
Prognostic value of miRNA-181a in human colorectal cancer evaluated by in situ hybridization
Introduction
Colorectal cancer (CRC) is the fourth most common factor for cancer-related deaths (1,2). Although new anticancer agents and molecular-targeted therapies for CRC have been developed, the prognosis for advanced stage CRC patients is very poor. This is because of the high incidence of tumor recurrence from distal metastasis (3,4). The 5-year survival rate for early stage cancer patients is approximately 80–90%, however, with tumor recurrence after distal metastasis, the survival rate drastically drops to 10–20% (5,6). The dysregulation of miRNAs has been reported to influence carcinogenesis, aggressiveness and metastasis. Hence, research to identify biomarkers involved with the progression of cancer is urgently needed.
MicroRNAs (miRNAs) comprise of a class of small non-coding RNA molecules, usually 20 to 22 nucleotides in length that regulate post-transcriptional processes of several genes. miRNAs regulate protein-coding genes in biological processes, including tumorigenesis and chemoresistance (7). miRNAs negatively regulate gene expression at the post-transcriptional level through the degradation of mRNA and inhibition of translation (8). Abnormal expression of oncogenic miRNAs can lead to tumor occurrence by facilitating tumor proliferation and apoptosis resistance (9). In CRC, it has been reported that dysregulation of miRNAs affects tumorigenesis, invasion and metastasis. MiRNA-143 and miRNA-145 were first identified as dysregulated miRNAs in CRC in 2003 (10). Since then, numerous miRNAs have been studied for their relevance in CRC prognosis. Recently, the higher expression level of miRNA-181a has been demonstrated in a number of cancers. MiRNA-181a has been identified as a prognostic biomarker for acute myeloid leukemia and non-small cell lung cancer (11,12). In addition, miRNA-181a levels have been correlated with cancer chemoresistance. Overexpression of miRNA-181a in prostate cancer cells contributes to tumor resistance to docetaxel and cabazitaxel (13). MiRNA-181a also promotes cancer resistance to radiation therapy by targeting the pro-apoptotic PRKCD gene (14) and modulates glioma tumor growth and chemotherapy resistance to temozolomide (TMZ) through the PTEN pathway (15). However, there have been limited studies regarding the association between miRNA-181a expression levels and the prognosis of CRC.
In our study, we investigated the expression of miRNA-181a using in situ hybridization (ISH) high-throughput platforms and the effect it had on the prognosis of CRC. In addition, we selected CRC patients undergoing chemotherapy to assess the ability of miRNA-181a to affect the prognosis of CRC patients.
Methods
Ethics statement
This study was approved by the Medical Ethics Committee of China Medical University at the First Affiliated Hospital of China Medical University. All patients agreed to the study protocols and signed informed consent forms to participate in the study. All tumor samples were anonymized before analysis using ISH.
Patients and tissue samples
CRC patients were recruited between June 2005 and March 2010 by the First Affiliated Hospital of China Medical University. Exclusion criteria were as follows: (I) incomplete patient medical records and/or no follow-up information; (II) lack of clinical samples for proper evaluation; (III) severe postoperative complications including intestinal fistulas and anastomotic obstructions; (IV) preoperative adjuvant chemotherapy and radiotherapy.
Tumor tissue and surrounding intestinal mucosa were collected during the surgery. The clinical staging was performed in line with the WHO and TNM staging guidelines (16). Based on tumor differentiation, patients were classified into grade 1 (poorly differentiated), grade 2 (medium differentiated) and grade 3 (well differentiated). All patients underwent enhanced CT examination of the pulmonary and abdominal regions prior to surgery. Patients received routine CT and colonoscopies during follow-up to detect efficacy, invasion and/or recurrence of the tumor. Tumor metastasis and vascular invasion were investigated by intraoperative and postoperative pathological examination respectively. The postoperative follow-up duration was 10 years.
Tissue microarray (TMA)
Fresh tissue from the central zone was paraformaldehyde-fixed, imbedded in paraffin and sliced into 4-µm sections. These sections were stained by hematoxylin and eosin (H & E) and placed on coverslips with mounting medium for analysis. The stained and paraformaldehyde-fixed tissues were then placed in a TMA. A total of 7 TMAs (6 from tumors and 1 from adjacent mucosa) were used for ISH analysis.
