
The antitumor effect of miR-448 in epithelial ovarian cancer
Introduction
Ovarian cancer (OC) is the seventh most commonly diagnosed cancer in world and the tenth most common in China (1). OC accounts for an estimated 239,000 annual new cases and 152,000 deaths worldwide (2). Although China has a relatively low incidence rate (4.1 per 100,000), the large population translated to an estimated 52,100 new cases and 22,500 related deaths in 2015 (3).
Tradition treatment of OC has included surgery, during which doctors remove all visible tumors, chemotherapy, and radiotherapy (4). Advances in traditional cytotoxic chemotherapies, such as intraperitoneal administration and dose-dense therapeutic regimens, are improving response rates, as are novel target agents such as bevacizumab or poly (ADP-ribose) polymerase (PARP) inhibitors (5-7).
However, these target agents have shown efficacy only in the early stages of some cases, and so more effective molecular targeting agents should be developed. Recently, microRNAs (miRNAs), which consist of short-sequence RNAs that do not encode a protein, have emerged as new biomarkers in the clinical diagnosis and treatment of OC (8).
MiRNA-448 exerts a tumor-suppressor role in numerous cancer types including breast (9-11), oral squamous cell (12), gastric (13), colorectal (14), and pancreatic ductal adenocarcinoma (15,16). However, the function of miR-448 in OC still remains unknown, and it is possible that the miRNA-448 also plays a tumor-suppressor role in OC. Epithelial ovarian cancer (EOC) is the most common type of OC. In this study, we collected 27 EOC samples and analyzed the function and expression of miR-448. We hope our findings may provide insight into the function of miR-448 in OC. We present the following article in accordance with the MDAR reporting checklist (available at http://dx.doi.org/10.21037/tcr-20-2464).
Methods
OC tissues samples
In total, the EOC samples and paired adjacent normal tissues from 27 female patients (age: 67±6.6 years) were collected from the Department of Gynecology and Obstetrics, Development and Related Diseases of Women and Children Key Laboratory of Sichuan Province, West China Women’s and Children’s Hospital.
The ethical approval of this study was confirmed by the medical ethics committee of Sichuan University (Approval ID: 20172356). The study was conducted in accordance with the Declaration of Helsinki (as revised in 2013). The 27 EOC patients from whom samples were obtained provided legal consent according to the relevant regulations.
Cell lines
One EOC cell line, SKOV3 (ATCC®HTB-77TM) was purchased from the American Type Culture Collection (ATCC, Manassas, VI, USA) and a normal epithelial cell control, IOSE29, (normal immortalized human ovarian surface epithelial cells) were acquired form the cell bank of Sichuan University. All the three cell lines were cultured in a laboratory CO2 incubator at 37 °C in 5% CO2. The cell culture medium was a mixture of RPMI-1640 and 10% fetal bovine serum (FBS) (v/v: 9:1) (RPMI-1640, cat. no 72400120; FBS cat. no 10100154. Thermo Fisher Scientific, Inc., Waltham, MA, USA) placed in 6-well plates.
Quantitative real-time polymerase chain (qRT-PCR)
The levels of miR-448 in the 27 EOC samples and SKOV3 and IOSE29 cell lines were detected by qRT-PCR analysis. For tissues, a mixture of 200 mg of tissue and 2 mL of TRIzol reagent was homogenized by a homogenizer in ice (cat.no.12183026, Thermo Fisher). Cells were rinsed with ice cold phosphate-buffered saline (PBS). Total RNA was extracted using TRIzolTM LS Reagent (cat.no.10296028, InvitrogenTM). The cDNA was synthesized by cDNA synthesis kit (cat.no.18091200, InvitrogenTM). The primer sequence for miR-448 and U6 were listed as followed: miR-448 F: 5'-TTATTGCGATGTGTTCCTTATG-3', R: 5'-ATGCATGCCACGGGCATATACACT-3'; U6 F: 5'-GCTTCGGCAGCACATATACTAAAAT-3', R: 5'-CGCTTCACGAATTTGCGTGTCAT-3'. The differences in target gene expression between the control group and the experimental group were determined using the 2−ΔΔCt method.
Transfection
miR-448 expression in cells was induced by miR-448 mimic transfection and suppressed by miR-448 antisense oligonucleotides (ASO) transfection. miR-448 mimics (UUGCAUAUGUAGGAUGUCCCAU) (miR10001532-1-5), miR-448 ASO (miR20001532-1-5), and control miRNA were designed and purchased from Shanghai Shengong Biotech Company (https://www.sangon.com/) (Shanghai, China). Lipofectamine® 2000 reagent (Thermo Fisher Scientific, USA) was applied for transfection. Seeds were 70–90% confluent at transfection, and Lipofectamine® reagent and miRNA-448 mimics or miRNA-448 ASO in Opti-MEM medium was diluted separately. Then, miRNA-lipid complex was added to the cells and co-cultured for 12 h.
