
The 1q21.3 region driver gene EFNA3 promotes disease progression via inhibition of lung adenocarcinoma cell apoptosis
Introduction
Lung cancer has the highest incidence and mortality rates among malignant tumors worldwide. As the most common pathological type of lung cancer, lung adenocarcinoma (LUAD) accounts for 80–85% of all lung cancer cases (1), placing a heavy economic burden on patients, families, and even society. Many researchers have attempted to identify the key genes involved in LUAD progression, so as to serve as molecular targets for the treatment and inhibition of disease progression in LUAD patients (2). However, as determined through advancements in research, there are a large number of differentially-expressed genes (DEGs) involved in LUAD progression. Identifying the extent to which DEGs are involved in disease progression has also become a particular focus of research. To understand the effects of DEGs on disease progression, researchers have proposed the concepts of “driver genes” and “passenger genes” (3,4). Driver genes refer to DEGs that promote cancer development and progression, while passenger genes refer to genes that do not significantly promote cancer development and progression. On this basis, effectively identifying and verifying the driver genes in LUAD patients has become an important aspect of lung cancer research (5).
It is possible to identify the driver genes in LUAD patients by analyzing the gene copy number variations (4). In rectal cancer, driver genes have been identified by screening the gene copy number variations coupled with the DEGs (6). Furthermore, a study has shown that the combination of copy number and DEGs can effectively predict the classification and drug resistance of cancers (7). However, variation in the gene copy number in the entire chromosome of cancer patients is relatively high, and there are still some passenger genes among the genes that intersect with DEGs (8). Therefore, it is necessary to further narrow the scope of screening for driver genes based on the gene copy number variation analysis.
The amplification of some regions of chromosomes is the main cause of an increase in copy number. For instance, amplification of the 9p24.1 region in triple-negative breast cancer significantly increases the copy number of immune-related genes, thereby promoting disease progression in patients (9). Also, amplification of the 1q21 region in patients with multiple myeloma is closely related to the clinical stage of the disease. Therefore, the driver genes in LUAD can be identified by determining the amplification region of specific chromosomes in LUAD patients and further clarifying the genes with copy number variations in the region (10). An existing study has confirmed that 1q21.3 is an amplified region in LUAD patients (11). Based on the above studies, there may be driver genes among copy variant genes amplified in the 1q21.3 region in LUAD patients.
With the development of genomics, LUAD driver genes can be preliminarily screened through The Cancer Genome Atlas (TCGA) database. This database collects clinical data and compiles genome and copy number variation data for each gene in various human cancers, and is a reliable database for screening LUAD driver genes. On this basis, the driver genes in LUAD patients were preliminarily screened in this study using TCGA database, and were subsequently analyzed and verified. We present the following article in accordance with the MDAR reporting checklist (available at https://tcr.amegroups.com/article/view/10.21037/tcr-22-979/rc).
Methods
Bioinformatics analysis
Acquisition of LUAD tissue
The transcriptome data, coefficient of variation (CV) data for copy number, and follow-up data for LUAD patients were obtained from TCGA database. LUAD cases with complete data from 1991 to 2013 and confirmed by pathological examination were screened. Additionally, corresponding data for a normal control group were obtained. The study was conducted in accordance with the Declaration of Helsinki (as revised in 2013).
Screening of LUAD driver genes
The edgeR package in R 4.0.3 (https://cran.r-project.org/web/packages/ROCR/index.html) was used to standardize the transcriptomics results obtained from TCGA database, and the threshold for DEGs was set to |log2FC| =1, P=0.01. The obtained CV data for gene copy number were annotated to obtain data for genes with a CV >10% in the 1q21.3 region. Using the follow-up outcomes of LUAD patients, Kaplan-Meier (KM) survival analysis was performed to obtain DEGs related to LUAD patient survival. The intersection of the above three datasets was used to obtain the driver genes in LUAD patients.
Mechanisms of action of driver genes
After obtaining the driver genes in LUAD patients, the LUAD patients were divided into a high expression group and a low expression group according to the optimal cutoff value for driver gene expression. KM survival analysis was conducted using the follow-up outcomes of the two groups of patients to determine the impact of driver gene expression levels on LUAD patient survival. Kyoto Encyclopedia of Genes and Genomes (KEGG) analysis was used to determine the main mechanisms of action of LUAD driver genes.
