
Mechanism of HBV-positive liver cancer cell exosomal miR-142-3p by inducing ferroptosis of M1 macrophages to promote liver cancer progression
Introduction
As a common malignant tumor, liver cancer has a very poor prognosis in developing countries (1), and its onset is a complex process with multiple factors and multiple steps (2). Hepatitis B virus (HBV) is the main pathogen that causes liver cancer (3). There are a large number of HBV patients in China, among which liver cancer caused by HBV infection accounts for about 80% of all liver cancer cases (4). Studies have reported at least eight genotypes (A–H) of HBV with different geographic distributions (5). In Europe, where genotype A and genotype D are predominant, genotype D infection is associated with more severe liver disease or hepatocellular carcinoma (HCC) than genotype A infection (6). In addition, HBV mutants are frequently found in HCC and are associated with an increased risk of HCC (7). During chronic HBV infection, recurrent liver inflammation caused by the host immune response can lead to severe liver fibrosis and cirrhosis (8). Although medical technology is continually improving, the early diagnosis rate of liver cancer in China is only about 20%, and the 5-year survival rate of liver cancer patients after treatment is about 15% in China (9). However, the relevant molecular mechanisms of HBV infection affecting the occurrence and development of liver cancer still warrant further study.
Tumor-associated macrophages (TAMs) are the main component of tumor immune infiltration and play an important role in the tumor microenvironment (10). TAMs include M1 type (classical activated macrophages) and M2 type (replacement of activated macrophages). M1-type TAMs can inhibit tumor growth and enhance immunity through acute pro-inflammatory response, immune activation response, and cell phagocytic function. M2-type TAMs can inhibit the proliferation and activation of T cells, promote the growth of tumor cells, and contribute to the occurrence of tumors (11). As liver cancer is an inflammatory tumor caused by multiple causes, TAMs play a vital role in chronic liver inflammation. Studies have shown that M1 TAMs regulate the metastasis of liver cancer cells through the nuclear factor кB/focal adhesion kinase (NF-кB/FAK) pathway (12). In addition, Macrophage-targeted immunotherapy can also provide new treatment avenues for patients with liver cancer (13). Therefore, we speculate that M1 TAMs are closely related to HBV-positive liver cancer.
Exosomes refer to small vesicles of 30–150 nm containing complex RNA and protein (14). A variety of cells can secrete exosomes under normal and pathological conditions. Many studies have shown that exosomes are important regulators of cell-to-cell communication, and have important regulatory roles in the body’s immune response, antigen presentation, and tumor invasion (15). Literature has confirmed that exosomal miRNAs secreted by tumor cells can affect tumor development by regulating the biological behavior of TAM cells. For example, treatment with epigallocin can induce breast cancer cells to secrete exosomes, and exosomal miR-16 can be regulated by M1-type TAM, which can have an anti-cancer effect (16).
Ferroptosis is a type of cell death that occurs due to the inhibition of the cystine-glutamate exchanger on the plasma membrane of the cell. It reduces the cell's acquisition of cystine, resulting in reactive oxygen-free radicals on membrane lipids. The accumulation of reactive oxygen species (ROS) eventually leads to cell death (17). Studies have shown that iron death is closely related to the occurrence, progression and suppression of cancer (18). The researchers also found that ferroptosis can promote tumor growth by driving the polarization of macrophages in tumor microenvironment (TME) (19). A variety of substances and external conditions can trigger cell ferroptosis. Of these, miRNAs have been found to regulate cell ferroptosis (20). For example, miR-9 inhibits ferroptosis of melanoma cells by targeting the downregulation of glutamate-oxaloacetate transaminase 1 expression (21). Recombinant solute carrier family 3, member 2 (SLC3A2) is a cystine-glutamate antiporter (xCT)-related protein. Its abnormal expression is closely related to cell ferroptosis. Regulating the expression of SLC3A2 can weaken the uptake of cystine by tumor cells, thereby promoting tumor cells’ lipid peroxidation and ferroptosis (22).
Early in our research, microarray experiments confirmed that miR-142-3p was abnormally highly expressed in exosomes of HBV-infected liver cancer cells. In the present study, we found that miR-142-3p can target the downregulation of SLC3A2 expression in M1 macrophages, thereby promoting ferroptosis in M1 macrophages, and ultimately leading to the further deterioration of liver cancer. We present the following article in accordance with the ARRIVE reporting checklist (available at https://tcr.amegroups.com/article/view/10.21037/tcr-22-96/rc).
Methods
Ethics statement
The samples were collected from HCC patients at the Calmette Affiliated Hospital of Kunming Medical University (Kunming, Yunnan, China). The study was conducted in accordance with the Declaration of Helsinki (as revised in 2013). The research protocol and the use of human samples were approved by the Medical Ethics Committee of the First People’s Hospital of Kunming (No. YLS2020-08) and all participants signed informed consent forms.
Cell culture
HBV stable replication cell lines (HepG2.2.15) were used in the present study. HepG2.215 cells were cultured in Dulbecco’s modified Eagle’s medium (DMEM; 31600034, Hyclone, Logan, UT, USA) containing 10% fetal bovine serum (FBS; Gibco, USA), 100 U/mL penicillin, and 100 µg/mL streptomycin (Gibco, USA) at 37 ℃ and 5% CO2.