Cell growth assay
The cell proliferation was tested at the indicated time point by MTT assay. SKOV3 cells were seeded according to the manufacturer’s instructions (cat.no. ab211091, abcam, Cambridge, United Kingdom). After miR-448 mimics or miR-448 ASO transfection, the medium were aspirated carefully. Then, 50 µL of serum-free medium and 50 µL of MTT solution were added into each well. The whole plate was incubated at 37 °C for 3 h. After incubation, 150 µL of MTT was added to the solvent in each well. The plates were wrapped in foil and shaken on an orbital shaker for 15 min. Occasionally, pipetting of the liquid was required to fully dissolve the MTT formazan. The measured absorbance at OD 590 nm is proportional to the number of viable cells., and the plate was read within 1 h.
Apoptosis
The apoptosis rate was assayed by annexin V-FITC and propidium iodide (PI) double staining. Specifically, cells (5×105) were collected by centrifugation and resuspended in 500 µL of 1× binding buffer. Then, 5 µL of annexin was added to V-FITC, which was followed by incubation for 30 min at room temperature, and the further addition of 1 µL of PI before analysis on a fluorescence-activated cell sorting (FACS) analyzer (FACSVerse, BD, Franklin Lakes, NJ, USA).
Prediction of the putative targets of miR-448
For bioinformatics prediction, Targetscan online software was used. TargetScan is a web server that predicts biological targets of miRNAs by searching for the presence of sites that match the seed region of each miRNA (17-19).
Dual luciferase reporter assays
Initially, the 3'UTR of zinc finger E-box-binding homeobox 2 (ZEB2) and mutated controls were prepared and inserted into the pGL3 vector (Promega, USA) with the restriction enzymes (BanII). Then, miR-448 mimics (50 nM) and wild type or pGL3 vector were transfected into the SKOV3 cells successively by Lipofectamine 2000. The assays for firefly luciferase activity and Renilla luciferase were performed sequentially using one reaction tube according to the manufacturer’s instructions (Invitrogen, USA).
Western blotting analysis
Cells were washed by phosphate-buffered saline (PBS) on ice, then scraped using a cold plastic cell scraper, and the cell suspension was gently transferred into a pre-cooled microcentrifuge tube with lysis buffer. Next, the cells were centrifuged at 12,000 rpm at 4 °C for 20 min. The protein concentration was determined by bicinchonic acid (BCA) disodium salt (ab146331, Abcam). Next, 20 µg of total protein was loaded into the SDS-PAGE gel wells (10%), and the electrophoresis were performed at 90 V for 2 h. Then, protein was transferred from the gel to the membrane. For ZEB2 analysis, the membrane was blocked at 4 °C overnight in blocking buffer, and washed by tris-buffered saline with Tween20 (TBST). After this, the membrane was incubated with ZEB2 antibody (cat.no, ab138222, Abcam, Cambridge, UK) at a dilution of 1:1,000 at room temperature for 2 h. Then, the whole membrane was incubated by the secondary antibody immunoglobulin G (IgG) (ab7090, 1:2,000 dilution, Abcam) at 4 °C overnight. After being washed by TBST three times, the membrane was mixed with ECL Western Blotting Substrate kit (cat.no, ab65623, Abcam) at a 1:1 ratio, incubated for 3 min, and get the blot in a X-ray film.
Statistical and analysis
Each experiment was performed independently 3 times. Data are shown as mean ± standard deviation (SD). A two-tailed Student’s t-test and an analysis of variance (ANOVA) test were used to test the mean value between two groups and the mean value across three or more groups or more, respectively. P value <0.05 indicated statistical significance. All statistical analyses were performed using the SPSS software (version 19.0, IBM Corporation, Armonk, NY, USA).
