
| Progressive (III–IV) | 202 | 45.29 |
| Differentiation level | ||
| Poorly | 63 | 14.13 |
| Moderate | 318 | 71.30 |
| Mild | 65 | 14.57 |
| Major subtypes | ||
| Lepidic | 28 | 6.28 |
| Acinar | 237 | 53.14 |
| Papillary | 77 | 17.26 |
| Solid | 71 | 15.92 |
| Mucinous adenocarcinoma | 17 | 3.81 |
| Special type* | 16 | 3.59 |
*, Special type includes those cases less than 1% in the enrolled patients, including adenocarcinoma in situ, minimally invasive adenocarcinoma, carcinoma combined small cell carcinoma, adenosquamous carcinoma, micropapillary adenocarcinoma, enteric adenocarcinoma and carcinosarcoma.
EGFR mutation and correlation with clinicopathological features
The overall EGFR mutation rate was 66.37% (296/446), specifically, with which located in exon 18 (3.04%, 9/296), exon 19 (42.56%, 126/296), exon 20 (3.04%, 9/296), exon 21 (48.31%, 143/296), co-existence in exons 18 and 20 (1/296, 0.33%), co-existence in 18 and 21 (1/296, 0.33%), co-existence in 20 and 21 (6/296, 2%); there’s one case with an unknown record of mutated site (1/296, 0.33%). Clinical features were compared by stratifying patients into EGFR mutant and EGFR wide type groups. Female patients harbored more mutations compared with male patients (41.98% vs. 27.92%, P<0.05). The mild and moderate differentiated tumors were more frequently mutated than the poorly differentiated tumors (80% and 69.81% vs. 34.92%, P<0.05). The mutation rate was also significantly correlated to lepidic, acinar and papillary histological subtypes (75% vs. 70.88% vs. 79.22%, P<0.05). However, there was no significant difference in pathological stages (P=0.99) (seen in Table 2).
Table 2
| Clinicopathological features | EGFR | χ2 | P value | |
|---|---|---|---|---|
| Wide type (n=150) | Mutant (n=296) | |||
| Age | 56.24±10.31 | 57.37±9.84 | 0.260 | |
| Gender | 9.531 | 0.002 | ||
| Male | 76 | 105 | ||
| Female | 74 | 191 | ||
| Pathological stage | 1.600×10–4 | 0.99 | ||
| Early (I–II) | 82 | 162 | ||
| Progressive (III–IV) | 68 | 134 | ||
| Differentiation | 35.013 | 2.495×10–10 | ||
| Poorly | 41 | 22 | ||
| Moderate | 96 | 222 | ||
| Mild | 13 | 52 | ||
| Major subtypes | 32.865 | 4.000×10–6 | ||
| Lepidic | 7 | 21 | ||
| Acinar | 69 | 168 | ||
| Papillary | 16 | 61 | ||
| Solid | 41 | 30 | ||
| Mucinous adenocarcinoma | 10 | 7 | ||
| Special type | 7 | 9 | ||
EGFR, epidermal growth factor receptor.
C-Met protein expression and correlation with clinicopathological features
C-Met protein was positively expressed in cytoplasm. Low expression and high expression of C-Met were observed in 27.35% (122/446) and 72.65% (324/446) of the cohort, respectively. The expression of C-Met protein showed a female predominance tendency (201 vs. 123, 45.67% vs. 27.57%) (seen in Table 3). There was no statistical difference in pathological stages, differentiation degrees and histological major subtypes (P>0.05).
Table 3
| Clinicopathological features | Low expression | High expression | χ2 | P value |
|---|---|---|---|---|
| Age | 57.54±9.91 | 56.78±10.01 | 0.474 | |
| Gender | 3.372 | 0.066 | ||
| Male | 58 | 123 | ||
| Female | 64 | 201 | ||
| Pathological stage | 0.343 | 0.558 | ||
| Early (I–II) | 64 | 180 | ||
| Progressive (III–IV) | 58 | 144 | ||
| Differentiation | 0.314 | 0.855 | ||
| Poorly | 17 | 46 | ||
| Moderate | 89 | 229 | ||
| Mild | 16 | 49 | ||
| Major subtypes | 4.317 | 0.505 | ||
| Lepidic | 7 | 21 | ||
| Acinar | 63 | 174 | ||
| Papillary | 20 | 57 | ||
| Solid | 21 | 50 | ||
| Mucinous adenocarcinoma | 8 | 9 | ||
| Special type | 3 | 13 |
C-Met, mesenchymal-epithelial transition factor.
Correlation between EGFR mutation and C-Met protein expression
EGFR mutation status showed a significantly positive correlation with C-Met protein expression (rs=0.095, P=0.044, see in Table 4).
