
MicorRNA-148b inhibits cell proliferation and facilitates cell apoptosis by regulating DNA Methyltransferase 1 in endometrial cancer
Introduction
Endometrial cancer (EC) is a gynaecologic malignancy and the sixth most common tumour in females (1). According to Global Cancer Statistics, more than 382,000 new cases of EC are diagnosed each year worldwide (2). The current therapeutic approaches for EC include surgical resection, brachytherapy, radiotherapy, and chemotherapy (3). However, despite substantial improvements in the treatment of EC in recent years, the survival rate of patients with EC is still poor (4). Therefore, exploring the molecular mechanisms that mediate the progression of EC is critical to improve the effectiveness of EC treatment.
MicroRNAs (miRNAs) are a large family of short, single-stranded, noncoding RNAs (19–25 nucleotides in length) that modulate target gene expression by directly pairing with the 3'-untranslated regions (3'-UTRs) of target genes, leading to mRNA cleavage or translational inhibition (5). Accumulating evidence has confirmed that miRNAs play important roles in various biological processes in tumours, such as cell growth, apoptosis, cell migration, the cell cycle, and carcinogenesis (6-8). Thus, analysis of miRNAs in cancer research is essential (9). Several studies have shown that the expression of several miRNAs is dysregulated in EC, which implicates these miRNAs in the development and progression of EC by influencing a wide range of biological processes (10). Thus, investigating the roles of these dysregulated miRNAs may be useful for identifying potential therapeutic targets and developing effective treatments for EC.
DNA methyltransferase (DNMT) is a kind of most abundant DNA methyltransferase and play significant roles in epigenetic processes, including maintaining genome integrity and gene expression (11). Previous studies have demonstrated that DNMTs mediate transcriptional silencing in various malignant tumours, such as lung (12) and ovarian cancers (13). Importantly, miR-148b has been shown to regulate DMNT1 in pancreatic cancer (14). miR-148b belongs to miR-148/152 family, and has a seed sequence of 6–7 nucleotides and mature structure of 21–22 nucleotides in length (15). A number of studies have demonstrated that miR-148b inhibits the progression of various cancers, such as colorectal (16), gastric (17), non-small cell lung (18), and cervical cancers (19). Nevertheless, the expression of miR-148b and its potential roles in EC are not well understood. In the present study, we evaluated the effects of miR-148b on EC cell proliferation and apoptosis. We also verified the targeting relationship between miR-148b and DNMT1. The findings of this research improve our understanding of the biological activities of miR-148b in EC.
Methods
Patients and tissue samples
From January 2016 to December 2018, total of 22 EC tissues and corresponding cancer-adjacent tissues were collected from EC patients at our hospital. No patients had received adjuvant treatment, such as chemotherapy or radiotherapy, before surgery. The cancerous and neighbouring (<3 cm away from cancerous tissues) tissues were collected during hysterectomy. The collected tissues were washed twice with ice-cold phosphate buffer saline (PBS) and then stored at −70 °C until use. The study was approved by the Ethics Committee of our hospital, and all patients provided informed consent.
Cell cultures
A normal human endometrial cell line (EMC) and two human EC cell lines (HEC-1A and HEC-1B) were obtained from American Type Culture Collection (ATCC; Manassas, VA, USA) and cultured in Dulbecco’s Modified Eagle Medium (DMEM; Gibco, USA) containing 10% fatal bovine serum (FBS) and 1% streptomycin/penicillin at 37 °C with 5% CO2.
Cell transfection and grouping
HEC-1A and HEC-1B cells (1×105 cells/well) were seeded in 6-well plates and then separately transfected with either miR-148b mimics or miR-148b inhibitor both obtained from GenePharma (Shanghai, China) using Lipofectamine® 3000 Reagent (Invitrogen, Carlsbad, CA, USA). DNMT1 siRNA (si-DMNT1) and a corresponding negative control siRNA (si-NC) were obtained from RIBO Bio (Guangzhou, China). The cells were randomly divided into the miR-148b mimics group (treated with miR-148b mimics), NC-mimics group (treated with the negative control for the miR-148b mimics), NC-inhibitor group (treated with the negative control for miR-148b inhibitor), miR-148b inhibitor group (treated with miR-148b inhibitor), si-DNMT1 group (treated with DNMT1 siRNA), si-NC group (treated with negative control siRNA), NC-inhibitor + si-NC group (treated with NC-inhibitor and si-NC), NC-inhibitor + si-DNMT1-1 group (treated with NC-inhibitor and si-DNMT1-1), miR-148b inhibitor + si-NC group (treated with miR-148b inhibitor and si-NC), miR-148b inhibitor + si-DNMT1 group (treated with miR-148b inhibitor and si-DNMT1), and BLANK group (no treatment). All cells were cultured in a 37 °C incubator for 48 h.