Quantitative reverse transcription polymerase chain reaction (qRT-PCR)
Total RNA was extracted using TRIzol (Invitrogen, Carlsbad, CA, USA); 5 µg of RNA was taken, and the RNA was converted to cDNA using an RT-PCR kit (Promega, Madison, WI, USA). cDNA was used as the template and was amplified by PCR (SYBR Green Real-time PCR Master Mix; TaKaRa, Otsu, Japan). After obtaining Ct values, relative expression levels were calculated by 2−ΔΔCt method.
Western blotting
After cell proteins were extracted, they were heated at 100 ℃ for 3 min to denature them. The protein was transferred to the negative control (NC) membrane, and the NC membrane was sealed in 5% skim milk powder [phosphate-buffered saline (PBS) preparation] at 37 ℃ for 2 h or 4 ℃ overnight. The primary antibody was then added to bind the target protein. The primary antibodies—CD9 (Abcam, Cambridge, UK), CD81 (Abcam, Cambridge, UK), Tumor susceptibility gene 101 (TSG101) (Abcam, Cambridge, UK), Heat Shock Protein 70 (HSP70) (Abcam, Cambridge, UK), ALG-2-interacting protein X (ALIX) (Abcam, Cambridge, UK), and flotillin-1 (Abcam, Cambridge, UK)—were incubated in a shaker at room temperature for 2 h or overnight at 4 ℃. The secondary antibody of horseradish peroxidase (HRP)-labeled goat anti-rabbit IgG (Abcam, Cambridge, UK) or goat anti-mouse (Abcam, Cambridge, UK) was incubated and incubated in a shaker for 1 h or overnight at 4 ℃. Diaminobenzidine (DAB) kit was used to develop color. ImageJ software (Bethesda, Maryland, USA) was used for gray value analysis of protein bands. Experimental results were recorded, and the NC film was dried, scanned, and preserved.
3-(4,5-dimethyl-2-thiazolyl)-2,5-diphenyl-2-H-tetrazolium bromide (MTT) assay
Single cell suspensions were inoculated on 96-well plates with 1,000–10,000 cells per well. After 3–5 days of cell culture, 20 µL MTT solution was added to each well. Cell culture continued for 4 h, and the supernatant in the hole was absorbed after the culture was stopped. For suspended cells, the supernatant in the hole was absorbed after centrifugation. A total of 150 µL Dimethyl sulfoxide (DMSO) was added to each well for oscillatory melting. A wavelength of 570 nm was selected to measure the light absorption value of each hole on an enzyme-linked immunosorbent monitor, and the results were recorded.
Immunohistochemistry
Paraffin-embedded tissue was sectioned, and the sections were dewaxed, soaked in 3% H2O2 for 10 min, and rinsed with water twice. Citric acid buffer was added, section cooked in microwave oven for 3 min, and cooled to room temperature, and then recooked and cooled to room temperature to complete antigen repair. The serum was dropped and sealed in a wet box at 37 ℃ for 30 min, and primary anti-CD68 Monoclonal antibody (mAb) (Abcam, UK) and CD163 mAb (NBP1-95135; Novus, Missouri, USA) were added. The mixture was then refrigerated at 4 ℃ overnight. Second antibody working solution was dropped and incubated at room temperature for 30 minutes. The second antibody was discarded and cleaned with PBS. The excess PBS on the slices was removed, then DAB dye solution was used for color rendering, followed by hematoxylin redyeing, alcohol gradient dehydration, xylene transparency, and finally resin sealing, and the experimental results could be observed under a microscope.
Luciferase assays
Huh7 cells were co-transfected with wild-type or mutant SLC3A2 3'-untranslated region (UTR) psiCHECK-2 plasmid (Promega, Madison, WI, USA) and miR-142-3p/mimics or control using Lipofectamine 2000 (Invitrogen, USA). Cells were collected 48 h after transfection, cell lysis was completed according to the instructions of the dual luciferase detection kit, and fluorescence intensity was detected using a luminometer fluorescence detector (Promega Corporation, Madison, WI, USA) or a device with a similar detection function.
Exosome isolation and identification
After the operation, fresh liver cancer tissue and adjacent tissue specimens were immersed in PBS containing double antibodies against streptomycin and penicillin once they were isolated. Under aseptic operating conditions, the tissue block was cut into 1–3-mm fragments using scissors. After rinsing the blood with PBS containing double antibody. The tissue fragments were added to DMEM/F12 medium, and type I collagenase (50–100 U/mL), type IV collagenase (20–50 U/mL), and hyaluronidase (20–50 U/mL) were all added at the same time. The extracellular matrix was degraded to facilitate the release of exosomes, and DNase (0.1 mg/mL) was added to digest the genomic DNA released by dead cells. Tissue fluid was incubated in a constant temperature shaker at 37 ℃ for 30–60 min to fully release the exosomes in the tumor tissue. The digestion fluid was filtered through a 70-µm filter to remove undigested tissue fragments. After centrifugation at 300 ×g for 10 min and 2,000 ×g for 20 min, the cell pellet was removed and the supernatant was obtained. The supernatant was centrifuged at 4 ℃, 16,500 ×g for 45 min, and 100,000 ×g for 2 h to obtain exosomes. Morphological characteristics of exosomes were observed by transmission electron microscope, and the expression of exosomal markers was detected by Western blotting.