Table 4
| Spearman correlation | EGRF | rs | P value | |
|---|---|---|---|---|
| Wild | Mutant | |||
| C-Met | 0.095 | 0.044 | ||
| Low expression | 50 | 72 | ||
| High expression | 100 | 224 | ||
EGFR, epidermal growth factor receptor; C-Met, mesenchymal-epithelial transition factor; rs, rank correlation coefficient of spearman.
Discussion
The molecular classification of lung adenocarcinoma and corresponding target therapy have effectively improved the clinical outcome of lung cancer, especially in patients with EGFR sensitive mutations (13). EGFR overexpression and/or mutation are associated with tumor cell proliferation, angiogenesis, tumor invasion, metastasis and inhibition of apoptosis. NSCLC patients with high expression of EGFR mutation are more prone to recurrence and metastasis (14). Likewise, in our study, 64.53% females had EGFR mutation. As there are interactions between the EGFR pathway and the estrogen receptor, EGFR mutations may be associated with estrogen (15). In adenocarcinomas, EGFR mutation correlated with estrogen receptor α expression (P=0.0029 to <0.0001) (16), and the expression of estrogen receptor β (P=0.029) (17). Meanwhile, the study showed a significant correlation of EGFR mutation with histological subtypes, which coincides with literatures (18). Ninomiya et al. (19) found that the hobnail cell type and a micropapillary morphology could predict a higher incidence of EGFR mutations in lung adenocarcinomas, meanwhile, Lee et al. (20) found EGFR mutations were frequent in well to moderately differentiated lung adenocarcinomas. Likewise, the current research showed that mild and moderate differentiated adenocarcinomas were more frequently mutated, and the mutation rate was also significantly correlated to lepidic, acinar and papillary histological subtypes, which indicated the differentiation degree and histological subtypes supported each other.
NSCLC are often intrinsically resistant to certain anticancer drugs, but our knowledge about drug resistance are still far from having a complete understanding of the underlying mechanisms. Currently, there are two specific mechanisms of acquired drug resistance, which includes T790M mutation and C-Met amplification, accounting for 50% and 20% of acquired resistant cases, respectively (7,21). Other possible mechanisms include a lack of phosphatase and tensin homolog deleted on chromosome ten (PTEN) (22), down-regulation of BIM (23), up-regulation of Integrin β1 (24), high expression of HGF (25), activation of ALK pathway (26). As for C-Met gene, gene amplification was found a mainly cause for EGFR-TKIs resistance which was first discovered by Engelman et al. at 2007 (7). Although, biologically, C-Met amplification may not necessarily lead to efficient protein expression for real function execution of the gene, which are possibly influenced by epigenetic factors, although not common. A study of Met protein expression and C-Met amplification in 316 surgically resected lung adenocarcinomas found that C-Met amplification was significantly associated with Met protein expression (P<0.001) (9). According to NCCN guidelines, the protein overexpression has not been routinely recommended for testing before EGFR-TKIs regimens. Instead, the newly updated NCCN guidelines recommend testing for C-Met gene amplification and exon 14 splice site mutation as of crizotinib targeting therapy (27). We experienced one advanced lung adenocarcinoma with C-Met protein overexpression benefitted from crizotinib (11), which suggests that the detection and analysis of C-Met protein in lung adenocarcinoma may play a role. Awad et al. found that patients with C-Met exon 14-mutation had a significant likelihood to have concurrent Met amplification and C-Met protein overexpression in NSCLC (28), meanwhile, C-Met IHC staining was a positive prognostic biomarker for overall survival in early stage NSCLC (29). In our study, the expression of C-Met protein showed a positive correlation tendency to gender (P=0.066), which is similar to Tsuta’s research (30). In another published larger investigation of 1,479 cases from our group, we identified a significant correlation of the expression of C-Met protein and gender (31). In this research, we also found a female concentration trend and positive correlation of both EGFR mutation and C-Met expression in lung adenocarcinoma, which was consistent with Nakamura et al. (32). Nakamura et al. identified phosphor-C-Met correlated with phospho-Akt (P=0.0381), and phospho-Akt expression was correlated with the expression of phospho-EGFR (P=0.0533). As activation of Akt is related to antiapoptotic of tumors, we can speculate that inhibition of the C-Met pathway may provide an alternative therapeutic approach in lung adenocarcinomas with resistance to EGFR inhibitors.