Quantitative real-time PCR (qRT-PCR)
Total RNA was extracted from EC tissues, normal tissues, and cell lines with TRIzol reagent (Invitrogen), and the RNA concentration was determined with a NanoDrop ND-1000 spectrophotometer (NanoDrop, USA). Then, an aliquot of the extracted total RNA (500 ng) was reverse-transcribed into cDNA using the PrimeScript RT kit (TaKaRa, Dalian, China) and analysed by qRT-PCR with the SYBR Green PCR kit (TaKaRa). The primers used are listed in Table 1. U6 and β-actin were used as internal controls for miR-148b and DNMT1, respectively.
Table 1
| Name of primer | Sequences (5'-3') |
|---|---|
| miR-148b-F | GCGCTCAGTGCATCACAGA |
| miR-148b-R | GTGCAGGGTCCGAGGT |
| DNMT1-F | TGTTTCACGCCCACTGGTTCTG |
| DNMT1-R | GCTGCCTTTGATGTAGTCGGAGTATTT |
| U6-F | CTTCGGCAGCACATATACT |
| U6-R | AAAATATGGAACGCTTCACG |
| si-DNMT1-1-F | CCCACAACAUGAAAGAAACTT |
| si-DNMT1-1-R | AUUUCUUUCAUGUUGUGGGTT |
| si-DNMT1-2-F | CAUGGUAGUCACUAACAUATT |
| si-DNMT1-2-R | UAUGUUAGUGACUACCAUGTT |
| si-NC-F | UUCUCCGAACGUGUCACGUTT |
| si-NC-R | ACGUGACACGUUCGGAGAATT |
| β-actin-F | TGAGACCTTCAACACCCCAG |
| β-actin -R | GCCATCTCTTGCTCGAAGTC |
Western blot
Total proteins were extracted with radio immunoprecipitation assay (RIPA) lysis buffer (Sigma, USA). Then, the extracted proteins (50 µg) were separated by 10% sodium dodecyl sulphate-poly acrylamide gel electrophoresis (SDS-PAGE) and transferred to a polyvinylidene difluoride membrane. The membrane was blocked with 5% non-fat milk for 2 h, and then incubated with the following primary antibodies: anti-DNMT1, anti-β-actin, anti-cyclin-dependent kinase 4 (anti-CDK4), anti-p16, anti-cyclin D1, anti-caspase-3, anti-B-cell lymphoma 2 (anti-Bcl-2), and anti-BCL2-associated X (anti-Bax) (1:1,000; Cell Signaling Technology, MA, USA) overnight at 4 °C. After three washes, the membranes were incubated with peroxidase-labelled anti-rabbit Immunoglobulin G (1:2,000, I5006MSDS; Sigma, USA) as a secondary antibody for 2 h. Finally, the protein bands were visualized with the ECL system (Thermo, USA) and analysed with Image Lab™ Software (Bio-Rad, USA).
Dual luciferase reporter assay
The targeting relationship between miR-148b and DNMT1 was predicted by TargetScan, and then the exact target binding sites were determined. We amplified a fragment of the 3'-UTR of DNMT1, including the miR-148b binding site, and then cloned this fragment into the pmirGLO vector (Promega, USA) to generate wild-type pmirGLO-WT-DNMT1-3'-UTR (DNMT1-Wt) and mutant pmirGLO-MUT-DNMT1-3'-UTR (DNMT1-Mut). Then, HEC-1A and HEC-1B cells were co-transfected with one of these reporter vectors along with either miR-148b mimics or the corresponding NC using Lipofectamine 3000. After 48 hours, luciferase activity was detected with the dual luciferase kit (Promega, USA).