Exosome labeling and tracking
The isolated purified exosomes were collected and labeled with PKH67 Green fluorescent membrane linker dye (Sigma-Aldrich, St Louis, MO, USA). The specific labeling method was carried out according to the manufacturer’s instructions. Then, the labeled exosome weight was suspended and added to macrophages. After incubation at 37 ℃ for 30 min, 2 h, or 12 h, the uptake of exosomes by macrophages was observed under a fluorescent microscope.
Glutathione (GSH) assay
Cell GSH level was assessed using a GSH luminesce assay kit (Promega, USA) following the manufacturer’s instructions.
Lipid peroxidation assay
Tissues or cells were homogenized or cleaved using lysate (Biyuntian, Biotechnology Shanghai, China). The supernatant was centrifuged at 10,000–12,000 ×g for 10 min for subsequent determination. A BCA protein concentration determination kit (Thermo Fisher Scientific, Waltham, MA, USA) was used to determine the protein concentration. After that, 0.1 mL homogenate was added into the centrifuge tube, a 0.1 mL sample was added for determination, and 0.2 mL malondialdehyde (MDA) detection working fluid was then added. After mixing, the mixture was heated at 100 ℃ or in a boiling water bath for 15 min, cooled to room temperature in the water bath, and centrifuged at 1,000 ×g for 10 min at room temperature. A total of 200 µL supernatant was added to the 96-well plate, and absorbance was measured at 532 nm with a microplate reader. Finally, the molar concentration of MDA was calculated according to the standard curve.
Lipid ROS assay
The cells inoculated in 6-well plates were treated with C11 BODIPY 581/591 (Glpbio, USA) (50 µM) and incubated for 1 h. Cells were washed twice with PBS to remove excess dye. Cells were digested with trypsin and resuspended in PBS containing 5% FBS. Lipid ROS were analyzed by flow cytometry.
Murine liver cancer induction and treatment
The liver cancer model was established by injecting diethylnitrosamine (DEN; Sigma-Aldrich, USA) into 2-week-old AlblHBV mice or wild-type C57BL/6J mice (23). The mice were kept in a Specific Pathogen Free (SPF) environment after intraperitoneal injection of 25 µg/g DEN. Fourteen weeks after DEN administration (24), precancerous clusters of hepatocytes and hepatocellular nodules formed at 6 weeks were observed. DEN-injected AlblHBV mice were then randomly divided into 3 groups. One group (n=5) only received DEN induction, another group (n=10) received DEN-induced posterior caudal vein injection of miR-142-3p inhibitor, and the other group (n=10) received DEN-induced posterior caudal vein injection of miR-142-3p inhibitor and erastin. Only the DEN-induced group was the control group. Similarly, DEN-injected C57BL/6J mice were also divided into 3 groups. One group (n=5) received DEN induction only, another group (n=10) received DEN induction followed by tail vein injection of miR-142-3p mimic, and the other group (n=10) received DEN induction followed by tail vein injection of miR-142-3p inhibitor and zileuton. Similarly, the group receiving DEN induction only was used as the control group. All the mice were killed 20 weeks after DEN administration. After the mice were sacrificed, the tumor size was measured, and the tumor tissue was taken to detect macrophage marker molecules by immunohistochemical method. Mice were treated with non-steroidal anti-inflammatory analgesic agents during or after operation. The index of the end of the experiment was to terminate the experiment and kill the animals when the maximum tumor volume and diameter of the mice reached 1 cm. The study protocol and the use of mice model were approved by the Medical Ethics Committee of the First People’s Hospital of Kunming (No. YLS2020-08), in compliance with the Medical Ethics Committee of the First People’s Hospital of Kunming guidelines for the care and use of animals.
Statistical analysis
The experimental results were statistically analyzed by GraphPad Prism software (La Jolla, CA, USA). Results were expressed as mean ± standard deviation. Unpaired Student’s t-test (two-tail test) was used for both groups, and one-way analysis of variance test was used for multiple groups. P<0.05 was considered statistically significant.
Results
HBV-positive infection promotes exosomal secretion and affects the differentiation of M1
To confirm the influence of HBV infection on the progression of liver cancer, and to determine that the influence mainly regulates the biological behavior of M1 macrophages through miRNAs in exosomes, we analyzed M1 macrophages and M2 macrophages in the cancer tissues of patients with HBV-positive and HBV-negative infection. Compared with patients with HBV-negative infection, M1 macrophages in cancer tissues of HBV-positive patients were significantly reduced, but there was no significant difference in the expression of M2 macrophages (Figure 1A). By extracting exosomes from cancer tissues of patients with HBV-positive and HBV-negative infection, we found that exosomes are enriched in liver cancer tissues of HBV-positive patients, and their expression is significantly higher than that of exosomes in liver cancer tissues of HBV-negative patients (Figure 1B). At the same time, Western blotting was used to detect the expression of exosomal proteins CD9, CD81, TSG101, HSP70, ALIX and flotillin-1. It was found that the expression level of exosomal proteins with HBV-positive infection was higher than the amount of exosomal proteins with HBV-negative infection (Figure 1C). Finally, the qRT-PCR method was used to detect miR-142-3p, and the Western blot method was used to detect the expression of SLC3A2. It was found that miR-142-3p was significantly upregulated in HBV-positive exosomes compared with HBV-negative exosomes. The expression of SLC3A2 was significantly downregulated (Figure 1D,1E).