In conclusion, from the current research, we identified that overexpression of C-Met protein was positively correlated with EGFR mutation, which indicated that a synergistic effect may exist between EGFR and C-Met in lung adenocarcinoma and may somehow play a role in primary drug resistance of EGFR-TKIs. Still, there are two limits lie in our study. Firstly, we failed to identify EGFR related target therapy due to the insufficiencies of enrolled advanced cases in the curatively surgical specimens. Secondly, since we choose surgical resection specimens to study, the correlation between the therapeutic effect of EGFR-TKIs and the expression of C-Met protein could not be observed, which will be extended for further study.
Acknowledgments
Funding: Supported by
Footnote
Reporting Checklist: The authors have completed the STROBE reporting checklist. Available at http://dx.doi.org/10.21037/tcr-20-2202
Data Sharing Statement: Available at http://dx.doi.org/10.21037/tcr-20-2202
Conflicts of Interest: All authors have completed the ICMJE uniform disclosure form (available at http://dx.doi.org/10.21037/tcr-20-2202). The authors have no conflicts of interest to declare.
Ethical Statement: The authors are accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved. The study was conducted in accordance with the Declaration of Helsinki (as revised in 2013). The study was approved by Ethics Committee of National Cancer Center/Cancer Hospital, Chinese Academy of Medical Sciences and Peking Union Medical College (No. 20/234-2430) and informed consent was taken from all the patients.
Open Access Statement: This is an Open Access article distributed in accordance with the Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License (CC BY-NC-ND 4.0), which permits the non-commercial replication and distribution of the article with the strict proviso that no changes or edits are made and the original work is properly cited (including links to both the formal publication through the relevant DOI and the license). See: https://creativecommons.org/licenses/by-nc-nd/4.0/.
References
- Siegel RL, Miller KD, Jemal A. Cancer statistics, 2019. CA Cancer J Clin 2019;69:7-34. [Crossref] [PubMed]
- Camidge DR, Doebele RC, Kerr KM. Comparing and contrasting predictive biomarkers for immunotherapy and targeted therapy of NSCLC. Nat Rev Clin Oncol 2019;16:341-55. [Crossref] [PubMed]
- Bodor JN, Boumber Y, Borghaei H. Biomarkers for immune checkpoint inhibition in non-small cell lung cancer (NSCLC). Cancer 2020;126:260-70. [Crossref] [PubMed]
- Wang S, Ma P, Ma G, et al. Value of serum tumor markers for predicting EGFR mutations and positive ALK expression in 1089 Chinese non-small-cell lung cancer patients: a retrospective analysis. Eur J Cancer 2020;124:1-14. [Crossref] [PubMed]
- Singh D, Attri BK, Gill RK, et al. Review on EGFR inhibitors: critical updates. Mini Rev Med Chem 2016;16:1134-66. [Crossref] [PubMed]
- Liu X, Wang P, Zhang C, et al. Epidermal growth factor receptor (EGFR): a rising star in the era of precision medicine of lung cancer. Oncotarget 2017;8:50209-20. [Crossref] [PubMed]
- Engelman JA, Zejnullahu K, Mitsudomi T, et al. MET amplification leads to gefitinib resistance in lung cancer by activating ERBB3 signaling. Science 2007;316:1039-43. [Crossref] [PubMed]
- Hass R, Jennek S, Yang Y, et al. c-Met expression and activity in urogenital cancers - novel aspects of signal transduction and medical implications. Cell Commun Signal 2017;15:10. [Crossref] [PubMed]
- Park S, Koh J, Kim DW, et al. MET amplification, protein expression, and mutations in pulmonary adenocarcinoma. Lung Cancer 2015;90:381-7. [Crossref] [PubMed]
- Park CK, Oh IJ, Choi YD, et al. A prospective observational study evaluating the correlation of c-MET expression and EGFR gene mutation with response to erlotinib as second-line treatment for patients with advanced/metastatic non-small-cell lung cancer. Oncology 2018;94:373-82. [Crossref] [PubMed]
- Song P, Liu L, Liu Y, et al. Effectiveness of crizotinib in a patient with mesenchymal-epithelial transition overexpression/fluorescence in situ hybridization-negative/next-generation sequencing-negative advanced lung adenocarcinoma: a case report. Transl Cancer Res 2019;8:705-8. [Crossref]
- Travis WD, Brambilla E, Burke AP, et al. WHO classification of tumours of lung, pleura, thymus and heart. 4th ed. Lyon: IARC, 2015.