Cell culture and grouping
In this experiment, the human LUAD cell lines, A549 and PC9, were selected as the study objects (purchased from the Stem Cell Bank of the Chinese Academy of Sciences). Cells in each group were cultured in Roswell Park Memorial Institute (RPMI)-1640 medium containing 10% fetal bovine serum (FBS) and 1% penicillin-streptomycin in a constant temperature incubator at 37 ℃ with 5% carbon dioxide(CO2). The medium was changed every day, and cell morphology was observed under an inverted microscope (Jeica MICROSYSTEM, Geman). Cells were passaged when they were approximately 80% confluent.
A549 and PC9 cells were divided into a control (negative control group), siRNA-control group (siEFNA3-negative control), and siRNA-ephrin-A3 (EFNA3) group (siEFNA3).
Establishment of a lentivirus knockdown cell line
Evaluation of lentivirus infection efficiency
In this experiment, A459 and PC9 cells were used as the study objects. EFNA3 siRNA was designed by Gene Pharma Inc., China. The specific EFNA3 siRNA sequences are provided in Table 1. The cells were cultured in a constant temperature incubator, and were harvested and seeded into six-well plates when they were approximately 80% confluent. The cells were cultured in a 37 ℃ constant temperature incubator for 24 h. A recombinant vector carrying siRNA was transfected into A459 and PC9 cells. Two micrograms of recombinant vector, 1.5 µL of PsPAX2 (lentiviral packaging plasmids) solution, and 0.6 µg of PMD2G (lentiviral packaging plasmids) were added to a 1.5-mL EP (eppendorf) tube, after which 400 µL of diluent was added. The solution was mixed well, and 8 µL of jetPRIME transfection reagent was added. This solution was subsequently mixed well at room temperature, allowed to stand for 10 min, and then added to the A459 and PC9 cells seeded in the six-well plate. The cells in each group were incubated at 37 ℃ for 24 h. The medium containing lentivirus was discarded. The efficiency of lentivirus infection was observed under a fluorescence microscope (Jeica MICROSYSTEM, Gemany).
Table 1
| siRNA | siRNA sequence |
|---|---|
| siEFNA3-1 | 5'-GGATTGTGGTTTGGATTGAAA-3' |
| siEFNA3-2 | 5'-GGCATGTACAGACTCTATATC-3' |
| siEFNA3-NC | 5'-GCTTCGCGCCGTAGTCTTA-3' |
Construction of the siEFNA3 cell line
A459 and PC9 cells were cultured in a constant temperature incubator, and were harvested and seeded into 6-well plates when they were approximately 80% confluent. When the cells were approximately 60–70% confluent, lentiviruses were diluted at a ratio of 1:10 in RPMI-1640 medium containing 10% FBS. After the culture medium in the six-well plate was discarded, 1 mL of the lentivirus solution was added to each well. The culture medium containing the lentivirus was discarded after 24 h of incubation in a constant temperature incubator at 37 ℃. Two milliliters of RPMI-1640 medium containing 10% FBS was added, and the cells continued to be cultured in a constant temperature incubator.
3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide (MTT) assay
A459 and PC9 cells were trypsinized and resuspended when they were approximately 80% confluent. The resuspended cells in each group were seeded in a 96-well plate at a density of 5×104 cells/well and continued to be cultured in a constant temperature incubator. The MTT assay was conducted at 0, 24, 48, and 72 h of culturing. After adding 20 µL of 5 g/L MTT solution to each well, the cells were cultured for 4 h, and then 100 µL of 10% sodium dodecyl sulfate (SDS) solution was added to each well. The OD (optical density) value of each group of cells at 490 nm was recorded.
Enzyme-linked immunosorbent assay (ELISA)
Cell proliferation
A459 and PC9 cells were trypsinized and resuspended when they were approximately 80% confluent. The resuspended cells in each group were seeded in a 96-well plate at a density of 5×104 cells/well. Twenty microliters of BrdU (5-bromo-2-deoxyuridine) labeling solution was added to each well, and the cells were cultured for 48 h. The proliferation of cells in each group was detected by ELISA, and the OD values of the cells at 450 nm were recorded.