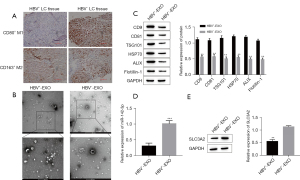
HBV-positive exosomes affect the polarization of macrophages
To further explore the relationship between HBV-positive exosomes and the abnormal expression of M1, when HBV-positive exosomes were co-cultured with macrophages, tumor-derived exosomes labeled with fluorescent PKH67 were internalized by unstained macrophages over time (Figure 2A). Compared with HBV-negative exosomes or PBS, the expression of M1 markers interleukin-6 (IL-6), tumor necrosis factor-α (TNF-α), CD-86, inducible nitric oxide synthase (iNOS) in macrophages incubated with HBV-positive exosomes was reduced (Figure 2B-2D). There was no significant difference in the expression of M2 markers arginase-1 (Arg-1), interleukin-10 (IL-10), CD206 (Figure 2E-2G). At the same time, it was found that the expression of miR-142-3p in the macrophages of the HBV-positive exosome co-culture group was significantly increased (Figure 2H), and the expression of SLC3A2 was significantly decreased (Figure 2I). There was no significant difference in the co-cultivation group of stain-negative exosomes (Figure 2H,2I).
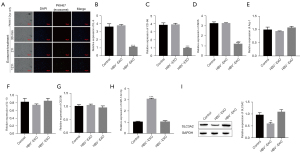
HBV-positive exosomes induce ferroptosis of M1 macrophages
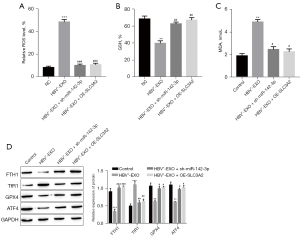
PBS and M1 macrophages were co-cultured as the control group, and HBV-positive or HBV-negative exosomes were co-cultured with M1 macrophages as the experimental group. The proliferation activity of M1 macrophages was detected by MTT. Compared with the control group, cell proliferation activity of the HBV-positive exosome co-culture group was significantly reduced, while the cell proliferation activity of the HBV-negative exosome co-culture group did not change significantly (Figure 3A). At the same time, the related indicators of the occurrence of ferroptosis were detected, and compared with the control group, the ROS level of the HBV-positive exosome co-culture group increased (Figure 3B), GSH level decreased (Figure 3C), lipid peroxidation level increased (Figure 3D), transferrin receptor 1 (TfR1) expression increased, and ferritin heavy chain 1 (FTH1), recombinant glutathione peroxidase 4 (GPX4), and activating transcription factor 4 (ATF4) expression decreased (Figure 3E), but the HBV-negative exosome co-culture group had no significant difference compared with the control group. When the ferroptosis inhibitor, zileuton, was added to the HBV-positive exosome co-cultivation group, the above phenomenon can be reversed, namely, compared with the HBV+-EXO group, the proliferation activity of M1 macrophages in HBV+-EXO + zileuton group increased (Figure 3F), ROS level decreased (Figure 3G), GSH level increased (Figure 3H), MDA content decreased (Figure 3I), the expression of TfR1 decreased (Figure 3J), and the expression of FTH1, GPX4, and ATF4 increased (Figure 3J).
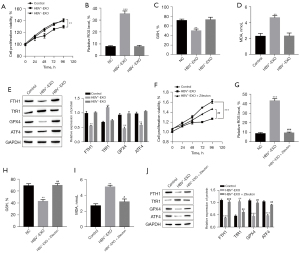
Exosomal miR-142-3p targets and negatively regulates the expression of SLC3A2
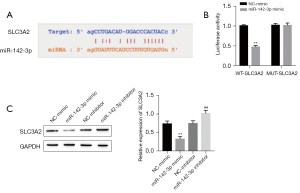
Bioinformatics database analysis indicated a potential miR-142-3p binding site on the 3'-UTR of the SLC3A2 gene (Figure 4A). The verification results of the dual luciferase reporter gene showed that the relative fluorescence intensity of SLC3A2 in the miR-142-3p mimic + SLC3A2–3'-UTR wild type (WT) group was significantly lower than that in the NC mimic + SLC3A2–3'-UTR WT group. After the miR-124 binding site was mutated, the inhibitory effect of miR-142-3p mimic on the reporter gene disappeared. The relative fluorescent intensity of SLC3A2 in the miR-142-3p mimic + SLC3A2–3'-UTR WUT group compared with the NC mimic + SLC3A2–3'-UTR WUT group was equivalent (Figure 4B). Western blot test results confirmed that compared with the NC mimic group, the miR-142-3p mimic group can significantly downregulate the SLC3A2 protein expression level. Compared with the NC-inhibitor group, the miR-142-3p inhibitor group SLC3A2 protein expression level was significantly increased (Figure 4C).
Exosomal miR-142-3p induces ferroptosis of M1 macrophages through targeted regulation of SLC3A2
To verify that miR-142-3p induces ferroptosis in M1 macrophages through targeted regulation of the expression of SLC3A2, we knock down miR-142-3p using the short hairpin structure, or transfect a plasmid overexpressing SLC3A2 in M1 macrophages, and then co-culture with HBV-infected positive exosomes. Results showed that knockdown of miR-142-3p or overexpression of SLC3A2 could effectively reverse the occurrence of ferroptosis induced by co-culture with HBV-positive exosomes, that is, compared with HBV+-EXO group, HBV+-EXO + sh-miR-142-3p group or HBV+-Exo + OE-SLC3A2 group, ROS level decreased (Figure 5A), GSH level increased (Figure 5B), MDA content decreased (Figure 5C), TfR1 expression down-regulated (Figure 5D), FTH1, GPX4, and ATF4 expression up-regulated (Figure 5D).