ISH
ISH was conducted as previously described (17). Briefly, tissues with treated with hydrogen peroxide at room temperature for 10 min. Subsequently, the sections were boiled (95 to 100 °C) in citric acid buffer for 15 min and incubated with protease at 37 °C for 8 min. The slides were then pre-hybridized in Exiqon hybridization buffer (Exiqon, Vedbæk, Denmark) at 65 °C for 2 h, hybridized with 50 nM miRNA-181a probe overnight and washed stringently with 5× SSC, 1× SSC and 0.2× SSC buffers at 65 °C for 30 min. DIG blocking reagent (Roche, Mannheim, Germany) was added for 15 min at 37 °C in maleic acid buffer with 2% sheep serum. After which, the alkaline phosphatase-conjugated anti-digoxigenin was added at 37°C for 20 min (1:500 in Roche blocking reagent). 4-nitroblue tetrazolium (NBT) and 5-brom-4-chloro-3'-Indolylphosphate (BCIP) substrate (Roche) were used for enzymatic development to form dark-blue NBT-formazan precipitates at 37 °C for 60 min. The sections were lightly counterstained with nuclear fast red (Vector Laboratories, Burlingame, CA) at 25 °C for 1 min and mounted. The probes for miRNA-181a detection was as follows: 5' Dig-AACAUUCAACGCUGUCGGUGAGU-Dig 3' (Exiqon, Vedbæk, Denmark).
Evaluation of ISH
TMAs were scanned using Pannoramic MIDI (3D HISTECH, Budapest, Hungary) and anonymized samples were then analyzed. The number of cells in each slice that had positive staining was recorded. It was based on staining intensity and classified using semi-quantified values. The process of semi-quantification was conducted as follows: H-score = Σ(PI×I) = (percentage of cells of negative intensity ×0) + (percentage of cells of weak intensity ×1) + (percentage of cells of moderate intensity ×2) + (percentage of cells of strong intensity ×3), where PI represents the percentage of positive cells to the total number of cells in the slice and I represents the intensity of the color. Five fields for each sample were selected randomly and the cutoff value for miRNA-181a levels was determined through receiver operating characteristic (ROC) curves.
Statistical analysis
Statistical analysis was conducted using SPSS 20.0 (Chicago, IL, USA). Pearson chi-square and Fisher’s tests was used to compare the statistical data. Disease-free survival (DFS) referred to the date from the day of diagnosis to tumor local recurrence or metastasis. Overall survival (OS) referred to the date from the day of diagnosis to tumor-related death. The Kaplan-Meier method and the log-rank test were used to evaluate survival probabilities. The association of multiple variables and prognosis was assessed using the multivariate Cox regression model. P values less than 0.05 was considered to have statistical significance.
The Cancer Genome Atlas (TCGA) database analysis
The expression data for miRNA-181a was obtained from the TCGA database for CRC, which contained 619 cases of colorectal tumors and 11 cases of adjacent normal tissues. The median age of these patients was 68 years, which included 328 (53.0%) males and 291 (47.0%) females. The number of patients with tumor grade 1 + grade 2 and tumor grade 3 + grade 4 were 90 (19.6%) and 369 (80.4%) respectively. Of the 619 patients, 185 (47.5%) patients had lymph node metastasis and 152 (24.9%) patients had distant metastasis, 510 (85.1%) patients had clinical stage I–III and 89 (14.9%) had clinical stage IV (Table S1). MiRNA-181a expression profiles were matched to the downloaded clinical data according to the TCGA code. Some patients had missing miRNA-181a expression data, no follow-up data, or missing clinical information. The t-test was conducted to analyze the miRNA-181a expression differences between cancer and normal tissues and the Kaplan-Meier method was performed to evaluate the association of miRNA-181a levels with OS.
Results
Clinicopathological characteristics of CRC patients
Table 1 summarizes the clinicopathological characteristics of the 329 CRC patients with miRNA-181a expression levels. The median age of these patients was 63 years (range, 17–88 years), which included 173 (52.6%) males and 156 (47.4%) females. The numbers of patients with histological grade G1 and G2–G3 tumors were 53 (16.1%) and 276 (83.9%) respectively. A total of 284 (86.3%) cases exhibited a tumor diameter of 30 mm and/or more, while 45 cases (13.7%) exhibited no more than 30 mm. Of the 329 patients, 43 (13.1%), 46 (14.0%), 22 (6.7%) and 29 (8.8%) patients had lymph node metastasis, liver, lung invasion and peritoneal dissemination respectively. A total of 238 (72.3%) patients had clinical stage I–III and 91 (27.7%) had clinical stage IV. In total, 160 patients received chemotherapy treatment, which were 48.6% of the total number of CRC patients.