Apoptosis
A459 and PC9 cells were trypsinized and resuspended when they were approximately 80% confluent. The resuspended cells in each group were seeded in a 12-well plate at a density of 5×104 cells/well. After culturing for 48 h, phosphate buffered saline (PBS) was used to wash the cells, and the supernatant was discarded. Two hundred microliters of lysis buffer was added to each well, and the cells were incubated for 20 min. The supernatant was obtained after centrifugation at 200 r/min for 10 min. A total of 20 µL of supernatant was added to each well, and 80 µL of premixed anti-DNA-POD (peroxidase horseradish, anti-histone-biotin, and culture medium were added, followed by centrifugation at 300 r/min for 30 min. One hundred microliters of ABTS (2,2-azino-bis-3-ethvlbenzothiaz0line-6-sulfonic acid) solution was added in the dark. After centrifugation at 150 r/min for 15 min, 100 µL of ABTS stop solution was added. The OD value of each group at 405 nm was recorded.
Reverse transcription polymerase chain reaction (RT-PCR)
A459 and PC9 cells were trypsinized, centrifuged, and harvested after 120 hours of treatment. Total RNA was extracted using the TRIzol method. The concentration and purity of RNA were measured in a microspectrophotometer (HACH, China) using 2 µL of dissolution liquid. To reverse transcribe RNA into cDNA (complementary DNA), a reverse transcription reaction system was prepared using a Takara reverse transcription kit. The cDNA reaction system was prepared using a qPCR kit. The reaction was performed at 95 ℃ for 10 min, 95 ℃ for 30 s, and 60 ℃ for 30 s, with 40 amplification cycles. The target genes were EFNA3, Bax, and caspase3, and the internal reference gene was glyceraldehyde-3-phosphate dehydrogenase (GADPH). The primer sequences are provided in Table 2. Additionally, the melting curve from 60 to 95 ℃ was analyzed. The messenger RNA (mRNA) expression levels of EFNA3, Bax, and caspase3 in each sample were statistically analyzed using the 2−∆∆Ct method.
Table 2
| Gene | Primers |
|---|---|
| EFNA3 | F: 5'-AACCGGCATGCGGTGTA-3' |
| R: 5'-ATCCAGATAGTCGTTCACGTTCA-3' | |
| Bax | F: 5'-TTGCTACAGGGTTTCATCC-3' |
| R: 5'-GTCCAGTTCATCGCCAAT-3' | |
| Caspase 3 | F: 5'-CGTGTCATAAAATACCAGTGGA-3' |
| R: 5'-AAATTCTGTTGCCACCTTTCG-3' | |
| GADPH | F: 5'-AGAAGGCTGGGGCTCATTTG-3' |
| R: 5'-AGGGGCCATCCACAGTCTTC-3' |
PCR, polymerase chain reaction.
Western blot
A459 and PC9 cells were trypsinized, centrifuged, and harvested after 120 h of treatment. The cell contents were collected after cell lysis, and total soluble protein was collected by centrifugation. The protein concentration was determined using the BCA (bicinchoninic acid) method. After the protein concentration was determined, sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE) was performed for each tissue/cell. Following electrophoresis, the proteins separated by electrophoresis were transferred to polyvinylidene fluoride (PVDF) membranes using a transfer device (millipore, USA). After transfer, the PVDF membrane was removed from the transfer device, blocked with 5% skim milk for 1 hour, and rinsed with PBS. Next, primary antibodies against EFNA3 (1:500), Bax (1:500), and caspase3 (1:500) were added, and the membranes were incubated at 4 ℃ overnight. Subsequently, a secondary goat anti-rabbit antibody (1:1,000) was added, and the membranes were incubated for 1 h. A chemiluminescence imaging system (Thermo Fisher SCIENTIFIC, USA) was used to expose and visualize the protein on the PVDF film.
Reagent information
The following reagents were used in this study: RPMI-1640 culture medium (Gibco, USA, catalog No. 31800022), FBS (HyClone, USA, catalog No. SH30396.03), PBS solution containing double antibodies (HyClone, USA, catalog No. SH30256.01), reverse transcription kit (Takara, Japan, catalog No. RR047A), qPCR kit (Takara, Japan, catalog No. RR430B), ELISA-BrdU kit (Roche, Switzerland, catalog No. 11647229001), ELISA-apoptosis kit (Roche, Switzerland, catalog No. 11774425001), rabbit anti-human Bax antibody (1:1,000, FineTest, Wuhan, China, catalog: FNab00811), rabbit anti-human EFNA3 antibody (1:1,000, FineTest, Wuhan, China, catalog No. FNab08194), and rabbit anti-human caspase 3 antibody (1:1,000, Abcam, USA, catalog No. ab44976).