Exosomal miR-142-3p induces ferroptosis of M1 macrophages in vivo
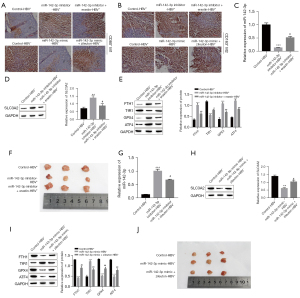
To further evaluate the effect of exosomal miR-142-3p on the ferroptosis of M1 macrophages, we injected miR-142-3p inhibitor or miR-142-3p inhibitor + Erastin into AlblHBV mice (HBV positive). At the same time, miR-142-3p mimic or miR-142-3p mimic + Zileuton was injected into C57BL/6J (HBV-negative) mice. After 6 weeks, it was found that, compared with C57BL/6J mice, the expression of M1 macrophage marker CD80 was significantly decreased in AlblHBV mice, while the M2 macrophage marker CD163 had no significant difference (Figure 6A,6B). In AlblHBV mice, the expression of CD80 increased after miR-142-3p inhibitor was injected, but when miR-142-3p inhibitor and the inducer of ferroptosis Erastin were injected, the expression of CD80 could be reversed, and the expression of CD163 was also not significant (Figure 6A,6B). We also detected the expression of miR-142-3p, SLC3A2, ferroptosis-related factors, and the size of subcutaneous tumors in mice. The results showed that compared with C57BL/6J (HBV−) mice, in AlblHBV mice (HBV+), the expression of miR-142-3p was up-regulated and the expression of SLC3A2 was down-regulated. At the same time, in AlblHBV mice, injection of miR-142-3p inhibitor can down-regulate the expression of miR-142-3p and up-regulate the protein expression of SLC3A2, inhibit the occurrence of ferroptosis and reduce the size of subcutaneous tumors, while simultaneously injecting miR-142-3p inhibitor and ferroptosis activator Erastin will reverse the above phenomenon, that is, compared with miR-142-3p inhibitor-HBV+ group, miR-142-3p inhibitor + Erastin-HBV+ group the expression of miR-142-3p is up-regulated (Figure 6C), SLC3A2 is down-regulated (Figure 6D), TfR1 is up-regulated, FTH1, GPX4, and ATF4 are down-regulated (Figure 6E), and the size of subcutaneous tumors increases (Figure 6F). In C57BL/6J mice, injection of miR-142-3p mimic can promote the expression of miR-142-3p and inhibit the protein expression of SLC3A2, and at the same time activate the occurrence of ferroptosis and increase the size of subcutaneous tumors. Simultaneous injection of miR-142-3p inhibitor and ferroptosis inhibitor Zileuton will reverse the above phenomenon. Compared with miR-142-3p mimic-HBV− group, miR-142-3p mimic + Erastin-HBV− group miR-142-3p expression is down-regulated (Figure 6G), SLC3A2 expression is up-regulated (Figure 6H), TfR1 expression is down-regulated, FTH1, GPX4, and ATF4 expression is up-regulated (Figure 6I), and subcutaneous tumor volume is reduced (Figure 6J).
Discussion
In the present study, we determined the regulatory effect of HBV-positive liver cancer exosomes on the iron death of M1 macrophages and the molecular mechanism of their existence. Intrahepatic HBV cccDNA is the main transcription template and necessary marker for virological cure (25). Macrophages in the liver play a key role in defending against HBV infection (26,27). A number of previously published studies have confirmed that, in the TEM, tumor cells and their surroundings are infiltrated with various immune cells, stromal cells, and other related cells (28). The TEM can interact with tumor cells through infiltrating cells, thereby regulating tumor growth, invasion, and metastasis (29). Because liver inflammation after HBV infection is an important factor in the development of liver cancer, studies have also confirmed that the continuous production of IL-23 caused by liver cell damage after chronic HBV infection can promote liver cancer by affecting macrophages (30). As the main component of tumor immune infiltration, TAMs play an important role in the TEM. Among them, M1 type macrophages can release a variety of pro-inflammatory factors, immune activating factors, and chemokines, which can inhibit tumor growth and enhance immunity. The preliminary test results of the present study confirmed that M1-type macrophages in tumor tissues of HBV-positive liver cancer patients were significantly less than those of HBV-negative liver cancer patients; there was no significant difference between M2-type macrophages in the tumor tissues of HBV-positive and HBV-negative liver cancer patients. This indicates that M1 macrophages are closely related to HBV-positive liver cancer.