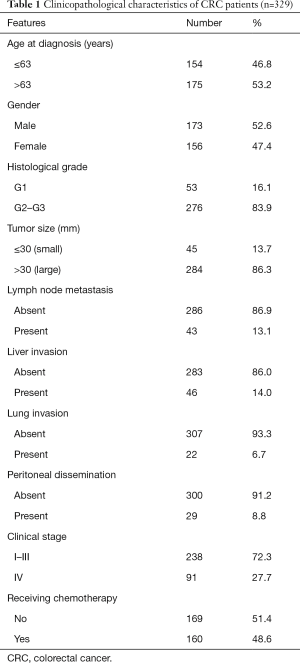
Full table
MiRNA-181a is overexpressed in CRC tissues
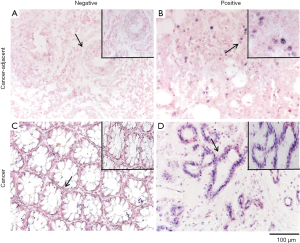
We evaluated the expression of miRNA-181a levels in 329 samples from patients with CRC and 45 tumor-adjacent control samples utilizing ISH (Figure 1). Among the 329 samples, 302 (91.8%) were positive for miRNA-181a expression in CRC tissues. In contrast, miRNA-181a was detected in only 6 (13.3%) samples out of 45 tumor-adjacent control samples. MiRNA-181a expression in CRC tissue samples was significantly higher than in corresponding tumor-adjacent samples (P<0.01).
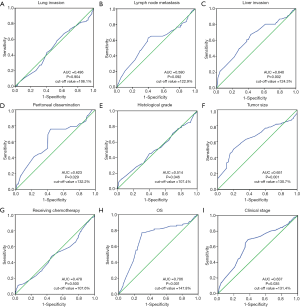
Selection of the cutoff value for miRNA-181a expression levels
ROC analysis was used to determine the appropriate cutoff score for miRNA-181a expression level. ROC curves demonstrated miRNA-181a expression levels were affected by liver invasion (P=0.002), peritoneal dissemination (P=0.029), histological grade (P=0.046), tumor size (P=0.001), OS (P<0.001) and clinical stage (P=0.035) (Figure 2). However, lung invasion, chemotherapy and lymph node metastasis were not associated (P=0.904, P=0.500, P=0.092). The largest area under the curve (AUC) of the ROC curve was corresponded to OS (Figure 2H). Based on these results, we defined the cut-off value of miRNA-181a expression level at 147.8%. Tumors with H-scores >147.8% and ≤147.8% were considered as high and low expression of miRNA-181a respectively. A total of 151 (45.9%) tumors had low expression of miRNA-181a and 178 (54.1%) tumors had high expression of miRNA-181a.
Association of miRNA-181a expression with clinicopathological characteristics of CRC
The association between miRNA-181a expression levels with clinical pathological parameters in CRC was investigated using Pearson χ2 test (Table 2). The expression levels of miRNA-181a were not correlated with the patients’ age, gender, histological grade, lymph node metastasis, lung invasion and receiving chemotherapy (Table 2, P>0.05). Tumors diameter >30 mm, liver invasion and peritoneal dissemination had significant association with miRNA-181a high expression (P<0.001, P=0.004, P=0.018, Table 2). The high expression of miRNA-181a was also significantly correlated with tumor clinical stage (P=0.003, Table 2).