Statistical analysis
Statistical analysis was performed using R 4.0.3 and SPSS 26.0 (https://www.ibm.com/support/pages/downloading-ibm-spss-statistics-26?mhsrc=ibmsearch_a&mhq=SPSS%2026.0%20). For data mining, the edgeR package of R software was used for the Bayesian analysis. KM survival analysis was performed using the survival package in R software. For basic experiments, SPSS 26.0 was used for the corresponding statistical analysis. The obtained data were expressed as the mean ± standard deviation (mean ± SD). Comparison between multiple groups was performed using analysis of variance (ANOVA). Indicators with statistically significant ANOVA results were subjected to LSD (least significance difference) post hoc tests. P<0.05 was considered statistically significant.
Results
Bioinformatics analysis preliminarily elucidated LUAD driver genes and their mechanisms in LUAD
After searching and screening TCGA database, a total of 483 LUAD samples and 59 normal control samples were obtained. The DEG analysis revealed 640 upregulated genes. A total of 154 genes with a CV >10% were identified when screening the chromosome 1q23.1 region. KM analysis revealed 5,968 genes significantly associated with patient OS. A Venn diagram suggested that HORMAD1, EFNA3, IVL, and SEFNA3 may be driver genes of LUAD (Figure 1A). The expression levels of HORMAD1, EFNA3, IVL, and SEFNA3 in the control samples and LUAD samples were extracted from TCGA database and compared. The results indicated that EFNA3 expression was significantly higher in LUAD tissues than in control tissues and significantly higher than the expression of the other three genes (P<0.05, Figure 1B; box plots of HORMAD1, IVL, and SEFNA3 are shown in Figure S1).
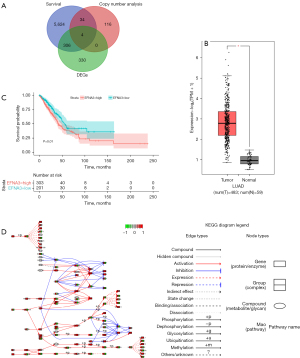
Using the optimal cutoff of EFNA3 expression, the LUAD samples were divided into high and low expression groups. The KM curve indicated that the OS of patients in the low EFNA3 expression group was significantly higher than that of patients in the high expression group (P<0.05) (Figure 1C). The results of the KEGG enrichment analysis of the DEGs in the two groups of patients indicated enrichment in apoptosis-related pathways (Figure 1D). The bioinformatics analysis preliminarily confirmed that EFNA3 was a driver gene in LUAD.
Effect of EFNA3 on LUAD cell proliferation
After infecting A459 and PC9 cells with siEFNA3 lentivirus, the MTT assay indicated that the viability of A459 and PC9 cells decreased markedly (P<0.05, Figure 2A,2B). The BrdU-ELISA results indicated that the proliferation of A459 and PC9 cells was significantly decreased after EFNA3 knockdown (P<0.05, Figure 2C,2D). The RT-PCR results showed that EFNA3 mRNA expression in each cell was notably reduced (Figure 2E,2F). The above results suggest that the knockdown of EFNA3 expression in LUAD cells significantly inhibits cell proliferation.
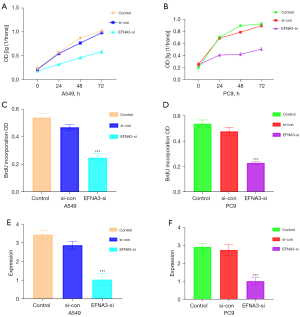
The effect of EFNA3 on LUAD cell apoptosis:
After EFNA3 knockdown in A549 and PC9 cells, apoptosis was markedly increased, as determined by ELISA (P<0.05, Figure 3A,3B). The RT-PCR results indicated that the mRNA expression of Bax and caspase3 increased considerably (P<0.05, Figure 3C-3F). The above results suggest that apoptosis significantly increased after EFNA3 knockdown.
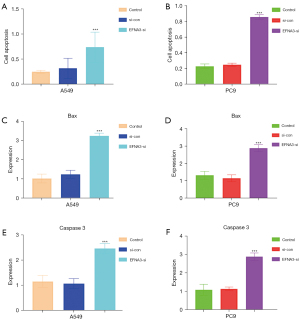
The effect of EFNA3 on the expression of apoptosis-related proteins in LUAD cells
The expression of apoptosis-related proteins (Bax and caspase 3) in the cells of each group was detected by western blot and subsequently analyzed. The results indicated that the expression levels of apoptosis-related proteins in LUAD cells (A549 and PC9) after EFNA3 knockdown were significantly lower than those in the normal control group (P<0.05, Figure 4).