Exosomes play a key role in cell-to-cell communication in diseases (31). For example, exosomes have been shown to participate in the spread of HBV components(32). The effect of tumor-derived exosomal miRNAs on liver cancer progression has also been reported in the literature, for example, exosomal miR-1247-3p secreted by highly metastatic hepatocellular carcinoma can induce the activation of cancer-associated fibroblasts to promote lung metastasis of liver cancer (33). Endoplasmic reticulum stress induces hepatocellular carcinoma (HCC) cells to release exosomal miR-23a-3p, which attenuates antitumor immunity by regulating the expression of programmed death ligand 1 (PD-L1) in macrophages (34). In addition, previously published studies have confirmed that exosomal miRNAs secreted by tumor cells can affect tumor development by regulating the biological behavior of TAM cells. Exosomal miRNAs can be transported from donor cells to recipient cells, and then regulate the replication of viruses to multiple viruses (35,36). For example, miR-21 in bladder cancer cell-derived exosomes regulates M2 TAM by activating the phosphatidylinositide 3-kinases (PI3K)/AKT signaling pathway in TAM, thereby promoting the migration and invasion of bladder cancer cells (37). Our research group confirmed, through chip experiments in the early stage, that miR-142-3p is abnormally highly expressed in the exosomes of HBV-infected liver cancer cells. By extracting exosomes from HBV-positive and HBV-negative liver cancer tissues, we found that miR-142-3p was highly expressed in HBV-positive exosomes. Co-culture with HBV-positive exosomes promoted ferroptosis and inhibited the proliferation activity of M1 macrophages.
Ferroptosis is defined as iron-dependent regulatory necrosis caused by membrane damage mediated by massive lipid peroxidation (38). Ferroptosis can lead to the death of white blood cell subsets and the corresponding loss of immune function. For example, lipid peroxidation-induced T-cell death promotes viral or parasitic infection (39). Therefore, ferroptosis is closely related to the occurrence of related inflammatory diseases. Studies have confirmed that when magnetosomes with FeO magnetic nanoclusters as the core are used for anti-cancer treatment, their entry into tumors can increase the content of H2O2 in polarized M1 macrophages and promote magnetism. The release of Fe ions in the corpuscles induces ferroptosis. Therefore, ferroptosis and immune regulation in tumors can produce effective treatments for tumors through a synergistic effect (40). In the present study, we found that ferroptosis inhibitors can reverse the inhibitory effect of CD80+ M1-type macrophages treated with HBV-positive liver cancer cell exosomes on the invasion ability of liver cancer cells. xCT functions in the process of transporting a molecule of cystine into the cell and releasing a molecule of glutamate to the outside of the cell (41), xCT-related proteins, and gluten. The abnormal expression of aminoamide transporter-related proteins is closely related to cell iron death. For example, the tumor suppressor gene, BRCA1-associated protein 1 (BAP1), promotes lipid peroxidation by downregulating the expression level of SLC7A11, thereby promoting tumor cell ferroptosis (42). Meanwhile, studies have shown that miR-137 can inhibit the occurrence of ferroptosis in melanoma cells by targeting the regulation of the expression of SLC1A5 (43). In this study, SLC3A2 expression level was significantly down-regulated after co-culture of M1 macrophages and HBV-positive exosomes, and HBV-positive liver cancer cell exosomal miR-142-3p have a targeting relationship with SLC3A2, and that miR-142-3p participates in the regulation of M1 macrophages ferroptosis by targeting SLC3A2, which in turn affects the progression of liver cancer. Tumor cells undergo various forms of regulated cell death during their growth. The activation or acceleration of these forms of regulated cell death has become a potential strategy for cancer treatment. For example, most anticancer drugs currently in clinical use are based on representative the specific apoptotic signaling pathway induces cancer cell death (44,45). Recent studies have shown that induction of ferroptosis is a beneficial and promising anticancer therapeutic strategy, and understanding the interaction between ferroptosis and TME may provide new and effective anticancer strategies (46). Therefore, the molecular mechanism proposed in this study enriches the theoretical basis of the interaction between ferroptosis and TME in the progression of liver cancer, and has a positive impact on the further treatment and prognosis of liver cancer, opening up a new way for live cancer treatment.
Conclusions
In the present study, our sample tissues, cell experiments, and mouse models indicated that miR-142-3p expression is upregulated in HBV-infected exosomes, SLC3A2 expression is downregulated. At the same time, M1-type macrophages in tumor tissues of HBV-positive patients were significantly less than those in HBV-negative patients. MiR-142-3p can significantly induce the ferroptosis of M1 macrophages, and regulates the expression of SLC3A2 to mediate the ferroptosis of M1 macrophages, affecting the proliferation of liver cancer.
Acknowledgments
Funding: The study was supported by
Footnote
Reporting Checklist: The authors have completed the ARRIVE reporting checklist. Available at https://tcr.amegroups.com/article/view/10.21037/tcr-22-96/rc
Data Sharing Statement: Available at https://tcr.amegroups.com/article/view/10.21037/tcr-22-96/dss
Conflicts of Interest: All authors have completed the ICMJE uniform disclosure form (available at https://tcr.amegroups.com/article/view/10.21037/tcr-22-96/coif). The authors have no conflicts of interest to declare.
Ethical Statement: The authors are accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved. The study was conducted in accordance with the Declaration of Helsinki (as revised in 2013). The study was approved by ethics board of the First People’s Hospital of Kunming (No. YLS2020-08) and informed consent was taken from all the patients. Animal experiments were performed under a project license (No. YLS2020-08) granted by the Medical Ethics Committee of the First People’s Hospital of Kunming, in compliance with the Medical Ethics Committee of the First People’s Hospital of Kunming guidelines for the care and use of animals.