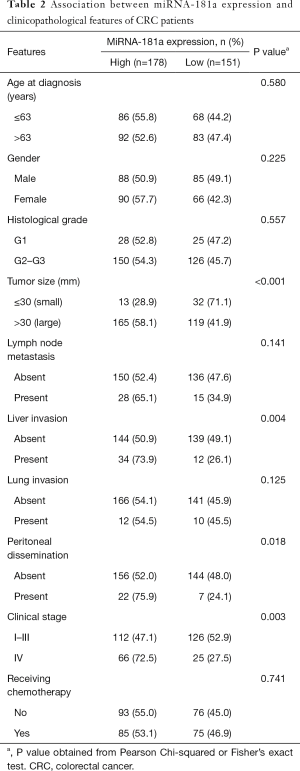
Full table
Association of miRNA-181a expression levels with survival
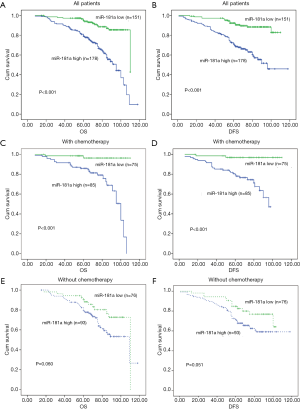
The association of miRNA-181a expression levels with OS and DFS were determined in CRC patients using Kaplan-Meier analysis. The high expression of miRNA-181a was correlated with shorter OS and DFS in all CRC patients (n=329, P<0.001, P<0.001, Figure 3A,B). Given that adjuvant chemotherapy influenced patients’ survival, we analyzed the association between miRNA-181a expression levels and its prognostic value in CRC patients receiving and not receiving chemotherapy. MiRNA-181a was highly associated with OS and DFS in patients who underwent chemotherapy than those who did not receive chemotherapy (Figure 3C,D,E,F). Moreover, the therapeutic efficacy of patients receiving adjuvant chemotherapy with high miRNA-181a expression was not beneficial compared to patients with low miRNA-181a expression levels (P<0.001, P<0.001, log-rank test, Figure 3C,D). These results suggested that miRNA-181a expression levels could influence the prognosis of CRC patients by affecting the chemotherapy effect. In addition, we investigated the correlation of miRNA-181a expression levels with OS and DFS in subgroups of CRC patients who were classified based on histological grade, liver invasion and peritoneal dissemination. High expression of miRNA-181a was significantly correlated with shorter OS, DFS in CRC patients with histological grade G2–G3 but not in G1 (P<0.001, P=0.215, log-rank test, Figure 4A,B). In liver metastatic-negative and peritoneal dissemination-negative patients, higher miRNA-181a expression levels were significantly associated with shorter OS (P<0.001, P<0.001, Figure 4C,D). However, the expression of miRNA-181a was not correlated with shorter OS in liver invasion-positive and peritoneal dissemination-positive patients (P=0.583, P=0.809, Figure 4E,F). This suggests that miRNA-181a may be an important factor affecting OS and DFS of CRC patients with histological grade G2–G3, without liver invasion and peritoneal dissemination.

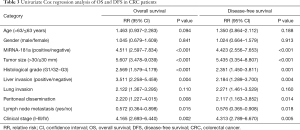
Next, we used the univariate and multivariate Cox survival risk regression model to analyze the correlation between common clinical pathology parameters with miRNA-181a expression levels for the prognosis of CRC patients. We found that the expression levels of miRNA-181a, together with lymph node metastasis, liver invasion, peritoneal dissemination, histological grade, tumor size, and clinical stage were significantly correlated with OS and DFS in CRC patients (P<0.001, P=0.015, P=0.004, P=0.008, P<0.001, P<0.001 and P=0.002 respectively for OS; P<0.001, P=0.018, P=0.004, P=0.014, P=0.001, P<0.001 and P=0.005 respectively for DFS; Table 3). We selected P value <0.05 variables to conduct the multivariate Cox regression analysis. Multivariate analysis indicated that miRNA-181a expression levels, tumor size, histological grade and clinical stage were independent prognostic factors for OS and DFS in CRC patients (P<0.001, P=0.035, P=0.003 and P<0.001 respectively for OS; P<0.001, P=0.046, P=0.005 and P<0.001 respectively for DFS; Table 4). This further demonstrated that miRNA-181a expression level was an independent risk factor for OS and DFS [risk ratio (RR) =2.872, 95% CI: 1.589–5.194, P<0.001 for OS; RR =3.024, 95% CI: 1.661–5.507, P<0.001 for DFS] (Table 4).