RNA pull-down assay
Biotinylated DNMT1 NC, DNMT1-Wt, and DNMT1-Mut (GenePharma, Shanghai, China) were transfected into HEC-1A and HEC-1B cells to generate the following three respective groups: Bio-NC, Bio-DNMT1-Wt, and Bio-DNMT1-Mut. After 48 h, the cells were lysed and then incubated with streptavidin agarose beads (ThermoFisher, USA) at 37 °C for 1 h. Finally, mRNA was detected by qRT-PCR.
Cell counting kit 8 (CCK-8)
At 24, 48, 72, and 96 h, 10 µL of CCK-8 was added to transfected HEC-1A and HEC-1B cells (2×104 cells/well) and incubated at room temperature for 1 h. Then, the absorbance was measured with a microplate reader (Bio-Rad, USA) at 450 nm.
Colony formation assay
Transfected HEC-1A and HEC-1B cells were seeded into 6-well plates at a density of 300 cells per well and cultivated for 14 d. After fixing with 4% cold formaldehyde for 30 min, the colonies were stained with 0.1% crystal violet (Beyotime, China) for 20 min. Then, the stained colonies were photographed and counted under an optical microscope (Olympus, Tokyo, Japan).
Flow cytometry
To assess the cell cycle status of the cells, transfected HEC-1A and HEC-1B cells were fixed in 70% pre-cooled ethanol and stained with Muse™ Cell Cycle Reagent (Millipore, USA) for 30 min in the dark. The number of cells in each phase of the cell cycle was detected with a MUSE Cell Analyzer (Millipore). To detect cell apoptosis, different groups of cells were sequentially stained with annexin V-Enhanced Green Fluorescent Protein (EGFP) and Propidium Iodide (PI) (Thermo Fisher Scientific, USA). After incubation for 15 min in the dark, the apoptotic rate was detected with a MUSE Cell Analyzer (Millipore).
Statistical analysis
Statistical analyses were performed using GraphPad Prism 7.0 software (San Diego, CA, USA) and SPSS 22.0 statistical software (Chicago, IL, USA). The results are presented as the mean ± standard deviation (SD). All experiments were performed at least three times. The results of two groups were compared using Student’s t-test, and the differences between several groups were analysed by One-way Analysis of Variance (ANOVA) followed by Tukey’s multiple comparisons test. A P value less than 0.05 was considered to be statistically significant.
Results
The expression of miR-148b is down-regulated in EC
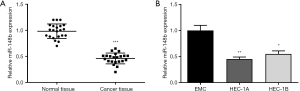
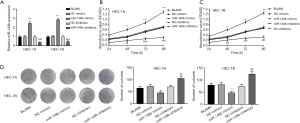
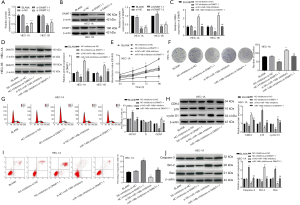
qRT-PCR was performed to assess the expression of miR-148b in EC. As shown in Figure 1A, miR-148b expression was significantly lower in EC tissues than in normal tissues (P<0.001). Similarly, miR-148b expression was lower in the EC cell lines HEC-1A and HEC-1B cells than in normal control EMC cells (P<0.01, P<0.05, respectively; Figure 1B). These results suggested that miR-148b is expressed at low levels in EC.

miR-148b inhibits the proliferation of EC cells
As shown in Figure 2A, miR-148b expression in the miR-148b mimics group was significantly higher than that in the NC-mimics and BLANK groups (P<0.001). In contrast, miR-148b expression in the miR-148b inhibitor group was significantly lower than that in the BLANK and NC-inhibitor groups (P<0.001), suggesting that the transfection was successful. Cell viability in the miR-148b mimics group was markedly lower than that in the BLANK and NC-mimics groups at 48 h (P<0.01), 72 h, and 96 h (P<0.001). When compared with the BLANK and NC-inhibitor groups, transfection of the miR-148b inhibitor significantly increased cell viability in a time-dependent manner (P<0.01; Figure 2B,C). Furthermore, transfection of miR-148b mimics markedly decreased the number of cell clones (P<0.01), while transfection of miR-148b inhibitor markedly increased the number of cell clones (P<0.01; Figure 2D). These data indicate that miR-148b inhibited the proliferation of EC cells.