Open Access Statement: This is an Open Access article distributed in accordance with the Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License (CC BY-NC-ND 4.0), which permits the non-commercial replication and distribution of the article with the strict proviso that no changes or edits are made and the original work is properly cited (including links to both the formal publication through the relevant DOI and the license). See: https://creativecommons.org/licenses/by-nc-nd/4.0/.
References
- Islami F, Miller KD, Siegel RL, et al. Disparities in liver cancer occurrence in the United States by race/ethnicity and state. CA Cancer J Clin 2017;67:273-89. [Crossref] [PubMed]
- Pittala S, Krelin Y, Shoshan-Barmatz V. Targeting Liver Cancer and Associated Pathologies in Mice with a Mitochondrial VDAC1-Based Peptide. Neoplasia 2018;20:594-609. [Crossref] [PubMed]
- Shih C, Yang CC, Choijilsuren G, et al. Hepatitis B Virus. Trends Microbiol 2018;26:386-7. [Crossref] [PubMed]
- Zhou Y, Wan Y, Ye ZW, et al. How Hepatitis B virus causes cirrhosis and liver cancer. Med Hypotheses 2017;108:52-3. [Crossref] [PubMed]
- Lin CL, Kao JH. The clinical implications of hepatitis B virus genotype: Recent advances. J Gastroenterol Hepatol 2011;26:123-30. [Crossref] [PubMed]
- Sánchez-Tapias JM, Costa J, Mas A, et al. Influence of hepatitis B virus genotype on the long-term outcome of chronic hepatitis B in western patients. Gastroenterology 2002;123:1848-56. [Crossref] [PubMed]
- Pollicino T, Cacciola I, Saffioti F, et al. Hepatitis B virus PreS/S gene variants: pathobiology and clinical implications. J Hepatol 2014;61:408-17. [Crossref] [PubMed]
- Xie Y, Hepatitis B. Virus-Associated Hepatocellular Carcinoma. Adv Exp Med Biol 2017;1018:11-21. [Crossref] [PubMed]
- Hilmi M, Vienot A, Rousseau B, et al. Immune Therapy for Liver Cancers. Cancers (Basel) 2019;12:77. [Crossref] [PubMed]
- Gajewski TF, Corrales L, Williams J, et al. Cancer Immunotherapy Targets Based on Understanding the T Cell-Inflamed Versus Non-T Cell-Inflamed Tumor Microenvironment. Adv Exp Med Biol 2017;1036:19-31. [Crossref] [PubMed]
- Xie L, Yang Y, Meng J, et al. Cationic polysaccharide spermine-pullulan drives tumor associated macrophage towards M1 phenotype to inhibit tumor progression. Int J Biol Macromol 2019;123:1012-9. [Crossref] [PubMed]
- Wang H, Wang X, Li X, et al. CD68(+)HLA-DR(+) M1-like macrophages promote motility of HCC cells via NF-κB/FAK pathway. Cancer Lett 2014;345:91-9. [Crossref] [PubMed]
- Hudson SV, Miller HA, Mahlbacher GE, et al. Computational/experimental evaluation of liver metastasis post hepatic injury: interactions with macrophages and transitional ECM. Sci Rep 2019;9:15077. [Crossref] [PubMed]
- Antimisiaris SG, Mourtas S, Marazioti A. Exosomes and Exosome-Inspired Vesicles for Targeted Drug Delivery. Pharmaceutics 2018;10:218. [Crossref] [PubMed]
- Li Y, Liu Y, Xiu F, et al. Characterization of exosomes derived from Toxoplasma gondii and their functions in modulating immune responses. Int J Nanomedicine 2018;13:467-77. [Crossref] [PubMed]
- Jang JY, Lee JK, Jeon YK, et al. Exosome derived from epigallocatechin gallate treated breast cancer cells suppresses tumor growth by inhibiting tumor-associated macrophage infiltration and M2 polarization. BMC Cancer 2013;13:421. [Crossref] [PubMed]
- Amaral EP, Costa DL, Namasivayam S, et al. A major role for ferroptosis in Mycobacterium tuberculosis-induced cell death and tissue necrosis. J Exp Med 2019;216:556-70. [Crossref] [PubMed]
- Wang Y, Wei Z, Pan K, et al. The function and mechanism of ferroptosis in cancer. Apoptosis 2020;25:786-98. [Crossref] [PubMed]
- Dai E, Han L, Liu J, et al. Autophagy-dependent ferroptosis drives tumor-associated macrophage polarization via release and uptake of oncogenic KRAS protein. Autophagy 2020;16:2069-83. [Crossref] [PubMed]
- Nemade H, Chaudhari U, Acharya A, et al. Cell death mechanisms of the anti-cancer drug etoposide on human cardiomyocytes isolated from pluripotent stem cells. Arch Toxicol 2018;92:1507-24. [Crossref] [PubMed]
- Zhang K, Wu L, Zhang P, et al. miR-9 regulates ferroptosis by targeting glutamic-oxaloacetic transaminase GOT1 in melanoma. Mol Carcinog 2018;57:1566-76. [Crossref] [PubMed]
- Wang W, Green M, Choi JE, et al. CD8+ T cells regulate tumour ferroptosis during cancer immunotherapy. Nature 2019;569:270-4. [Crossref] [PubMed]