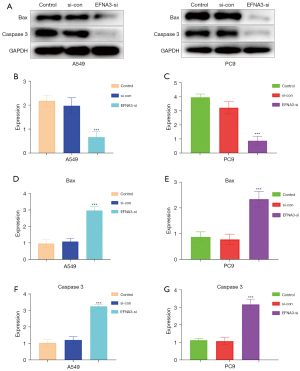
Discussion
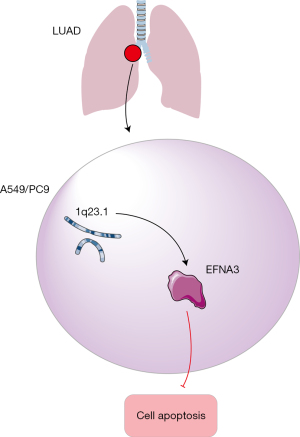
In this study, we used bioinformatics analysis to preliminarily screen EFNA3 as a driver gene for LUAD, concluding that EFNA3 regulates the progression of LUAD by inhibiting apoptosis. The results obtained through cell-based experiments indicated that after EFNA3 knockdown in LUAD cells, cell viability was significantly decreased, proliferation was markedly inhibited, and apoptosis was notably increased (Figure 5).
LUAD patients have a poor prognosis and high mortality. Eighty percent of LUAD patients have progressed to the middle and advanced stage at the time of diagnosis, which seriously affects the prognosis of LUAD patients. With the in-depth research on the diagnosis and treatment of LUAD, the research and application of targeted drugs have significantly improved the prognosis of patients. Epidermal growth factor receptor (EGFR)-TKI, as an epidermal growth factor-targeted drug, can significantly improve the prognosis of patients. A study has shown that the objective response rate of LUAD patients reached 56.3% after EGFR-TKI treatment (12). Other targeted drugs such as PD-1, PD-L1 immune checkpoint inhibitors, VEGF inhibitors and other targeted drugs have shown excellent clinical application value (13,14). However, with the application of various targeted drugs, LUAD will further mutate and become resistant. Therefore, it is necessary to further clarify the targets that have the advantages of targeted therapy for LUAD, so as to provide an effective clinical treatment for LUAD.
In this study, the driver genes of LUAD were determined by identifying the genes that intersect among LUAD DEGs, genes with high copy number variations in the 1q21.3 region, and DEGs obtained by KM analysis. Changes in copy number CV in tumor driver genes are the main genetic variation patterns in the process of tumor formation and development (15).
Copy variation refers to the variation in DNA fragments larger than 1 kb. Variation modes include deletion, duplication, inversion, and ectopic variation. Therefore, the occurrence of copy variation significantly enriches the diversity of genetic variation in genomes (16). The occurrence of copy variation is also closely related to the occurrence and development of tumors. In neuroblastoma, copy variation in the 1q21.1 region is highly correlated with tumor development (17). In breast cancer patients, copy variation in the human epidermal growth factor receptor-2 (HER2) gene on chromosome 17 promotes the development and progression of breast cancer (18). Additionally, EGFR gene copy variation amplification on the short arm of chromosome 7 is closely related to the occurrence of gastrointestinal cancer and non-small cell lung cancer (19). A previous study has also confirmed that the amplification of the 1q21.3 region is closely related to tumor progression (20). Furthermore, amplification of the interleukin-2 enhancer binding factor 2 (ILF2) gene in the 1q21.3 region is closely related to the invasiveness of skin melanoma (21), and amplification of the Jun Proto-Oncogene (JUN) gene in the 1q21.3 region is closely related to the occurrence and development of thyroid cancer (22). Moreover, a study has also shown that pi4kii β amplification in the 1q21.3 region is closely associated with LUAD (9). In this study, the copy number coefficient of variation (CV) of the genes in the 1q21.3 region was screened, and the statistical analysis indicated that ENFA3 was an LUAD driver gene.
EFNA3 is located in the 1q21.3 region of the human chromosome (23). The epidermal differentiation complex gene cluster is located in this region. Previous studies have shown that the EFNA3 gene primarily regulates the development of skin keratinocytes, and EFNA3 is also involved in lymphocyte chemotaxis and glutamate translocation (24,25). With advancements in research, EFNA3 has been shown to play an important role in the regulation of malignant tumors (26). A study has shown that high EFNA3 expression in breast cancer suggests a poor prognosis (27). In oral cancer, EFNA3 is mainly involved in the regulation of tumor angiogenesis (28). A study has also shown that high EFNA3 expression can promote the migration, invasion, and proliferation of gastric cancer cells (29). However, whether EFNA3 has a regulatory effect on tumor cell function in LUAD patients has not yet been confirmed.