Full table

Full table
MiRNA-181a expression levels are associated with CRC progression and prognosis based on the TCGA database
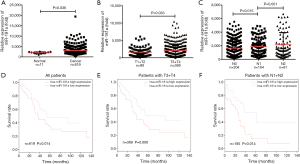
In order to further confirm that miRNA-181a expression levels had a high correlation with CRC, we analyzed miRNA-181a expression in CRC tissues of 619 cases and 11 adjacent normal tissues using the TCGA database. The results demonstrated an obvious increase in miRNA-181a expression levels in CRC tissues (n=619) compared with normal tissues (n=11, P=0.036, Figure 5A). We then analyzed the miRNA-181a expression in subgroups of CRC tissues classified based on T stage and lymph node metastases. The expression of miRNA-181a in T3 + T4 (n=369) stage was remarkably higher than in T1 + T2 stage (n=90, P=0.033, Figure 5B) and miRNA-181a expression was significantly lower in N0 (n=204) stage than those in N1 (n=104) and N2 (n=81) stage (P=0.015, P=0.001, Figure 5C). The results demonstrated that miRNA-181a expression was increased significantly in advanced T stage and lymph node metastases, which was consistent with its effect on colorectal tumor progression. We next analyzed the correlation between miRNA-181a expression and the prognosis of 619 cases of CRC patients based on the TCGA clinical database. The results showed that high expression of miRNA-181a was significantly correlated with poor OS in CRC patients (P=0.014, Figure 5D). Furthermore, high expression of miRNA-181a was associated with shorter OS in CRC patients with T3 + T4 stage tumors (P=0.006, Figure 5E). However, miRNA-181a expression levels had no correlation in patients with T1 + T2 stage tumors (P=0.821, data not shown). The high expression of miRNA-181a was also associated with a poor survival prognosis in CRC patients with lymph node metastases (P=0.014, Figure 5F) and had no association in CRC patients without lymph node metastases (P=0.302, data not shown). These results demonstrate that miRNA-181a may play an important role in the development and progression of CRC and may be an indicator for CRC diagnosis and prognosis.
Discussion
Numerous studies have confirmed the significant role that miRNA-181a plays in regulating tumorigenesis, proliferation and distant metastasis. However, the association of miRNA-181a with regards to CRC has yet to be determined. In the present study, we first conducted high-throughput TMA using ISH to determine the prognostic value of miRNA-181a expression levels with CRC. We found that CRC had higher expression of miRNA-181a than tumor-adjacent control samples. CRC patients with higher expression levels of miRNA-181a had a higher clinical stage, increased liver metastasis, peritoneal dissemination and larger tumor size. Higher expression of miRNA-181a was also correlated with shorter OS and DFS in CRC patients and was an independent prognostic factor for CRC progression.
MicroRNAs are stably expressed in tumor tissues and are attributed to them being frequently used for detection (18,19). The majority of previous studies on miRNA detection have been carried out using microarrays with RNA extracted from tumor tissues and tumor-adjacent stromal tissues, which may include a mixture of tumor cells and tumor related stromal cells, thus making the analysis difficult to interpret. However, ISH can detect positive signals precisely at the cellular level. For example, some miRNAs are expressed abundantly in tumor related stromal cells but have no expression in tumor cells (20). Analysis using microarrays with RNA extracted from tumors will not be able to precisely delineate these differences.
In addition, several miRNAs act as tumor suppressors or oncogenes (“onco-miRNA”), which would not have been detected if molecular and cellular composition is not utilized. Taking this into context, a pivotal but controversial role has been attributed to miRNA-181a as an onco-miRNA in different human cancers, including cervical, prostate, gastric, hepatocellular carcinoma and CRCs (13,21-24). Conversely, some studies have suggested that miRNA-181a plays an opposite role as a tumor suppressor in other types of cancers (11,25,26). For example, miRNA-181a has been identified as frequently downregulated, contributes to apoptosis, and inhibition of growth and invasion in gliomas (27). However, in another study, low miRNA-181a expression was significantly associated with poor cancer-specific survival in patients with CRC (28).
Several studies have demonstrated that miRNA-181a is overexpressed in CRC (23,29,30). As far as we know, only one publication has reported the prognostic value of miRNA-181a expression levels in CRC using ISH (31). Consistent with our results, high miRNA-181a expression was an independent and significant prognostic factor for CRC. Our study was also consistent with the report from Nishimura et al. (32). Nishimura et al. showed that high miRNA-181a expression levels detected by RT-qPCR had a significantly poorer prognosis in CRC patients. However, in the report by Nishimura et al., the expression of miRNA-181a had no obvious differences between tumor-adjacent tissue and CRC tissue and had no association with any clinical pathological parameters. We hypothesized that these results were probably due to the different origin of tumor-adjacent tissues and the number and detection methods used between our two studies.