- Maeda S, Kamata H, Luo JL, et al. IKKbeta couples hepatocyte death to cytokine-driven compensatory proliferation that promotes chemical hepatocarcinogenesis. Cell 2005;121:977-90. [Crossref] [PubMed]
- Chen K, Wu Z, Zang M, et al. Immunization with glypican-3 nanovaccine containing TLR7 agonist prevents the development of carcinogen-induced precancerous hepatic lesions to cancer in a murine model. Am J Transl Res 2018;10:1736-49. [PubMed]
- Shi A, Zhang X, Xiao F, et al. CD56bright natural killer cells induce HBsAg reduction via cytolysis and cccDNA decay in long-term entecavir-treated patients switching to peginterferon alfa-2a. J Viral Hepat 2018;25:1352-62. [Crossref] [PubMed]
- Han M, Li Y, Wu W, et al. Altered expression of interferon-stimulated genes is strongly associated with therapeutic outcomes in hepatitis B virus infection. Antiviral Res 2017;147:75-85. [Crossref] [PubMed]
- Wu D, Wang P, Han M, et al. Sequential combination therapy with interferon, interleukin-2 and therapeutic vaccine in entecavir-suppressed chronic hepatitis B patients: the Endeavor study. Hepatol Int 2019;13:573-86. [Crossref] [PubMed]
- Abadjian MZ, Edwards WB, Anderson CJ. Imaging the Tumor Microenvironment. Adv Exp Med Biol 2017;1036:229-57. [Crossref] [PubMed]
- Goswami KK, Ghosh T, Ghosh S, et al. Tumor promoting role of anti-tumor macrophages in tumor microenvironment. Cell Immunol 2017;316:1-10. [Crossref] [PubMed]
- Zang M, Li Y, He H, et al. IL-23 production of liver inflammatory macrophages to damaged hepatocytes promotes hepatocellular carcinoma development after chronic hepatitis B virus infection. Biochim Biophys Acta Mol Basis Dis 2018;1864:3759-70. [Crossref] [PubMed]
- Théry C, Zitvogel L, Amigorena S. Exosomes: composition, biogenesis and function. Nat Rev Immunol 2002;2:569-79. [Crossref] [PubMed]
- Yang Y, Han Q, Hou Z, et al. Exosomes mediate hepatitis B virus (HBV) transmission and NK-cell dysfunction. Cell Mol Immunol 2017;14:465-75. [Crossref] [PubMed]
- Fang T, Lv H, Lv G, et al. Tumor-derived exosomal miR-1247-3p induces cancer-associated fibroblast activation to foster lung metastasis of liver cancer. Nat Commun 2018;9:191. [Crossref] [PubMed]
- Liu J, Fan L, Yu H, et al. Endoplasmic Reticulum Stress Causes Liver Cancer Cells to Release Exosomal miR-23a-3p and Up-regulate Programmed Death Ligand 1 Expression in Macrophages. Hepatology 2019;70:241-58. [Crossref] [PubMed]
- Lack JB, Weider LJ, Jeyasingh PD. Whole genome amplification and sequencing of a Daphnia resting egg. Mol Ecol Resour 2018;18:118-27. [Crossref] [PubMed]
- Chaput N, Théry C. Exosomes: immune properties and potential clinical implementations. Semin Immunopathol 2011;33:419-40. [Crossref] [PubMed]
- Lin F, Yin HB, Li XY, et al. Bladder cancer cell secreted exosomal miR 21 activates the PI3K/AKT pathway in macrophages to promote cancer progression. Int J Oncol 2020;56:151-64. [PubMed]
- Stockwell BR, Friedmann Angeli JP, Bayir H, et al. Ferroptosis: A Regulated Cell Death Nexus Linking Metabolism, Redox Biology, and Disease. Cell 2017;171:273-85. [Crossref] [PubMed]
- Matsushita M, Freigang S, Schneider C, et al. T cell lipid peroxidation induces ferroptosis and prevents immunity to infection. J Exp Med 2015;212:555-68. [Crossref] [PubMed]
- Zhang F, Li F, Lu GH, et al. Engineering Magnetosomes for Ferroptosis/Immunomodulation Synergism in Cancer. ACS Nano 2019;13:5662-73. [Crossref] [PubMed]
- Li C, Chen H, Lan Z, et al. mTOR-dependent upregulation of xCT blocks melanin synthesis and promotes tumorigenesis. Cell Death Differ 2019;26:2015-28. [Crossref] [PubMed]
- Zhang Y, Shi J, Liu X, et al. BAP1 links metabolic regulation of ferroptosis to tumour suppression. Nat Cell Biol 2018;20:1181-92. [Crossref] [PubMed]
- Luo M, Wu L, Zhang K, et al. miR-137 regulates ferroptosis by targeting glutamine transporter SLC1A5 in melanoma. Cell Death Differ 2018;25:1457-72. [Crossref] [PubMed]
- Pistritto G, Trisciuoglio D, Ceci C, et al. Apoptosis as anticancer mechanism: function and dysfunction of its modulators and targeted therapeutic strategies. Aging (Albany NY) 2016;8:603-19. [Crossref] [PubMed]
- Olechowska-Jarząb A, Ptak-Belowska A, Brzozowski T. Terapeutic importance of apoptosis pathways in pancreatic cancer. Folia Med Cracov 2016;56:61-70. [PubMed]
- Binnewies M, Roberts EW, Kersten K, et al. Understanding the tumor immune microenvironment (TIME) for effective therapy. Nat Med 2018;24:541-50. [Crossref] [PubMed]
(English Language Editor: R. Scott)