On this basis, the present study used a lentiviral infection technique to knock down the EFNA3 gene in A459 and PC9 cells. Cell functions were assessed following RT-PCR verification of the EFNA3 gene knockdown efficiency. The MTT assay results indicated that the viability of A459 and PC9 cells was significantly decreased after EFNA3 knockdown. In addition, the ELISA results also showed that the proliferation of A459 and PC9 cells was markedly reduced. On this basis, our team believes that EFNA3 is a driver gene for LUAD. However, the mechanism through which EFNA3 regulates the functions of LUAD cells has not been elucidated.
To clarify the specific molecular mechanism by which EFNA3 regulates the functional changes in LUAD cells, this study divided the patients into high group and low expression groups based on the optimal cutoff values for EFNA3 expression in LUAD patients included in TCGA database. The DEGs of the two groups of patients were comparatively analyzed, and were subjected to KEGG enrichment pathway analysis. The results indicated that EFNA3 regulated the functional changes in LUAD cells mainly by regulating the expression of apoptosis-related pathways. Apoptosis refers to programmed cell death (30,31). The inhibition of apoptosis in LUAD tissues leads to infinite tumor cell proliferation. Multiple previous studies have attempted to effectively control the course of LUAD by promoting tumor cell apoptosis (32,33). A study has shown that anlotinib can effectively improve the sensitivity of A549 LUAD cells to radiotherapy, thereby promoting LUAD cell apoptosis and ultimately controlling of the disease course of LUAD (34). Cisplatin effectively treats LUAD by inducing A549 cell apoptosis (35). The above studies have shown that promoting tumor cell apoptosis is an effective means of treating and controlling the progression of LUAD.
Yet, the current treatment methods of promoting cell apoptosis have certain limitations. For example, cisplatin promotes tumor cell apoptosis by binding to intracellular DNA, thereby treating LUAD. However, tumor cells exposed to cisplatin treatment can activate adaptive resistance mechanisms, thereby resulting in treatment failure and tumor recurrence events (36). Therefore, it is necessary to identify new therapeutic targets that promote LUAD cell apoptosis. The results of the bioinformatics analysis in this study showed that EFNA3 may regulate LUAD progression by regulating tumor cell apoptosis. On this basis, EFNA3 might be a key target for promoting LUAD cell apoptosis. In this study, cell apoptosis was assessed by ELISA after EFNA3 knockdown. The results showed that apoptosis significantly increased after EFNA3 knockdown. The expression of apoptosis-related genes (Bax and caspase3) was assessed by RT-PCR and western blot. The results showed that the expression of apoptosis-related genes (Bax and caspase3) decreased considerably after EFNA3 knockdown. Therefore, we believe that EFNA3 is a driver gene in LUAD progression and that it can promote the progression of LUAD by inhibiting tumor cell apoptosis. This study will provide a new therapeutic target for effective clinical treatment of LUAD and control the progression of LUAD, which has far-reaching clinical research significance.
Acknowledgments
Funding: This study was funded by
Footnote
Reporting Checklist: The authors have completed the MDAR reporting checklist. Available at https://tcr.amegroups.com/article/view/10.21037/tcr-22-979/rc
Data Sharing Statement: Available at https://tcr.amegroups.com/article/view/10.21037/tcr-22-979/dss
Conflicts of Interest: All authors have completed the ICMJE uniform disclosure form (available at https://tcr.amegroups.com/article/view/10.21037/tcr-22-979/coif). All authors report that this study was funded by the Qiqihar Science and Technology Innovation Incentive Project (No. CSFFG-2021330). The authors have no other conflicts of interest to declare.
Ethical Statement: The authors are accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved. The study was conducted in accordance with the Declaration of Helsinki (as revised in 2013).
Open Access Statement: This is an Open Access article distributed in accordance with the Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License (CC BY-NC-ND 4.0), which permits the non-commercial replication and distribution of the article with the strict proviso that no changes or edits are made and the original work is properly cited (including links to both the formal publication through the relevant DOI and the license). See: https://creativecommons.org/licenses/by-nc-nd/4.0/.