The underlying mechanism how miRNA-181a influences poor survival remains to be elucidated. MiRNA-181a has been shown to inhibit the expression of several tumor suppressor genes, such as PTEN (23), MTMR3 (22), GPD1L (33) and WIF-1 (31,34). A recent study found that miRNA-181a expression was transcriptionally regulated by signal transducers and activators of transcription 1 (STAT1) in CRC. STAT1/miRNA-181a/PTEN pathway in CRC adds new insights regarding the carcinogenesis and development of CRC (23). In addition, the Wnt/β-catenin signaling pathway is deregulated in CRC and miRNA-181a could be directly up-regulated through the activation of the Wnt/β-catenin pathway (30). These results/findings could explain the reason for the high expression of miRNA-181a in CRC. Regulating the expression of miRNA-181a through modulating transcription factors may be a therapeutic option.
Our research also analyzed the potential association between miRNA-181a expression and a variety of clinical pathological parameters in CRC patients, as well as its prognostic value for survival. Our results demonstrated that high expression of miRNA-181a was associated with tumor size, liver metastasis, peritoneal dissemination, clinical stage and shorter prognostic survival in CRC patients. We also showed that the expression of miRNA-181a was significantly associated with poor prognosis, however, it did not have significant association with prognosis if patients had liver metastasis and peritoneal dissemination. This suggested that although the expression of miRNA-181a had an obvious effect on proliferation of tumor cells and poor prognosis, it was not the dominant factor when liver metastasis or peritoneal dissemination was present.
Furthermore, miRNA-181a had a significant impact on regulating tumor drug-resistance in various cancers. Higher expression of miRNA-181a increased tumor resistance to docetaxel and cabazitaxel in prostate cancer cells and inhibition of miRNA-181a expression could restore treatment efficacy (13). TMZ-based chemotherapy is the standard treatment for glioma and miRNA-181a reduced the sensitivity to TMZ in glioma cancers through the inhibition of PTEN (15). In this study, we found that the prognostic value of miRNA-181a-high patients receiving adjuvant chemotherapy was poor, which suggested that high expression of miRNA-181a may influence the progression and prognosis of CRC. Our previous published work demonstrated that miRNA-181a played a significant role in regulating macrophage polarization through directly targeting KLF6 and C/EBPα to induce macrophage polarization to the M2 phenotype, which leads to tumor development and drug resistance (35). These results further demonstrated that the expression of miRNA-181a could be a potential therapeutic marker for CRC patients. Our results demonstrated for the first time that miRNA-181a expression was significantly correlated with the survival and efficacy of chemotherapy treatment of CRC patients.
Conclusions
In summary, our results demonstrated that miRNA-181a expression was significantly associated with tumor size, distant metastasis, clinical stage and prognostic survival of CRC patients, indicating that miRNA-181a expression has a significant effect on the proliferation and progression of tumor cells of the disease. We also demonstrated that the expression of miRNA-181a was significantly correlated with OS and DFS and may be an important indicator for prognosis of CRC patients.

Full table
Acknowledgments
Funding: This work was supported by grants from National Natural Science Foundation of China (No. 30973559, 81572898), Program for Liaoning Innovative Research Team in University (No. LT2014016), Program for Liaoning Excellent Talents in University (No. LJQ2015118), Key Laboratory Foundation from Shenyang S&T Projects (F16-094-1-00).
Footnote
Conflicts of Interest: All authors have completed the ICMJE uniform disclosure form (available at http://dx.doi.org/10.21037/tcr.2018.01.18). The authors have no conflicts of interest to declare.
Ethical Statement: The authors are accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved. The study was conducted in accordance with the Declaration of Helsinki (as revised in 2013). This study was approved by the Medical Ethics Committee of China Medical University at the First Affiliated Hospital of China Medical University (ID: AF-SOP-07-1.0-01). All patients agreed to the study protocols and signed informed consent forms to participate in the study.
Open Access Statement: This is an Open Access article distributed in accordance with the Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License (CC BY-NC-ND 4.0), which permits the non-commercial replication and distribution of the article with the strict proviso that no changes or edits are made and the original work is properly cited (including links to both the formal publication through the relevant DOI and the license). See: https://creativecommons.org/licenses/by-nc-nd/4.0/.