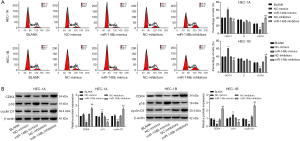
Effect of miR-148b on the cell cycle in EC cells
Transfection of miR-148b mimics dramatically increased the percentage of cells in G0/G1 phase (P<0.01) and reduced the percentages of cells in S phase (P<0.01) and G2/M phase (P<0.01). In contrast, transfection of miR-148b inhibitor dramatically decreased the percentage of cells in G0/G1 phase (P<0.01) and markedly increased the percentage of cells in G2/M phase (P<0.01) (Figure 3A). Furthermore, we detected the expression of CDK4, p16, and cyclin D1 proteins by western blotting. The expression levels of cyclin D1 and CDK4 in the miR-148b mimics group were significantly lower than those in the NC-mimics and BLANK groups, whereas p16 expression was markedly higher (P<0.01). Conversely, transcription of miR-148b inhibitor dramatically decreased CDK4 and cyclin D1 expression levels and increased p16 expression levels (P<0.01; Figure 3B). These results showed that miR-148b arrested the cell cycle of EC cells at G1/S-phase.
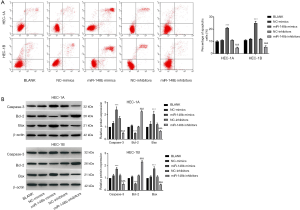
miR-148b promotes apoptosis in EC cells
As shown in Figure 4A, the apoptotic rate in the miR-148b mimics group was significantly higher than the rates in the BLANK and NC-mimics groups (P<0.001). However, transfection of miR-148b inhibitor significantly decreased apoptosis (P<0.001). The western blotting results showed that transfection of miR-148b mimics markedly increased the expression levels of Bax and caspase-3 but decreased the expression of Bcl-2 (P<0.001). In contrast, transfection of miR-148b inhibitor markedly decreased the expression of caspase-3 and Bax and increased Bcl-2 expression (P<0.001; Figure 4B). These results suggested that miR-148b promotes apoptosis in EC cells.
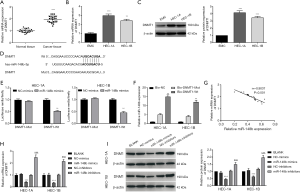
miR-148b negatively regulates the expression of DNMT1
The qRT-PCR results showed that DNMT1 mRNA expression was dramatically increased in EC tissues (P<0.001; Figure 5A). Similarly, DNMT1 mRNA and protein levels were higher in HEC-1A (P<0.001) and HEC-1B (P<0.01) cells than in EMC cells (Figure 5B,C). TargetScan predicted DNMT1 as a potential target gene of miR-148b, and a binding site for miR-148b was detected in the 3'-UTR region of DNMT1 mRNA (Figure 5D). The results of a dual luciferase reporter assay showed that transfection of miR-148b mimics dramatically reduced luciferase activity in DNMT1-Wt-transfected HEC-1A and HEC-1B cells (P<0.01; Figure 5E), suggesting that DNMT1 is a miR-148b target in EC cells. In addition, the results of an RNA pull-down assay showed that miR-148b was significantly pulled down by Bio-DNMT1-Wt (P<0.01) but not by Bio-DNMT1-Mut (P<0.01; Figure 5F). We also found that miR-148b expression was negatively correlated with DNMT1 in EC tissues (r=−0.9037, P<0.001; Figure 5G). In addition, DNMT1 mRNA and protein expression in the miR-148b mimics group was markedly lower than that in the BLANK and NC-mimics groups (P<0.001). DNMT1 expression was higher in the miR-148b inhibitor group than in the NC-inhibitor and BLANK groups (P<0.001; Figure 5H,I). These results revealed that miR-148b negatively regulates DNMT1 expression in EC cells.