Results
Low miR-448 expression was associated with low survival rate
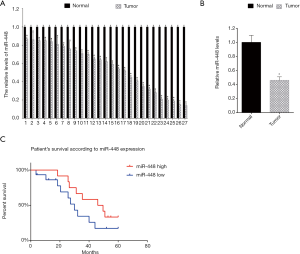
Initially, we analyzed the miR-448 expression in 27 EOC samples. The miR-448 levels were analyzed by qRT-PCR. The data showed that miR-448 showed lower levels in tumor tissues compared with the adjacent normal tissues (Figure 1A). Then, we analyzed the mean value of the levels of miR-448 in EOC tissues and adjacent normal tissues, and found that the mean levels of miR-448 in EOC tissues was lower than the mean levels of adjacent normal tissues (Figure 1B). Next, we analyzed patient survival according to the miR-448 levels. The 27 EOC patients were followed-up for 60 months from the day when they received the surgery. We separated the 27 EOC patients into two groups: the miR-488 high expression group and the miR-448 low expression group. We found that the EOC patient group with high miR-448 expression showed a higher patient survival rate (Figure 1C).
miR-448 inhibited SKOV3 proliferation and promoted apoptosis
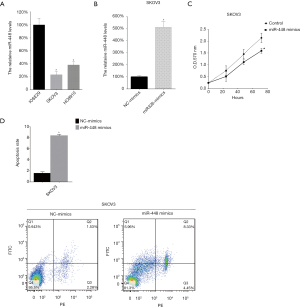
We further analyzed the miR-448 level in two OC cell lines (SKOV3). The miR-448 levels in cells were tested by qRT-PCR. We found that, compared with normal ovarian epithelial cells (NOE), there were lower miR-448 levels in SKOV3 (Figure 2A). Then, we introduced miR-448 mimics to the two cell lines to upregulate the miR-448 levels, and this effect was tested by qRT-PCR (Figure 2B). After this, we tested the cellular proliferation of SKOV3 cells following miR-448 mimic transfection and found that the upregulation of miR-448 inhibited the growth of SKOV3 cells (Figure 2C). Subsequently, we tested the cell apoptosis rate of SKOV3 cells following miR-448 up-regulation and found that miR-448 mimics promoted the cell apoptosis of the two cell lines (Figure 2D).
Downregulation of miR-448 promoted SKOV3 proliferation
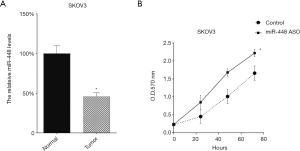
We transfected miR-448 ASO into the two cell lines to downregulate miR-448 expression, and its effect was tested by qRT-PCR (Figure 3A). Then, we tested the cellular proliferation of SKOV3 cells following miR-448 ASO transfection and found that transfection of miR-448 ASO promoted the growth of SKOV3 cells (Figure 3B).
ZEB2 was a target gene of miR-448
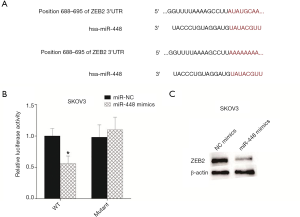
miRNA performs its role by binding the 3'UTR of mRNA. A previous study showed that miR-448 inhibited breast cancer by targeting zinc finger E-box binding protein homeobox 2 (ZEB2) (11). Thus, it is highly possible that miR-448 may inhibit cellular growth by targeting ZEB2. We listed the binding sites between miR-448 and ZEB2 and mutated the binding site in 3'UTR of mRNA (Figure 4A). Then, this mutated site was cloned into the reported plasmid. For detecting the interaction between miR-448 and the ZEB2, miR-448 mimics and reporter plasmid were transfected into SKOV3 cells sequentially. The activity data were tested 16 h following transfection. The data showed that the upregulation of miR-448 suppressed luciferase activity, whereas the mutated binding site restored it, indicating that miR-448 targets ZEB2 in SKOV3 cells (Figure 4B). As the luciferase assay showed that miR-448 could target the 3'UTR of ZEB2, we wondered whether miR-448 inhibits ZEB2 protein levels, and thus transfected miR-448 to SKOV3 cells and assayed the ZEB2 protein via Western blot. Indeed, this showed that miR-488 suppresses ZEB2 expression (Figure 4C).
Discussion
In this study, we tested the role of miR-448 in EOC. This study found that miR-448 plays a suppressive role in EOC. Enforced miR-448 expression inhibited cell growth and the promoted cell apoptosis rate. Our data indicate that the ZEB2 may be a target gene of miR-448.
Our data also point to miR-448 as an indicator of EOC patient prognosis. The patient group with high miR-448 levels showed a higher survival rate. Our data also proved that ZEB2 is at least one of the target genes of miR-448. Consistent with this, previous studies showed that an elevated level of ZEB2 correlated with poor prognosis in ovarian cancer (20,21). Importantly, ZEB2 is a key factor in epithelial-mesenchymal transition (EMT) (22-24). Importantly, growing evidence shows that EMT induces chemotherapy resistance and stemness, and the underlying molecular mechanism of EMT inducing chemotherapy resistance involved with EMT results lower drug uptake, EMT enhanced repair of carboplatin-induced DNA damage, and decreased apoptosis (25).