References
- Jemal A, Bray F, Center MM, et al. Global cancer statistics. CA Cancer J Clin 2011;61:69-90. [Crossref] [PubMed]
- Ferlay J, Shin HR, Bray F, et al. Estimates of worldwide burden of cancer in 2008: GLOBOCAN 2008. Int J Cancer 2010;127:2893-917. [Crossref] [PubMed]
- Heinemann V, von Weikersthal LF, Decker T, et al. FOLFIRI plus cetuximab versus FOLFIRI plus bevacizumab as first-line treatment for patients with metastatic colorectal cancer (FIRE-3): a randomised, open-label, phase 3 trial. Lancet Oncol 2014;15:1065-75. [Crossref] [PubMed]
- Loupakis F, Cremolini C, Masi G, et al. Initial therapy with FOLFOXIRI and bevacizumab for metastatic colorectal cancer. N Engl J Med 2014;371:1609-18. [Crossref] [PubMed]
- Mehlen P, Puisieux A. Metastasis: a question of life or death. Nat Rev Cancer 2006;6:449-58. [Crossref] [PubMed]
- Fidler IJ. The pathogenesis of cancer metastasis: the 'seed and soil' hypothesis revisited. Nat Rev Cancer 2003;3:453-8. [Crossref] [PubMed]
- Zhang B, Pan X, Cobb GP, et al. microRNAs as oncogenes and tumor suppressors. Dev Biol 2007;302:1-12. [Crossref] [PubMed]
- Filipowicz W, Bhattacharyya SN, Sonenberg N. Mechanisms of post-transcriptional regulation by microRNAs: are the answers in sight? Nat Rev Genet 2008;9:102-14. [Crossref] [PubMed]
- Esquela-Kerscher A, Slack FJ. Oncomirs - microRNAs with a role in cancer. Nat Rev Cancer 2006;6:259-69. [Crossref] [PubMed]
- Michael MZ. Reduced accumulation of specific microRNAs in colorectal neoplasia. Mol Cancer Res 2003;1:882-91. [PubMed]
- Schwind S, Maharry K, Radmacher MD, et al. Prognostic significance of expression of a single microRNA, miR-181a, in cytogenetically normal acute myeloid leukemia: a Cancer and Leukemia Group B study. J Clin Oncol 2010;28:5257-64. [Crossref] [PubMed]
- Gao W, Yu Y, Cao H, et al. Deregulated expression of miR-21, miR-143 and miR-181a in non small cell lung cancer is related to clinicopathologic characteristics or patient prognosis. Biomed Pharmacother 2010;64:399-408. [Crossref] [PubMed]
- Armstrong CM, Liu C, Lou W, et al. MicroRNA-181a promotes docetaxel resistance in prostate cancer cells. Prostate 2017;77:1020-8. [Crossref] [PubMed]
- Ke G, Liang L, Yang JM, et al. MiR-181a confers resistance of cervical cancer to radiation therapy through targeting the pro-apoptotic PRKCD gene. Oncogene 2013;32:3019-27. [Crossref] [PubMed]
- Liao Y, Shen L, Zhao H, et al. LncRNA CASC2 Interacts With miR-181a to Modulate Glioma Growth and Resistance to TMZ Through PTEN Pathway. J Cell Biochem 2017;118:1889-99. [Crossref] [PubMed]
- Dindo D, Demartines N, Clavien PA. Classification of surgical complications: a new proposal with evaluation in a cohort of 6336 patients and results of a survey. Ann Surg 2004;240:205-13. [Crossref] [PubMed]
- Wang F, Flanagan J, Su N, et al. RNAscope: a novel in situ RNA analysis platform for formalin-fixed, paraffin-embedded tissues. J Mol Diagn 2012;14:22-9. [Crossref] [PubMed]
- Xi Y, Nakajima G, Gavin E, et al. Systematic analysis of microRNA expression of RNA extracted from fresh frozen and formalin-fixed paraffin-embedded samples. Rna 2007;13:1668-74. [Crossref] [PubMed]