DNMT1 reverses the effects of miR-148b on proliferation and apoptosis in EC cells
The results in Figure 6A,B showed that DNMT1 was dramatically down-regulated following transfection with si-DNMT1 (si-DNMT1-1 or si-DNMT1-2) when compared to the levels in the BLANK and si-NC groups (P<0.001). To determine whether miR-148b exerted its effects in EC by regulating DNMT1, HEC-1A and HEC-1B cells were co-transfected with si-DNMT1-1 and miR-148b inhibitor. The results showed that when compared with the DMNT1 expression in the BLANK and NC-inhibitor + si-NC groups, expression was significantly decreased in the NC-inhibitor + si-DNMT1-1 group (P<0.01), but increased in the si-NC + miR-148b inhibitor group (P<0.01). NEAT1 expression was lower in the miR-148b inhibitor + si-DNMT1-1 group than in the si-NC + miR-148b inhibitor group (P<0.01; Figure 6C,D). Evaluation of cell proliferation using the CCK-8 and colony formation assays (Figure 6E,F) showed that the proliferation of HEC-1A cells in the miR-148b inhibitor + si-DNMT1-1 group was lower than that in si-NC + miR-148b inhibitor group (P<0.01). In addition, we confirmed that the percentage of HEC-1A cells in G0/G1 phase was dramatically increased in the miR-148b inhibitor + si-DNMT1-1 group (P<0.01), and the percentage of cells in G2/M phase was markedly decreased when compared with that in the si-NC + miR-148b inhibitor group (P<0.01; Figure 6G). Western blotting showed that CDK4 and cyclin D1 expression levels in were markedly lower in the miR-148b inhibitor + si-DNMT1-1 group than in the si-NC + miR-148b inhibitor group (P<0.01), whereas p16 expression was significantly higher (P<0.01; Figure 6H). The apoptotic rate of HEC-1A cells transfected with miR-148b inhibitor and si-DNMT1-1 was markedly higher than that in cells transfected with miR-148b inhibitor and si-NC (P<0.01; Figure 6I). Furthermore, caspase-3 and Bax expression levels were higher in HEC-1A cells transfected with miR-148b inhibitor and si-DNMT1-1 than in cells transfected with miR-148b inhibitor and si-NC, while Bcl-2 expression was lower (P<0.01; Figure 6J). These results showed that si-DNMT1 abrogated the effect of miR-148b inhibitor on cell cycle progression in HEC-1A cells. Thus, miR-148b might suppress EC by targeting DNMT1.
Discussion
EC is mainly treated with surgery, chemotherapy, and radiotherapy; however, the long-term survival rate of EC patients is low (3). Thus, research into the novel molecular mechanisms involved in the occurrence and progression of EC is desperately needed to identify new therapeutic targets and improve the treatment of this disease. The results of this study demonstrated that miR-148b was down-regulated in EC tissues and cells and that miR-148b inhibits cell proliferation and the cell cycle and facilitates apoptosis in EC likely by modulating DNMT1.
Several studies have suggested that abnormal miRNA expression is involved in different cancers, including EC (20), which is the most commonly reported cancer of the female genital tract (21). Dysregulation of miRNAs may play an important role in the development and progression of EC by regulating a variety of processes, including cell proliferation, apoptosis, the cell cycle, metastasis, and differentiation (20). For example, when miR-183 expression is increased in EC, it inhibits apoptosis but induces epithelial-mesenchymal transition (EMT), cell proliferation, migration, and invasion by targeting cytoplasmic polyadenylation element-binding protein 1 (CPEB1) (22). miR-873 also plays an important role in EC, as it acts as a tumour suppressor by targeting heparin binding growth factor (HDGF) (23). Several studies focused on the tumour-suppressive mechanisms of miR-148b have shown that it is down-regulated in a number of cancers, such as colorectal (24), pancreatic (14) and gastric cancers (25). In the current study, miR-148b expression was lower in EC, and miR-148b inhibited cell proliferation and facilitated apoptosis. Consistent with previous reports, our findings indicated that miR 148b could be developed as a therapeutic target for the treatment of patients with EC.