It is highly possible that miR-448 plays an important role in the EMT of EOC. Our findings further show that the miR-448 level in EOC tissues could be used as an indicator of prognosis, but it is not appropriate for the diagnosis of EOC. To confirm our speculations, further research will involve collecting fresh serum from EOC patients to test whether miR-448 in serum is related to EOC patient prognosis.
Besides ZEB2, miR-448 may have other target genes. In hepatocellular carcinoma (HCC) miR-448 was also found to inhibit EMT by regulating rho-associated coiled-coil-containing protein kinase (ROCK2) (26). Interestingly, ROCK2 is targeted by miR-139-5p in OC (27), and miR-139-5p could also target ZEB2 in HCC (28). Thus, it is possible that miR-139-5p and miR-448 regulate ROCK2 and ZEB2 in OC.
Previous studies have indicated that a miRNA-gene network exists in OC (29). We surmise that miR-448 is a key miRNA and plays an important role with miR-139-5p by targeting ZEB2 and/or ROCK2. Overall, this study indicates that miR-448 has an anti-tumor role in EOC.
Acknowledgments
Funding: None.
Footnote
Reporting Checklist: The authors have completed the MDAR reporting checklist. Available at http://dx.doi.org/10.21037/tcr-20-2464
Data Sharing Statement: Available at http://dx.doi.org/10.21037/tcr-20-2464
Conflicts of Interest: All authors have completed the ICMJE uniform disclosure form (available at http://dx.doi.org/10.21037/tcr-20-2464). The authors have no conflicts of interest to declare.
Ethical Statement: The authors are accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved. The ethical approval of this study was confirmed by the medical ethics committee of Sichuan University (No. 20182356). The study was conducted in accordance with the Declaration of Helsinki (as revised in 2013). The 27 EOC patients from whom samples were obtained provided legal consent according to the regulations of Sichuan University.
Open Access Statement: This is an Open Access article distributed in accordance with the Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License (CC BY-NC-ND 4.0), which permits the non-commercial replication and distribution of the article with the strict proviso that no changes or edits are made and the original work is properly cited (including links to both the formal publication through the relevant DOI and the license). See: https://creativecommons.org/licenses/by-nc-nd/4.0/.
References
- Reid BM, Permuth JB, Sellers TA. Epidemiology of ovarian cancer: a review. Cancer Biol Med 2017;14:9-32. [Crossref] [PubMed]
- Gorodnova TV, Sokolenko AP, Kuligina E, et al. Principles of clinical management of ovarian cancer. Chin Clin Oncol 2018;7:56. [Crossref] [PubMed]
- Said N. Screening of three-dimensional spheroids of ovarian cancer: identification of novel therapeutics targeting stemness and chemoresistance. Ann Transl Med 2018;6:S26. [Crossref] [PubMed]
- Cannistra SA. Cancer of the ovary. N Engl J Med 2004;351:2519-29. [Crossref] [PubMed]
- Xue C, Zhu D, Chen L, et al. Expression and prognostic value of PD-L1 and PD-L2 in ovarian cancer. Transl Cancer Res 2019;8:111-9. [Crossref]
- Meehan RS, Chen AP. New treatment option for ovarian cancer: PARP inhibitors. Gynecol Oncol Res Pract 2016;3:3. [Crossref] [PubMed]
- Kim JY, Cho CH, Song HS. Targeted therapy of ovarian cancer including immune check point inhibitor. Korean J Intern Med 2017;32:798-804. [Crossref] [PubMed]
- Deb B, Uddin A, Chakraborty S. miRNAs and ovarian cancer: An overview. J Cell Physiol 2018;233:3846-54. [Crossref] [PubMed]
- Bamodu OA, Huang WC, Lee WH, et al. Aberrant KDM5B expression promotes aggressive breast cancer through MALAT1 overexpression and downregulation of hsa-miR-448. BMC Cancer 2016;16:160. [Crossref] [PubMed]