- Li J, Smyth P, Flavin R, et al. Comparison of miRNA expression patterns using total RNA extracted from matched samples of formalin-fixed paraffin-embedded (FFPE) cells and snap frozen cells. BMC Biotechnol 2007;7:36. [Crossref] [PubMed]
- Hu Y, Correa AM, Hoque A, et al. Prognostic significance of differentially expressed miRNAs in esophageal cancer. Int J Cancer 2011;128:132-43. [Crossref] [PubMed]
- Yang M, Zhai X, Ge T, et al. MiR-181a-5p Promotes Proliferation and Invasion, and Inhibits Apoptosis of Cervical Cancer Cells via Regulating Inositol Polyphosphate-5-Phosphatase A (INPP5A). Oncol Res 2017; [Epub ahead of print]. [Crossref] [PubMed]
- Lin Y, Zhao J, Wang H, et al. miR-181a modulates proliferation, migration and autophagy in AGS gastric cancer cells and downregulates MTMR3. Mol Med Rep 2017;15:2451-6. [Crossref] [PubMed]
- Zhang X, Li X, Tan F, et al. STAT1 Inhibits MiR-181a Expression to Suppress Colorectal Cancer Cell Proliferation Through PTEN/Akt. J Cell Biochem 2017;118:3435-43. [Crossref] [PubMed]
- Nishida N, Arizumi T, Hagiwara S, et al. MicroRNAs for the Prediction of Early Response to Sorafenib Treatment in Human Hepatocellular Carcinoma. Liver Cancer 2017;6:113-25. [Crossref] [PubMed]
- Cao Y, Zhao D, Li P, et al. MicroRNA-181a-5p Impedes IL-17-Induced Nonsmall Cell Lung Cancer Proliferation and Migration through Targeting VCAM-1. Cell Physiol Biochem 2017;42:346-56. [Crossref] [PubMed]
- Lin F, Li Y, Yan S, et al. MicroRNA-181a inhibits tumor proliferation, invasiveness, and metastasis and is downregulated in gastric cancer. Oncol Res 2015;22:75-84. [Crossref] [PubMed]
- Shi L, Cheng Z, Zhang J, et al. hsa-mir-181a and hsa-mir-181b function as tumor suppressors in human glioma cells. Brain Res 2008;1236:185-93. [Crossref] [PubMed]
- Pichler M, Winter E, Ress AL, et al. miR-181a is associated with poor clinical outcome in patients with colorectal cancer treated with EGFR inhibitor. J Clin Pathol 2014;67:198-203. [Crossref] [PubMed]
- Hai Ping P, Feng Bo T, Li L, et al. IL-1beta/NF-kb signaling promotes colorectal cancer cell growth through miR-181a/PTEN axis. Arch Biochem Biophys 2016;604:20-6. [Crossref] [PubMed]
- Han P, Li JW, Zhang BM, et al. The lncRNA CRNDE promotes colorectal cancer cell proliferation and chemoresistance via miR-181a-5p-mediated regulation of Wnt/beta-catenin signaling. Mol Cancer 2017;16:9. [Crossref] [PubMed]
- Ji D, Chen Z, Li M, et al. MicroRNA-181a promotes tumor growth and liver metastasis in colorectal cancer by targeting the tumor suppressor WIF-1. Mol Cancer 2014;13:86. [Crossref] [PubMed]
- Nishimura J, Handa R, Yamamoto H, et al. microRNA-181a is associated with poor prognosis of colorectal cancer. Oncol Rep 2012;28:2221-6. [Crossref] [PubMed]
- Zhai X, Meng R, Li H, et al. miR-181a Modulates Chondrocyte Apoptosis by Targeting Glycerol-3-Phosphate Dehydrogenase 1-Like Protein (GPD1L) in Osteoarthritis. Med Sci Monit 2017;23:1224-31. [Crossref] [PubMed]
- Lyu X, Li J, Yun X, et al. miR-181a-5p, an inducer of Wnt-signaling, facilitates cell proliferation in acute lymphoblastic leukemia. Oncol Rep 2017;37:1469-76. [Crossref] [PubMed]
- Bi J, Zeng X, Zhao L, et al. miR-181a Induces Macrophage Polarized to M2 Phenotype and Promotes M2 Macrophage-mediated Tumor Cell Metastasis by Targeting KLF6 and C/EBPalpha. Mol Ther Nucleic Acids 2016;5:e368 [Crossref] [PubMed]