DNA methylation is an important aspect of mammalian epigenetic gene regulation, and it is mainly catalysed by enzymes of the DNMT family, such as DNMT1, DNMT3A, and DNMT3B (26). Previous studies showed that epigenetic alterations are involved in the development and progression of human cancers, and these alterations are mediated by dysregulated expression of DNMT1 (27-29). DNMT1 is a target of miR-148a/152, and DNMT1 expression is inversely correlated with the expression of miR-148a/152 in breast cancer (30). Sawada et al. (31) reported that progressively increasing DNMT1 expression is involved in an early step of cervical carcinogenesis. Wang et al. (32) reported that miR-148a inhibits EMT and cell proliferation by establishing regulatory circuits between DNMT1 and epidermal growth factor receptor 3 (ERBB3)/AKT serine/threonine kinase 2 (AKT2)/c-myc in bladder cancer. Accumulating evidence suggests that several miRNAs, including miR-21, -152, -140, -148a, and -148b, target DNMT1 expression (14,33-35). Our findings showed that DNMT1 is also a target of miR-148b in EC, which is consistent with previous research results. Based on previous research findings, the role of miR-148b and DNMT1 in EC was further explored. The data showed that knock-down of DNMT1 partially rescued the miR-148b inhibitor-mediated effects on EC cell proliferation, apoptosis, and the expression of apoptosis-associated proteins.
We measured caspase-3 and cyclin D1 protein expression levels to investigate the mechanisms underlying the effects of miR-148b on cell proliferation and apoptosis. Based on previous research results, we know that caspases-3 plays an important role in the treatment of malignancies and that cyclin D1 is a significant cell cycle regulator in G1 phase (36-38). Many miRNAs have been reported to be involved in various cancers by regulating cyclin D1 expression; for example, miR-195 suppresses cell proliferation and cell cycle progression by directly targeting cyclin D1a in cervical cancer (39). Similarly, previous research has shown that inhibition of miR-643 expression could reduce apoptosis and caspase-3 activation in human epithelial colon cells (40). In this study, we also confirmed that miR-148b inhibited cyclin D1 expression and markedly decreased caspase-3. These results further demonstrated that miR-148b inhibits proliferation and promotes apoptosis in EC cells by down-regulating DNMT1.
Conclusions
Our work confirmed that miR-148b expression was down-regulated in both EC tissues and cells. We also demonstrated that miR-148b reduced cell proliferation and promoted apoptosis in EC cells by regulating DNMT1. Our research provides an innovative regulatory mechanism involving miR-148b in EC as well as a new potential target for EC treatment.
Acknowledgments
Funding: None.
Footnote
Conflicts of Interest: All authors have completed the ICMJE uniform disclosure form (available at http://dx.doi.org/10.21037/tcr.2019.12.79). The authors have no conflicts of interest to declare.
Ethical Statement: The authors are accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved. The study was conducted in accordance with the Declaration of Helsinki (as revised in 2013). The study was approved by the Ethics Committee of our hospital, and all patients provided informed consent.
Open Access Statement: This is an Open Access article distributed in accordance with the Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License (CC BY-NC-ND 4.0), which permits the non-commercial replication and distribution of the article with the strict proviso that no changes or edits are made and the original work is properly cited (including links to both the formal publication through the relevant DOI and the license). See: https://creativecommons.org/licenses/by-nc-nd/4.0/.
References
- Lee YC, Lheureux S, Oza AM. Treatment strategies for endometrial cancer: current practice and perspective. Curr Opin Obstet Gynecol 2017;29:47-58. [Crossref] [PubMed]
- Bray F, Ferlay J, Soerjomataram I, et al. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin 2018;68:394-424. [Crossref] [PubMed]
- Bendifallah S, Ballester M, Darai E. Endometrial cancer: Predictive models and clinical impact. Bull Cancer 2017;104:1022-31. [Crossref] [PubMed]
- Vale CL, Tierney J, Bull SJ, et al. Chemotherapy for advanced, recurrent or metastatic endometrial carcinoma. Cochrane Database Syst Rev 2012;CD003915. [PubMed]
- Bartel DP. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 2004;116:281-97. [Crossref] [PubMed]
- Feng Q, Zhang H, Nie X, et al. miRNA-149* suppresses liver cancer progression by down-regulating TRADD protein expression. Am J Pathol 2019; [Epub ahead of print].