- Zucali PA, Ruiz MG, Giovannetti E, et al. Role of cMET expression in non-small-cell lung cancer patients treated with EGFR tyrosine kinase inhibitors. Ann Oncol 2008;19:1605-12. [Crossref] [PubMed]
- Mitra AK, Sawada K, Tiwari P, et al. Ligand-independent activation of c-Met by fibronectin and α(5)β(1)integrin regulates ovarian cancer invasion and metastasis. Oncogene 2011;30:1566-76. [Crossref] [PubMed]
- Hsu LH, Chu NM, Kao SH. Estrogen, estrogen receptor and lung cancer. Int J Mol Sci 2017;18:1713. [Crossref] [PubMed]
- Raso MG, Behrens C, Herynk MH, et al. Immunohistochemical expression of estrogen and progesterone receptors identifies a subset of NSCLCs and correlates with EGFR mutation. Clin Cancer Res 2009;15:5359-68. [Crossref] [PubMed]
- Deng F, Li M, Shan WL, et al. Correlation between epidermal growth factor receptor mutations and the expression of estrogen receptor-beta in advanced non-small cell lung cancer. Oncol Lett 2017;13:2359-65. [Crossref] [PubMed]
- Olivo-Marston SE, Mechanic LE, Mollerup S, et al. Serum estrogen and tumor-positive estrogen receptor-alpha are strong prognostic classifiers of non-small-cell lung cancer survival in both men and women. Carcinogenesis 2010;31:1778-86. [Crossref] [PubMed]
- Ninomiya H, Hiramatsu M, Inamura K, et al. Correlation between morphology and EGFR mutations in lung adenocarcinomas Significance of the micropapillary pattern and the hobnail cell type. Lung Cancer 2009;63:235-40. [Crossref] [PubMed]
- Lee B, Lee T, Lee SH, et al. Clinicopathologic characteristics of EGFR, KRAS, and ALK alterations in 6,595 lung cancers. Oncotarget 2016;7:23874-84. [Crossref] [PubMed]
- Bean J, Brennan C, Shih JY, et al. MET amplification occurs with or without T790M mutations in EGFR mutant lung tumors with acquired resistance to gefitinib or erlotinib. Proc Natl Acad Sci U S A 2007;104:20932-7. [Crossref] [PubMed]
- Bianco R, Shin I, Ritter CA, et al. Loss of PTEN/MMAC1/TEP in EGF receptor-expressing tumor cells counteracts the antitumor action of EGFR tyrosine kinase inhibitors. Oncogene 2003;22:2812-22. [Crossref] [PubMed]
- Li Z, Zhou S, Zhang L, et al. BIM induction of apoptosis triggered by EGFR-sensitive and resistance cell lines of non-small-cell lung cancer. Med Oncol 2011;28:572-7. [Crossref] [PubMed]
- Ju L, Zhou C, Li W, et al. Integrin beta1 over-expression associates with resistance to tyrosine kinase inhibitor gefitinib in non-small cell lung cancer. J Cell Biochem 2010;111:1565-74. [Crossref] [PubMed]
- Wang W, Li Q, Yamada T, et al. Crosstalk to stromal fibroblasts induces resistance of lung cancer to epidermal growth factor receptor tyrosine kinase inhibitors. Clin Cancer Res 2009;15:6630-8. [Crossref] [PubMed]
- Tiseo M, Gelsomino F, Boggiani D, et al. EGFR and EML4-ALK gene mutations in NSCLC: a case report of erlotinib-resistant patient with both concomitant mutations. Lung Cancer 2011;71:241-3. [Crossref] [PubMed]
- NCCN guidelines Version 3.2020 Non-Small Cell Lung Cancer. NCCNorg.
- Awad MM, Oxnard GR, Jackman DM, et al. MET exon 14 mutations in non-small-cell lung cancer are associated with advanced age and stage-dependent MET genomic amplification and c-Met overexpression. J Clin Oncol. 2016;34:721-30. [Crossref] [PubMed]
- Tsakonas G, Botling J, Micke P, et al. c-MET as a biomarker in patients with surgically resected non-small cell lung cancer. Lung Cancer 2019;133:69-74. [Crossref] [PubMed]
- Tsuta K, Kozu Y, Mimae T, et al. c-MET/phospho-MET protein expression and MET gene copy number in non-small cell lung carcinomas. J Thorac Oncol 2012;7:331-9. [Crossref] [PubMed]
- Yang L, Che Y, Guo L, et al. Correlation analysis of mesenchymal-epithelial transition factor protein and human epidermal growth receptor 2 protein expression in 1479 cases of lung adenocarcinoma in China. Thorac Cancer 2018;9:439-44. [Crossref] [PubMed]
- Nakamura Y, Niki T, Goto A, Morikawa T, et al. c-Met activation in lung adenocarcinoma tissues: an immunohistochemical analysis. Cancer Sci 2007;98:1006-13. [Crossref] [PubMed]