References
- Subbarayan K, Ulagappan K, Wickenhauser C, et al. Expression and Clinical Significance of SARS-CoV-2 Human Targets in Neoplastic and Non-Neoplastic Lung Tissues. Curr Cancer Drug Targets 2021;21:428-42. [Crossref] [PubMed]
- Li Y, Gu J, Xu F, et al. Molecular characterization, biological function, tumor microenvironment association and clinical significance of m6A regulators in lung adenocarcinoma. Brief Bioinform 2021;22:bbaa225.
- Zhang X, Yang H, Zhang J, et al. HSD17B4, ACAA1, and PXMP4 in Peroxisome Pathway Are Down-Regulated and Have Clinical Significance in Non-small Cell Lung Cancer. Front Genet 2020;11:273. [Crossref] [PubMed]
- Chang S, Yim S, Park H. The cancer driver genes IDH1/2, JARID1C/ KDM5C, and UTX/ KDM6A: crosstalk between histone demethylation and hypoxic reprogramming in cancer metabolism. Exp Mol Med 2019;51:1-17. [Crossref] [PubMed]
- Dietlein F, Weghorn D, Taylor-Weiner A, et al. Identification of cancer driver genes based on nucleotide context. Nat Genet 2020;52:208-18. [Crossref] [PubMed]
- Halaburkova A, Cahais V, Novoloaca A, et al. Pan-cancer multi-omics analysis and orthogonal experimental assessment of epigenetic driver genes. Genome Res 2020;30:1517-32. [Crossref] [PubMed]
- Wang G, Wang Q, Liang N, et al. Oncogenic driver genes and tumor microenvironment determine the type of liver cancer. Cell Death Dis 2020;11:313. [Crossref] [PubMed]
- Molina-Sánchez P, Ruiz de Galarreta M, Yao MA, et al. Cooperation Between Distinct Cancer Driver Genes Underlies Intertumor Heterogeneity in Hepatocellular Carcinoma. Gastroenterology 2020;159:2203-2220.e14. [Crossref] [PubMed]
- Shi L, Tan X, Liu X, et al. Addiction to Golgi-resident PI4P synthesis in chromosome 1q21.3-amplified lung adenocarcinoma cells. Proc Natl Acad Sci U S A 2021;118:e2023537118. [Crossref] [PubMed]
- Luo G, Tang M, Zhao Q, et al. Bone marrow adipocytes enhance osteolytic bone destruction by activating 1q21.3(S100A7/8/9-IL6R)-TLR4 pathway in lung cancer. J Cancer Res Clin Oncol 2020;146:2241-53. [Crossref] [PubMed]
- Goh JY, Feng M, Wang W, et al. Chromosome 1q21.3 amplification is a trackable biomarker and actionable target for breast cancer recurrence. Nat Med 2017;23:1319-30. [Crossref] [PubMed]
- Zhang Q, Liu M, Zhong Y, et al. Long-term survival in a patient with advanced non-smoking lung adenocarcinoma and no EGFR mutation after comprehensive therapy with EGFR-TKI-based therapy A case report. Medicine 2022;101:e28583. [PubMed]
- Duan JC, Wang ZJ, Lin L, et al. Apatinib, a novel VEGFR inhibitor plus docetaxel in advanced lung adenocarcinoma patients with wild-type EGFR: a phase I trial. Invest New Drugs 2019;37:731-7. [Crossref] [PubMed]
- Huang D, Cui P, Huang Z, et al. Anti-PD-1/L1 plus anti-angiogenesis therapy as second-line or later treatment in advanced lung adenocarcinoma. J Cancer Res Clin Oncol 2021;147:881-91. [Crossref] [PubMed]
- Bailey MH, Tokheim C, Porta-Pardo E, et al. Comprehensive Characterization of Cancer Driver Genes and Mutations. Cell 2018;173:371-385.e18. [Crossref] [PubMed]
- Xu Y, Wu G, Li J, et al. Screening and Identification of Key Biomarkers for Bladder Cancer: A Study Based on TCGA and GEO Data. Biomed Res Int 2020;2020:8283401. [Crossref] [PubMed]
- Wu M, Shang X, Sun Y, et al. Integrated analysis of lymphocyte infiltration-associated lncRNA for ovarian cancer via TCGA, GTEx and GEO datasets. PeerJ 2020;8:e8961. [Crossref] [PubMed]