- Yang M, Xiaoli T, Wang Z, et al. miR-125 inhibits colorectal cancer proliferation and invasion by targeting TAZ. Biosci Rep 2019; [Crossref] [PubMed]
- Dai Y, Lin X, Xu W, et al. miR-210-3p protects endometriotic cells from oxidative stress-induced cell cycle arrest by targeting BARD1. Cell Death Dis 2019;10:144. [Crossref] [PubMed]
- Cummins JM, Velculescu VE. Implications of micro-RNA profiling for cancer diagnosis. Oncogene 2006;25:6220-7. [Crossref] [PubMed]
- Rarani FZ, Borhani F, Rashidi B. Endometrial pinopode biomarkers: Molecules and microRNAs. J Cell Physiol 2018;233:9145-58. [Crossref] [PubMed]
- Herman JG, Baylin SB. Gene silencing in cancer in association with promoter hypermethylation. N Engl J Med 2003;349:2042-54. [Crossref] [PubMed]
- Parbin S, Pradhan N, Das L, et al. DNA methylation regulates Microtubule-associated tumor suppressor 1 in human non-small cell lung carcinoma. Exp Cell Res 2019;374:323-32. [Crossref] [PubMed]
- Liu J, Zhang X, Huang Y, et al. miR-200b and miR-200c co-contribute to the cisplatin sensitivity of ovarian cancer cells by targeting DNA methyltransferases. Oncol Lett 2019;17:1453-60. [PubMed]
- Azizi M, Teimoori-Toolabi L, Arzanani MK, et al. MicroRNA-148b and microRNA-152 reactivate tumor suppressor genes through suppression of DNA methyltransferase-1 gene in pancreatic cancer cell lines. Cancer Biol Ther 2014;15:419-27. [Crossref] [PubMed]
- Chen Y, Song YX, Wang ZN. The microRNA-148/152 family: multi-faceted players. Mol Cancer 2013;12:43. [Crossref] [PubMed]
- Song Y, Xu Y, Wang Z, et al. MicroRNA-148b suppresses cell growth by targeting cholecystokinin-2 receptor in colorectal cancer. Int J Cancer 2012;131:1042-51. [Crossref] [PubMed]
- Ding X, Liu J, Liu T, et al. miR-148b inhibits glycolysis in gastric cancer through targeting SLC2A1. Cancer Med 2017;6:1301-10. [Crossref] [PubMed]
- Lu L, Liu Q, Wang P, et al. MicroRNA-148b regulates tumor growth of non-small cell lung cancer through targeting MAPK/JNK pathway. BMC Cancer 2019;19:209. [Crossref] [PubMed]
- Mou Z, Xu X, Dong M, et al. MicroRNA-148b Acts as a Tumor Suppressor in Cervical Cancer by Inducing G1/S-Phase Cell Cycle Arrest and Apoptosis in a Caspase-3-Dependent Manner. Med Sci Monit 2016;22:2809-15. [Crossref] [PubMed]
- Yanokura M, Banno K, Iida M, et al. MicroRNAS in endometrial cancer: recent advances and potential clinical applications. Excli J 2015;14:190-8. [PubMed]
- Morice P, Leary A, Creutzberg C, et al. Endometrial cancer. Lancet 2016;387:1094-108. [Crossref] [PubMed]
- Xiong H, Chen R, Liu S, et al. MicroRNA-183 induces epithelial-mesenchymal transition and promotes endometrial cancer cell migration and invasion in by targeting CPEB1. J Cell Biochem 2018;119:8123-37. [Crossref] [PubMed]
- Wang Q, Zhu W. MicroRNA-873 inhibits the proliferation and invasion of endometrial cancer cells by directly targeting hepatoma-derived growth factor. Exp Ther Med 2019;18:1291-8. [PubMed]
- Wang G, Cao X, Lai S, et al. Altered p53 regulation of miR-148b and p55PIK contributes to tumor progression in colorectal cancer. Oncogene 2015;34:912-21. [Crossref] [PubMed]