- Jiang X, Zhou Y, Sun AJ, et al. NEAT1 contributes to breast cancer progression through modulating miR-448 and ZEB1. J Cell Physiol 2018;233:8558-66. [Crossref] [PubMed]
- Ma P, Ni K, Ke J, et al. miR-448 inhibits the epithelial-mesenchymal transition in breast cancer cells by directly targeting the E-cadherin repressor ZEB1/2. Exp Biol Med (Maywood) 2018;243:473-80. [Crossref] [PubMed]
- Shen L, Liu L, Ge L, et al. miR-448 downregulates MPPED2 to promote cancer proliferation and inhibit apoptosis in oral squamous cell carcinoma. Exp Ther Med 2016;12:2747-52. [Crossref] [PubMed]
- Wu X, Tang H, Liu G, et al. miR-448 suppressed gastric cancer proliferation and invasion by regulating ADAM10. Tumour Biol 2016;37:10545-51. [Crossref] [PubMed]
- Li B, Ge L, Li M, et al. miR-448 suppresses proliferation and invasion by regulating IGF1R in colorectal cancer cells. Am J Transl Res 2016;8:3013-22. [PubMed]
- Yu DL, Zhang T, Wu K, et al. MicroRNA-448 suppresses metastasis of pancreatic ductal adenocarcinoma through targeting JAK1/STAT3 pathway. Oncol Rep 2017;38:1075-82. [Crossref] [PubMed]
- Zhao L, Kong H, Sun H, et al. LncRNA-PVT1 promotes pancreatic cancer cells proliferation and migration through acting as a molecular sponge to regulate miR-448. J Cell Physiol 2018;233:4044-55. [Crossref] [PubMed]
- Agarwal V, Bell GW, Nam JW, et al. Predicting effective microRNA target sites in mammalian mRNAs. Elife. 2015;4:e05005. [Crossref] [PubMed]
- Garcia DM, Baek D, Shin C, et al. Weak seed-pairing stability and high target-site abundance decrease the proficiency of lsy-6 and other microRNAs. Nat Struct Mol Biol 2011;18:1139-46. [Crossref] [PubMed]
- Friedman RC, Farh KK, Burge CB, et al. Most mammalian mRNAs are conserved targets of microRNAs. Genome Res 2009;19:92-105. [Crossref] [PubMed]
- Prislei S, Martinelli E, Zannoni GF, et al. Role and prognostic significance of the epithelial-mesenchymal transition factor ZEB2 in ovarian cancer. Oncotarget 2015;6:18966-79. [Crossref] [PubMed]
- Yan Z, Tian X, Wang R, et al. Title Prognosis Significance of ZEB2 and TGF-β1 as well as Other Clinical Characteristics in Epithelial Ovarian Cancer. Int J Gynecol Cancer 2017;27:1343-9. [Crossref] [PubMed]
- DaSilva-Arnold SC, Kuo CY, Davra V, et al. ZEB2, a master regulator of the epithelial-mesenchymal transition, mediates trophoblast differentiation. Mol Hum Reprod 2019;25:61-75. [Crossref] [PubMed]
- Hanrahan K, O'Neill A, Prencipe M, et al. The role of epithelial-mesenchymal transition drivers ZEB1 and ZEB2 in mediating docetaxel-resistant prostate cancer. Mol Oncol 2017;11:251-65. [Crossref] [PubMed]
- Korpal M, Lee ES, Hu G, et al. The miR-200 family inhibits epithelial-mesenchymal transition and cancer cell migration by direct targeting of E-cadherin transcriptional repressors ZEB1 and ZEB2. J Biol Chem 2008;283:14910-4. [Crossref] [PubMed]
- Loret N, Denys H, Tummers P, et al. The Role of Epithelial-to-Mesenchymal Plasticity in Ovarian Cancer Progression and Therapy Resistance. Cancers (Basel) 2019;11:838. [Crossref] [PubMed]
- Zhu H, Zhou X, Ma C, et al. Low Expression of miR-448 Induces EMT and Promotes Invasion by Regulating ROCK2 in Hepatocellular Carcinoma. Cell Physiol Biochem 2015;36:487-98. [Crossref] [PubMed]
- Wang Y, Li J, Xu C, et al. MicroRNA-139-5p Inhibits Cell Proliferation and Invasion by Targeting RHO-Associated Coiled-Coil-Containing Protein Kinase 2 in Ovarian Cancer. Oncol Res 2018;26:411-20. [Crossref] [PubMed]
- Qiu G, Lin Y, Zhang H, et al. miR-139-5p inhibits epithelial-mesenchymal transition, migration and invasion of hepatocellular carcinoma cells by targeting ZEB1 and ZEB2. Biochem Biophys Res Commun 2015;463:315-21. [Crossref] [PubMed]
- Zhang S, Ng MK. Gene-microRNA network module analysis for ovarian cancer. BMC Syst Biol 2016;10:117. [Crossref] [PubMed]
(English Language Editor: J. Gray)