- Xu M, Li Y, Li W, et al. Immune and Stroma Related Genes in Breast Cancer: A Comprehensive Analysis of Tumor Microenvironment Based on the Cancer Genome Atlas (TCGA) Database. Front Med (Lausanne) 2020;7:64. [Crossref] [PubMed]
- Lv J, Zhu Y, Ji A, et al. Mining TCGA database for tumor mutation burden and their clinical significance in bladder cancer. Biosci Rep 2020;40:BSR20194337. [Crossref] [PubMed]
- Tomczak K, Czerwińska P, Wiznerowicz M. The Cancer Genome Atlas (TCGA): an immeasurable source of knowledge. Contemp Oncol (Pozn) 2015;19:A68-77. [Crossref] [PubMed]
- Zhang X, Bustos MA, Gross R, et al. Interleukin enhancer-binding factor 2 promotes cell proliferation and DNA damage response in metastatic melanoma. Clin Transl Med 2021;11:e608. [Crossref] [PubMed]
- Trimarchi G, Caraffi SG, Radio FC, et al. Adducted Thumb and Peripheral Polyneuropathy: Diagnostic Supports in Suspecting White-Sutton Syndrome: Case Report and Review of the Literature. Genes (Basel) 2021;12:950. [Crossref] [PubMed]
- Deng M, Tong R, Zhang Z, et al. EFNA3 as a predictor of clinical prognosis and immune checkpoint therapy efficacy in patients with lung adenocarcinoma. Cancer Cell Int 2021;21:535. [Crossref] [PubMed]
- Gong W, Qie S, Huang P, et al. Deletion of long noncoding RNA EFNA3 aggravates hypoxia-induced injury in PC-12 cells by upregulation of miR-101a. J Cell Biochem 2019;120:836-47. [Crossref] [PubMed]
- Wang H, Wang L, Zhou X, et al. OSCC Exosomes Regulate miR-210-3p Targeting EFNA3 to Promote Oral Cancer Angiogenesis through the PI3K/AKT Pathway. Biomed Res Int 2020;2020:2125656. [Crossref] [PubMed]
- Shukla A, Dahiya S, Onteru SK, et al. Differentially expressed miRNA-210 during follicular-luteal transition regulates pre-ovulatory granulosa cell function targeting HRas and EFNA3. J Cell Biochem 2018;119:7934-43. [Crossref] [PubMed]
- Gómez-Maldonado L, Tiana M, Roche O, et al. EFNA3 long noncoding RNAs induced by hypoxia promote metastatic dissemination. Oncogene 2015;34:2609-20. [Crossref] [PubMed]
- Duffy SL, Steiner KA, Tam PP, et al. Expression analysis of the Epha1 receptor tyrosine kinase and its high-affinity ligands Efna1 and Efna3 during early mouse development. Gene Expr Patterns 2006;6:719-23. [Crossref] [PubMed]
- Wang Z, Yin B, Wang B, et al. MicroRNA-210 promotes proliferation and invasion of peripheral nerve sheath tumor cells targeting EFNA3. Oncol Res 2013;21:145-54. [Crossref] [PubMed]
- Miura H, Kusakabe Y, Hashido K, et al. The glossopharyngeal nerve controls epithelial expression of Sprr2a and Krt13 around taste buds in the circumvallate papilla. Neurosci Lett 2014;580:147-52. [Crossref] [PubMed]
- Specht S, Isse K, Nozaki I, et al. SPRR2A expression in cholangiocarcinoma increases local tumor invasiveness but prevents metastasis. Clin Exp Metastasis 2013;30:877-90. [Crossref] [PubMed]
- Mizuguchi Y, Specht S, Lunz JG 3rd, et al. SPRR2A enhances p53 deacetylation through HDAC1 and down regulates p21 promoter activity. BMC Mol Biol 2012;13:20. [Crossref] [PubMed]
- Wang H. MicroRNAs and Apoptosis in Colorectal Cancer. Int J Mol Sci 2020;21:5353. [Crossref] [PubMed]
- Carneiro BA, El-Deiry WS. Targeting apoptosis in cancer therapy. Nat Rev Clin Oncol 2020;17:395-417. [Crossref] [PubMed]
- Warren CFA, Wong-Brown MW, Bowden NA. BCL-2 family isoforms in apoptosis and cancer. Cell Death Dis 2019;10:177. [Crossref] [PubMed]
- Burke PJ. Mitochondria, Bioenergetics and Apoptosis in Cancer. Trends Cancer 2017;3:857-70. [Crossref] [PubMed]
(English Language Editor: A. Kassem)