- Wu M, Ye X, Wang S, et al. MicroRNA-148b suppresses proliferation, migration, and invasion of nasopharyngeal carcinoma cells by targeting metastasis-associated gene 2. Onco Targets Ther 2017;10:2815-22. [Crossref] [PubMed]
- Hervouet E, Peixoto P, Delage-Mourroux R, et al. Specific or not specific recruitment of DNMTs for DNA methylation, an epigenetic dilemma. Clin Epigenetics 2018;10:17. [Crossref] [PubMed]
- Qi X, Yu XJ, Wang XM, et al. Knockdown of KCNQ1OT1 Suppresses Cell Invasion and Sensitizes Osteosarcoma Cells to CDDP by Upregulating DNMT1-Mediated Kcnq1 Expression. Mol Ther Nucleic Acids 2019;17:804-18. [Crossref] [PubMed]
- Yu H, Zhang S, Ibrahim AN, et al. RCC2 promotes proliferation and radio-resistance in glioblastoma via activating transcription of DNMT1. Biochem Biophys Res Commun 2019;516:999-1006. [Crossref] [PubMed]
- Wang Y, Hu Y, Guo J, et al. miR-148a-3p Suppresses the Proliferation and Invasion of Esophageal Cancer by Targeting DNMT1. Genet Test Mol Biomarkers 2019;23:98-104. [Crossref] [PubMed]
- Xu Q, Jiang Y, Yin Y, et al. A regulatory circuit of miR-148a/152 and DNMT1 in modulating cell transformation and tumor angiogenesis through IGF-IR and IRS1. J Mol Cell Biol 2013;5:3-13. [Crossref] [PubMed]
- Sawada M, Kanai Y, Arai E, et al. Increased expression of DNA methyltransferase 1 (DNMT1) protein in uterine cervix squamous cell carcinoma and its precursor lesion. Cancer Lett 2007;251:211-9. [Crossref] [PubMed]
- Wang X, Liang Z, Xu X, et al. miR-148a-3p represses proliferation and EMT by establishing regulatory circuits between ERBB3/AKT2/c-myc and DNMT1 in bladder cancer. Cell Death Dis 2016;7:e2503. [Crossref] [PubMed]
- Wang XX, Zhang H, Li Y. Preliminary study on the role of miR148a and DNMT1 in the pathogenesis of acute myeloid leukemia. Mol Med Rep 2019;19:2943-52. [PubMed]
- Shi YK, Guo YH. miR-139-5p suppresses osteosarcoma cell growth and invasion through regulating DNMT1. Biochem Biophys Res Commun 2018;503:459-66. [Crossref] [PubMed]
- Azizi M, Fard-Esfahani P, Mahmoodzadeh H, et al. miR-377 reverses cancerous phenotypes of pancreatic cells via suppressing DNMT1 and demethylating tumor suppressor genes. Epigenomics 2017;9:1059-75. [Crossref] [PubMed]
- Garcia IJP, de Oliveira GC, de Moura Valadares JM, et al. New bufadienolides extracted from Rhinella marina inhibit Na,K-ATPase and induce apoptosis by activating caspases 3 and 9 in human breast and ovarian cancer cells. Steroids 2019;152:108490. [Crossref] [PubMed]
- John RR, Malathi N, Ravindran C, et al. Mini review: Multifaceted role played by cyclin D1 in tumor behavior. Indian J Dent Res 2017;28:187-92. [Crossref] [PubMed]
- Jeong CH, Ryu H, Kim DH, et al. Piperlongumine Induces Cell Cycle Arrest via Reactive Oxygen Species Accumulation and IKKbeta Suppression in Human Breast Cancer Cells. Antioxidants (Basel) 2019; [Crossref] [PubMed]
- Chang W, Wang Y, Li W, et al. MicroRNA-551b-3p inhibits tumour growth of human cholangiocarcinoma by targeting Cyclin D1. J Cell Mol Med 2019;23:4945-54. [Crossref] [PubMed]
- López-Rosas I, Lopez-Camarillo C, Salinas-Vera YM, et al. Entamoeba histolytica Up-Regulates MicroRNA-643 to Promote Apoptosis by Targeting XIAP in Human Epithelial Colon Cells. Front Cell Infect Microbiol 2019;8:437. [Crossref] [PubMed]


